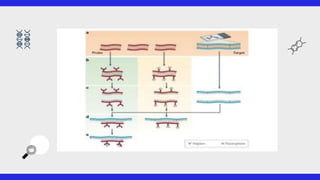
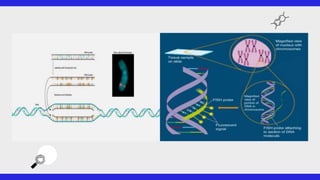
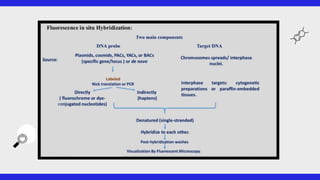
This document provides an overview of FISH (fluorescence in situ hybridization), SNPs (single nucleotide polymorphisms), and ESTs (expressed sequence tags). FISH allows visualization of specific DNA sequences on chromosomes. SNPs are variations in a single DNA nucleotide that can provide information about disease risk. ESTs are short DNA sequences that can be used to identify genes being expressed in a cell. All three are important tools in molecular biology and genetics research.