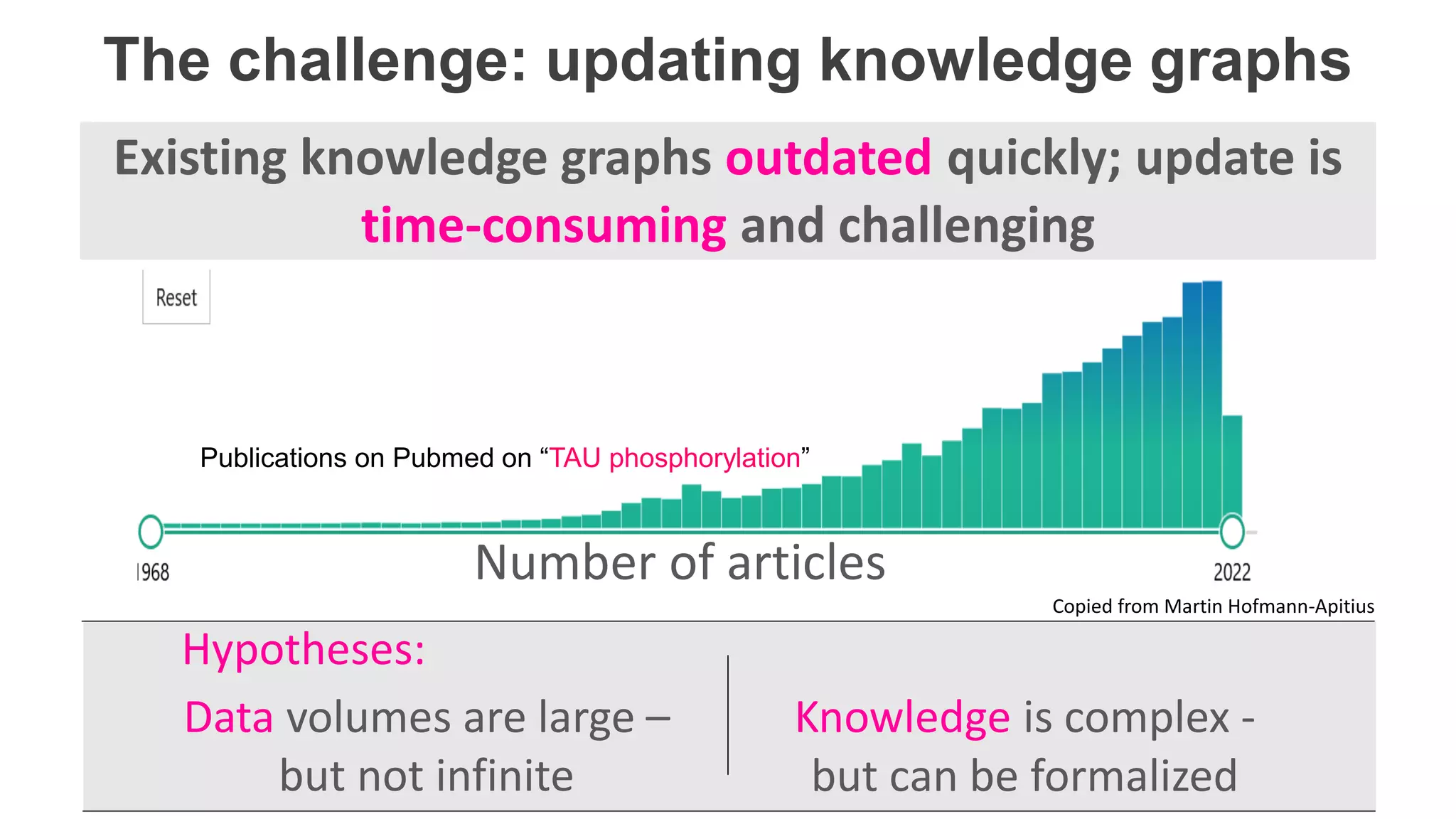
