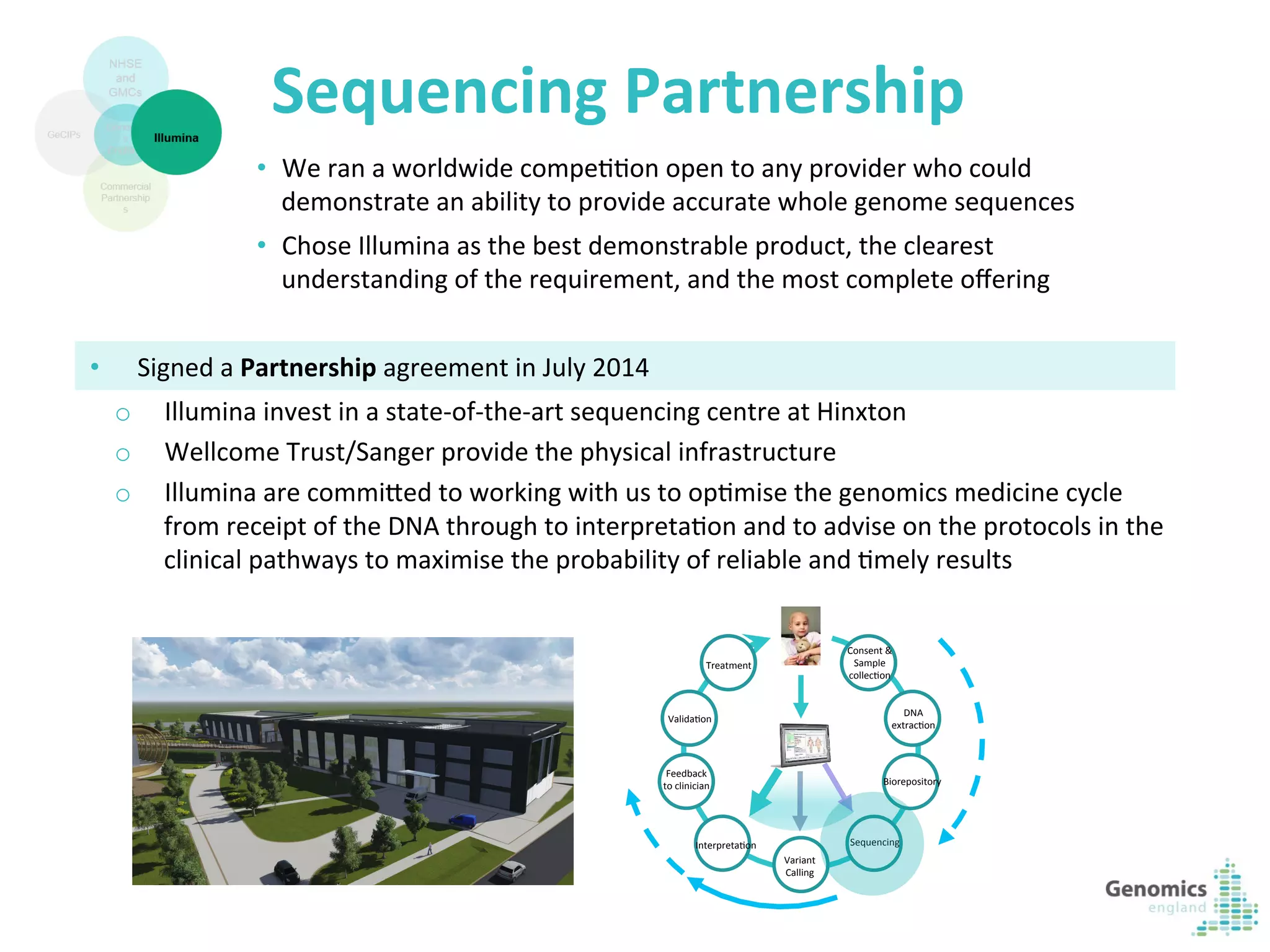
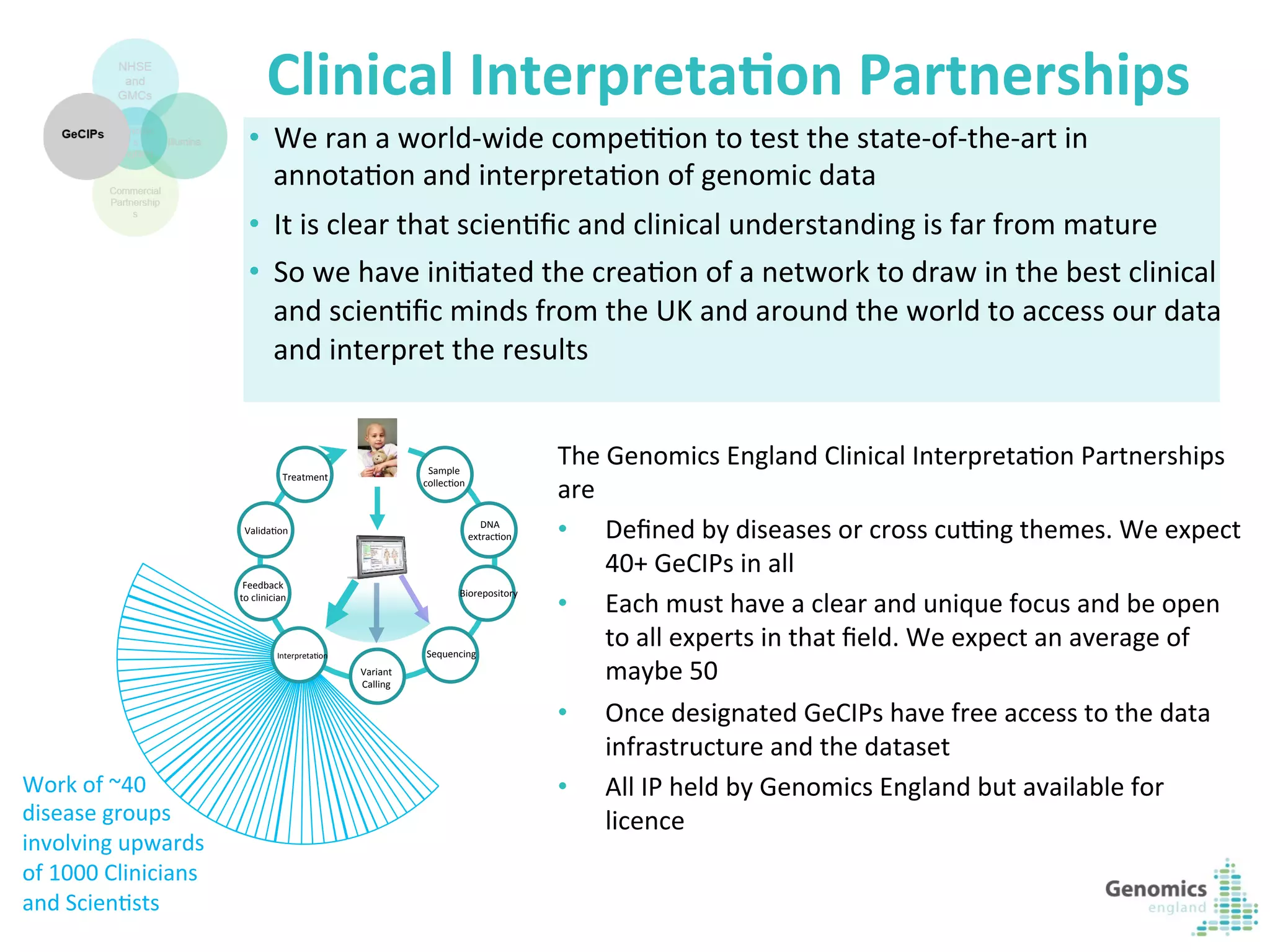
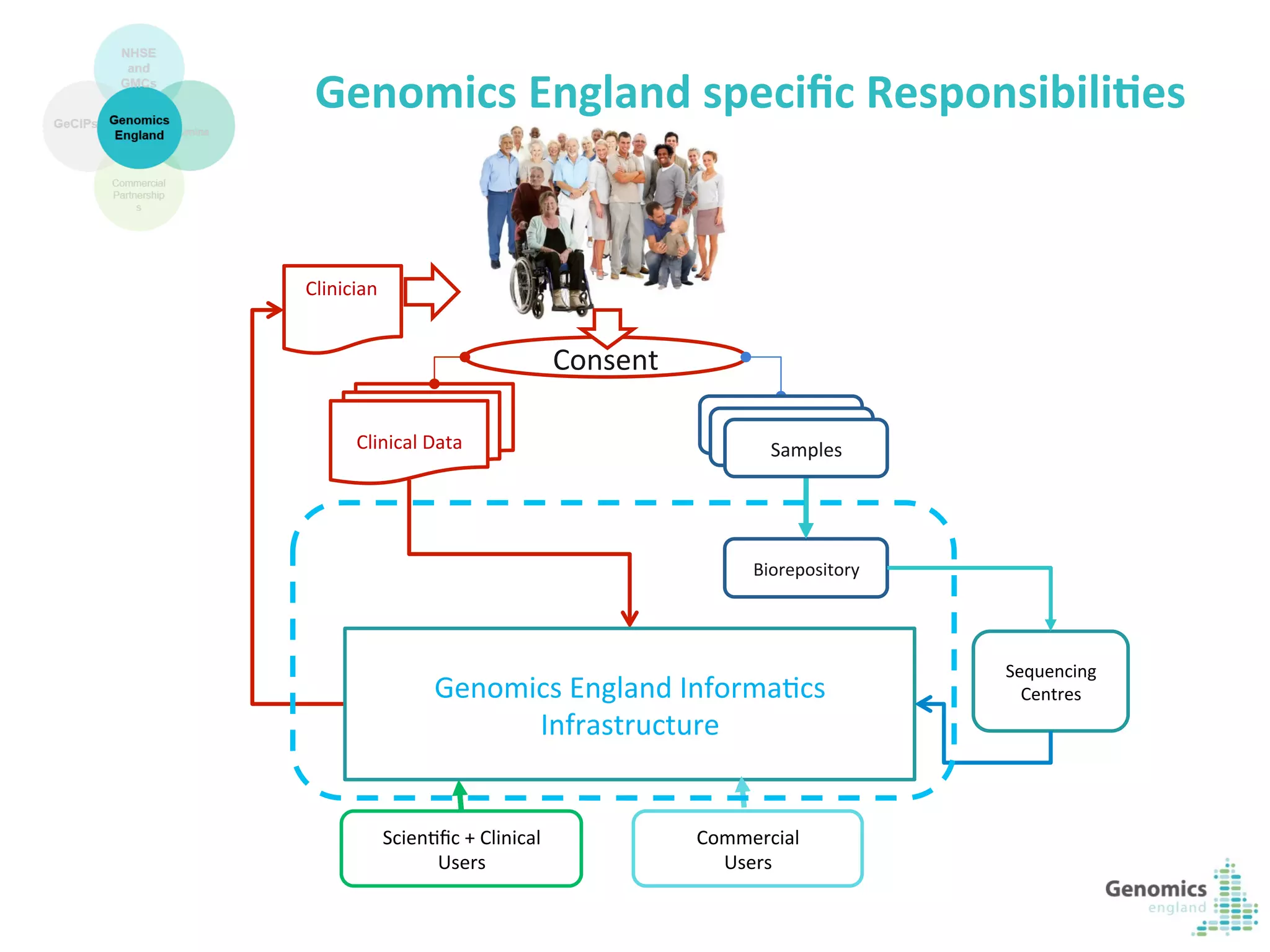