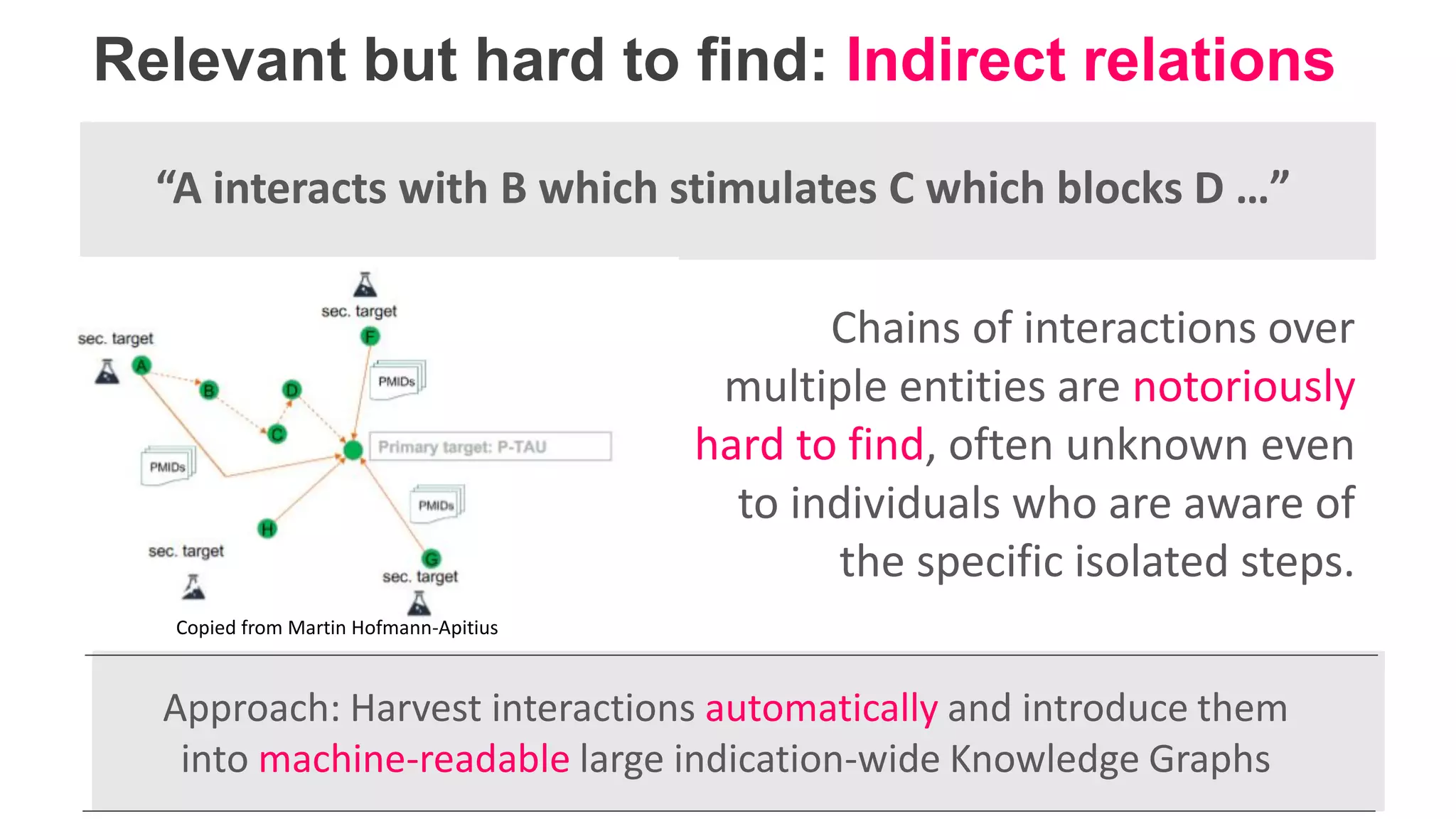
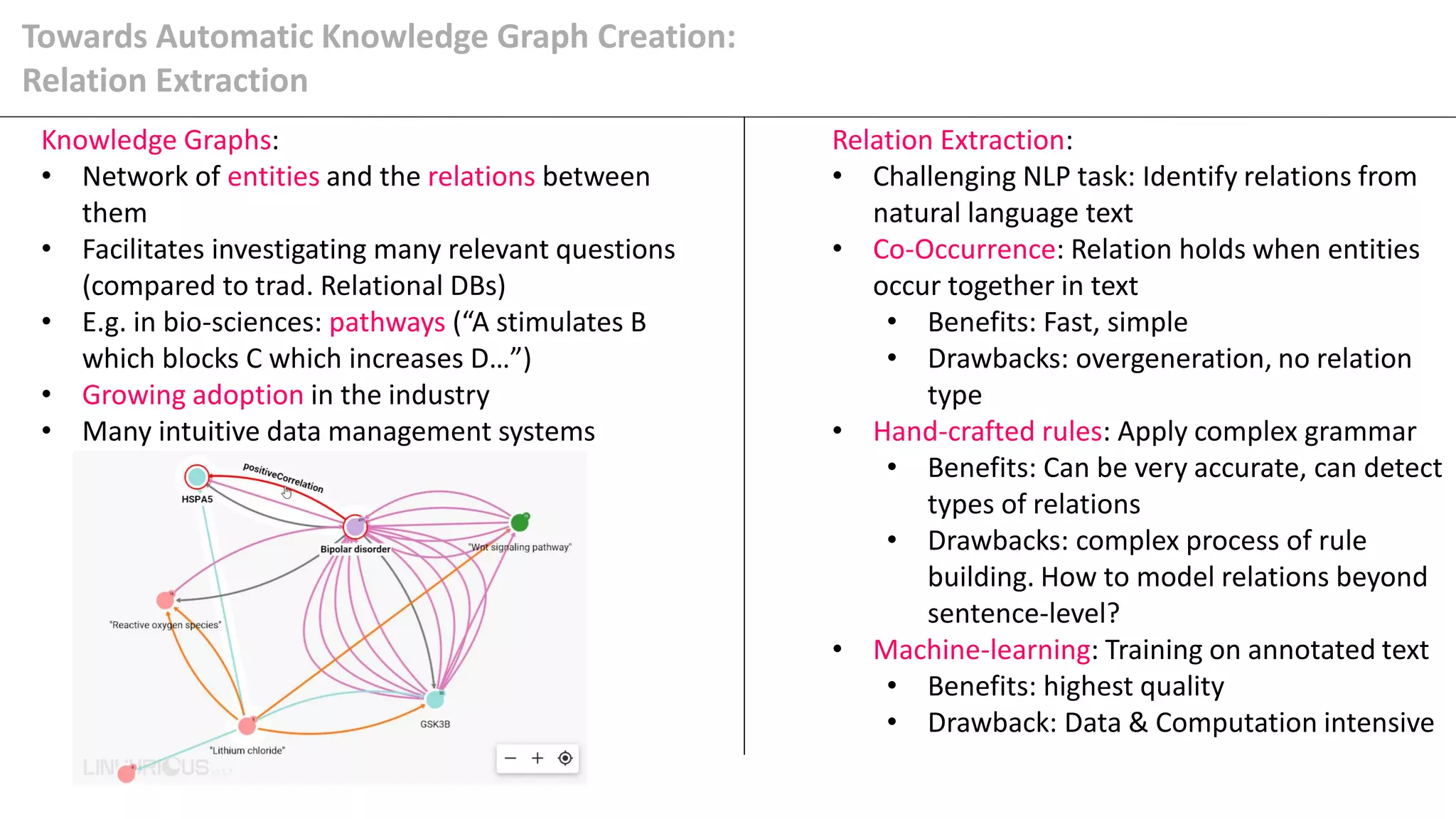