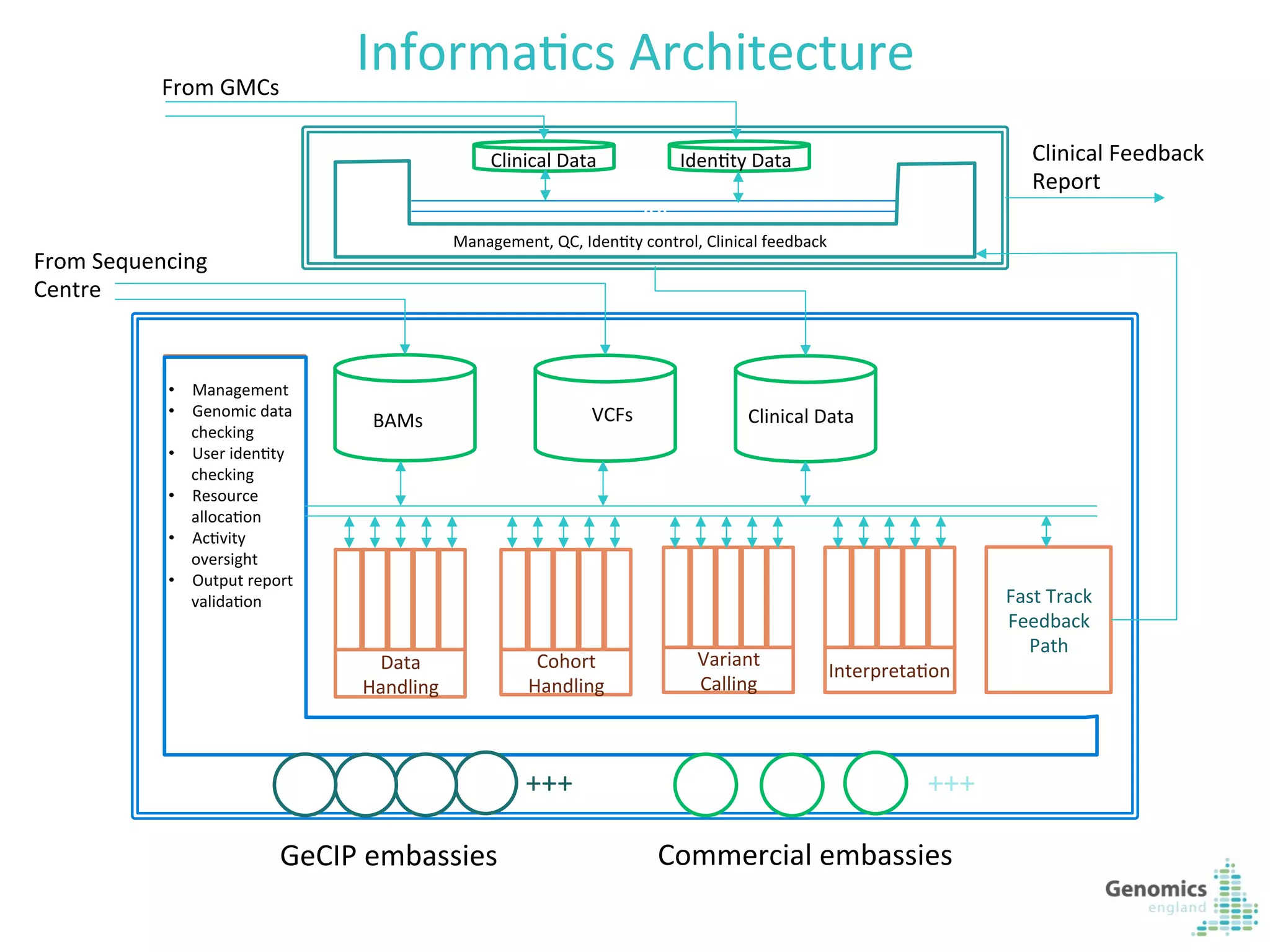
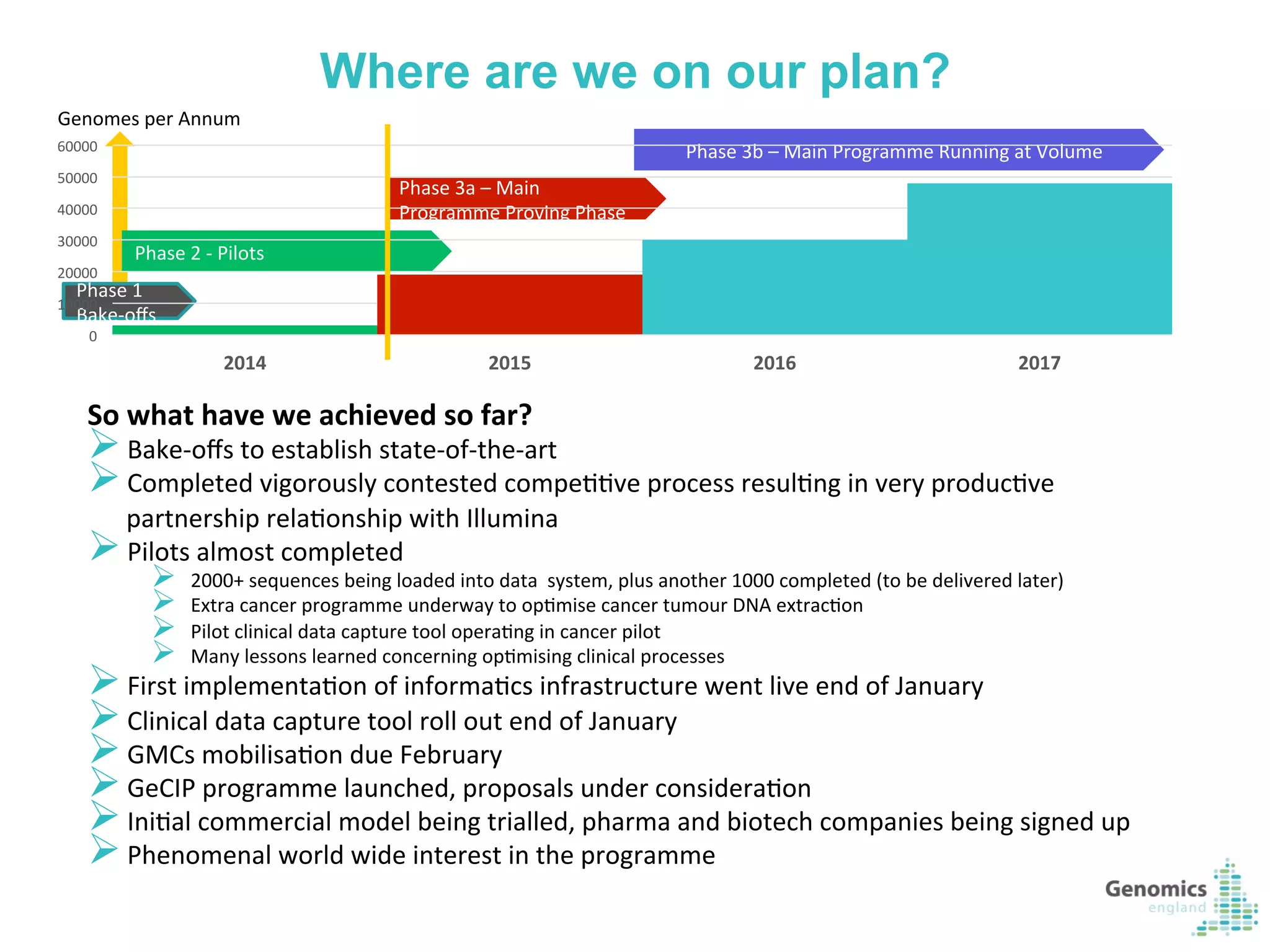
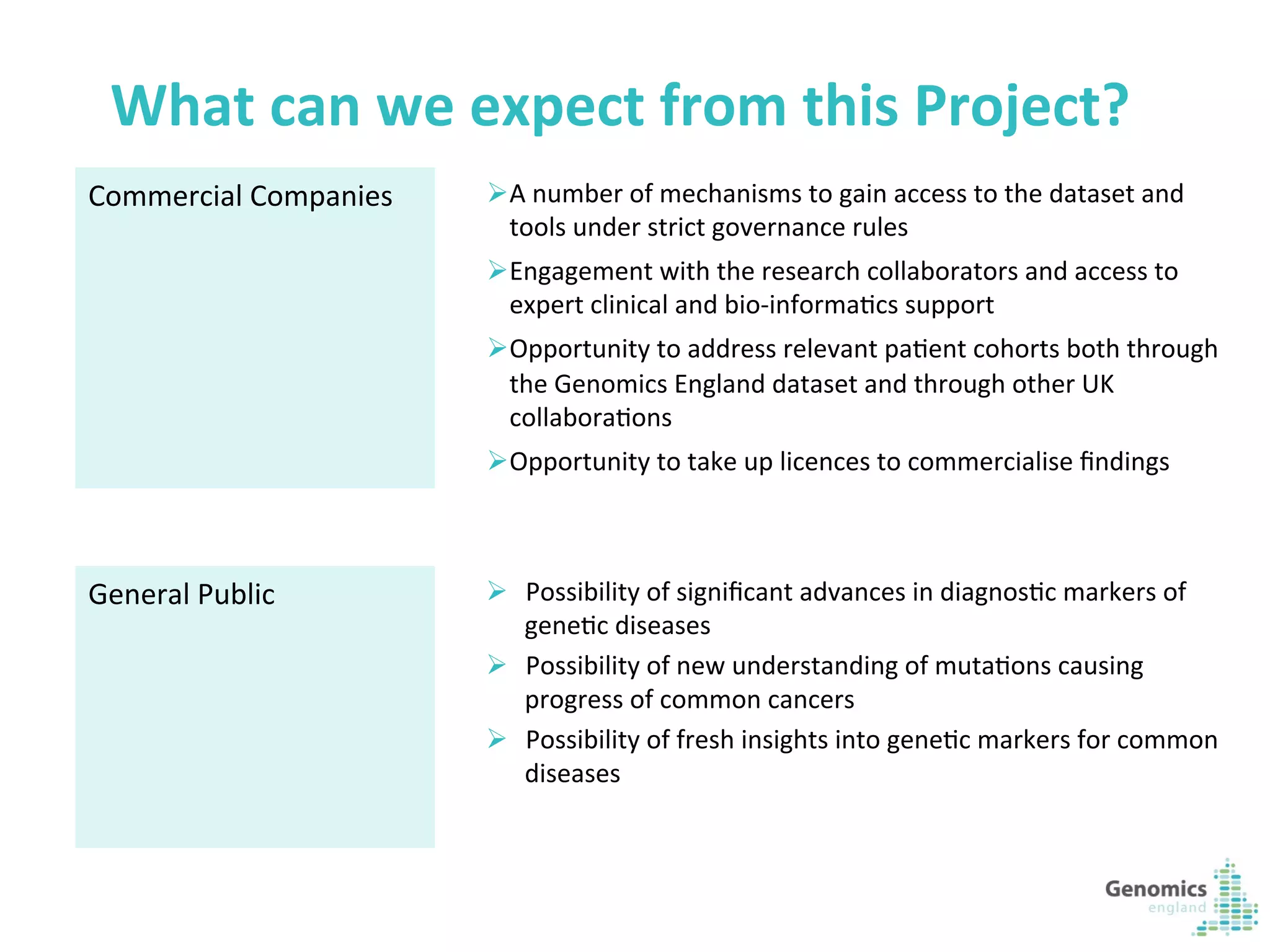
The document describes the UK 100,000 Genomes Project, which aims to sequence 100,000 genomes from NHS patients by 2017. It will provide benefits to patients through new treatments and insights, contribute to the UK economy through commercial activity in genomics, and help integrate genomic medicine into clinical practice in the NHS. Genomics England was established to lead the project and work with partners including NHS England, 11 Genomic Medicine Centres, and commercial companies. The project involves genome sequencing, analysis, and returning results to clinicians to inform treatment.