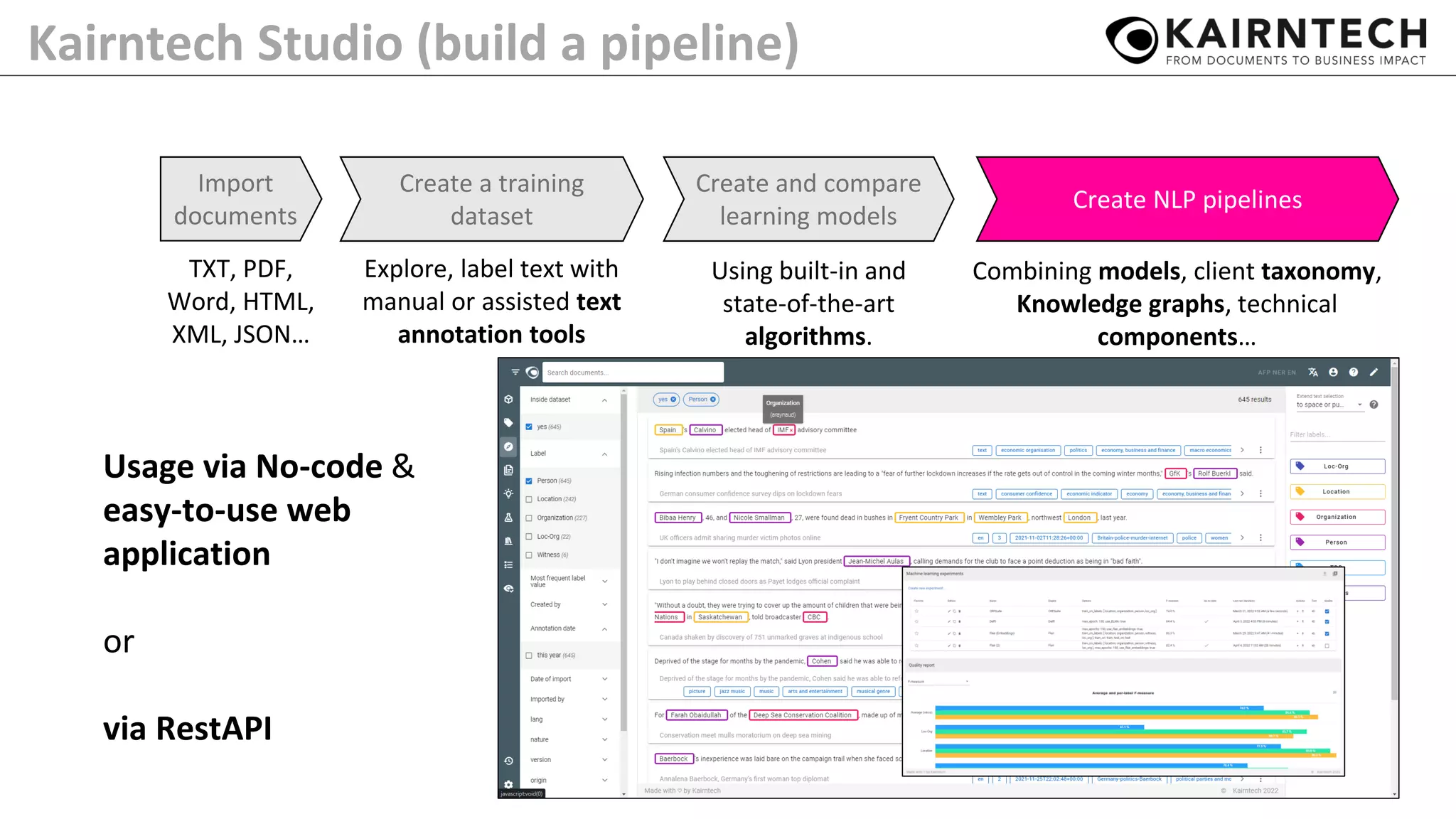
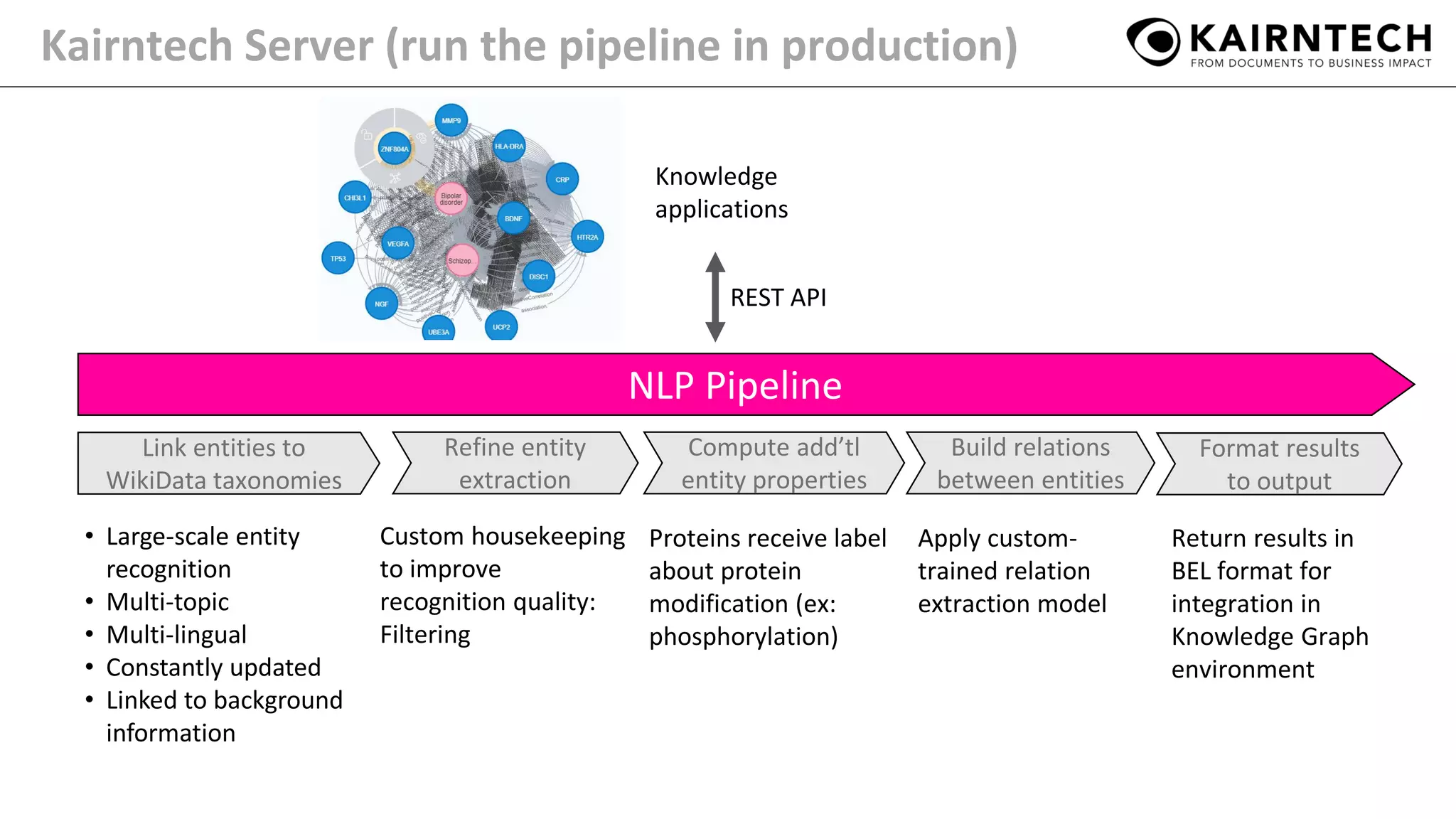
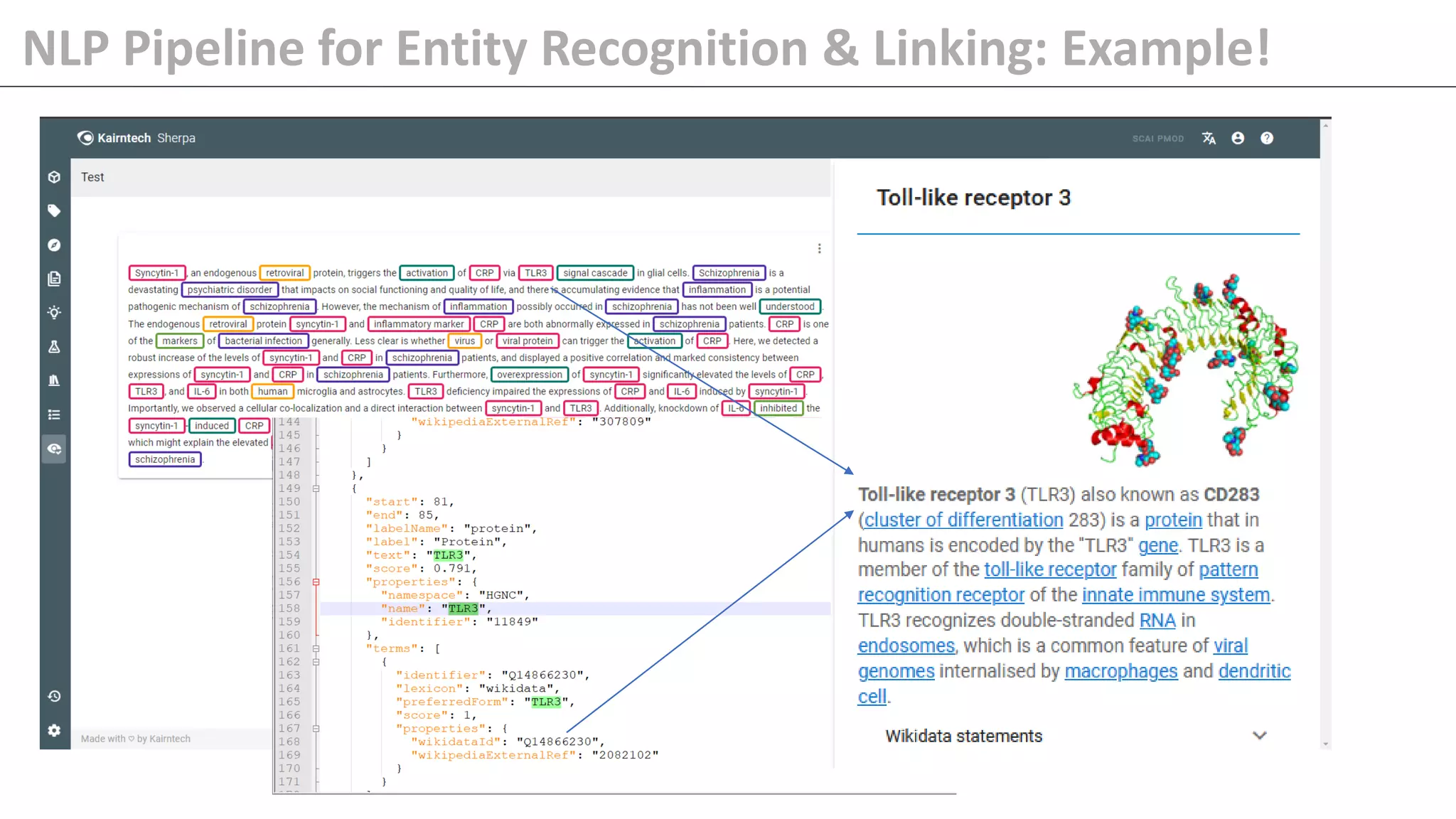
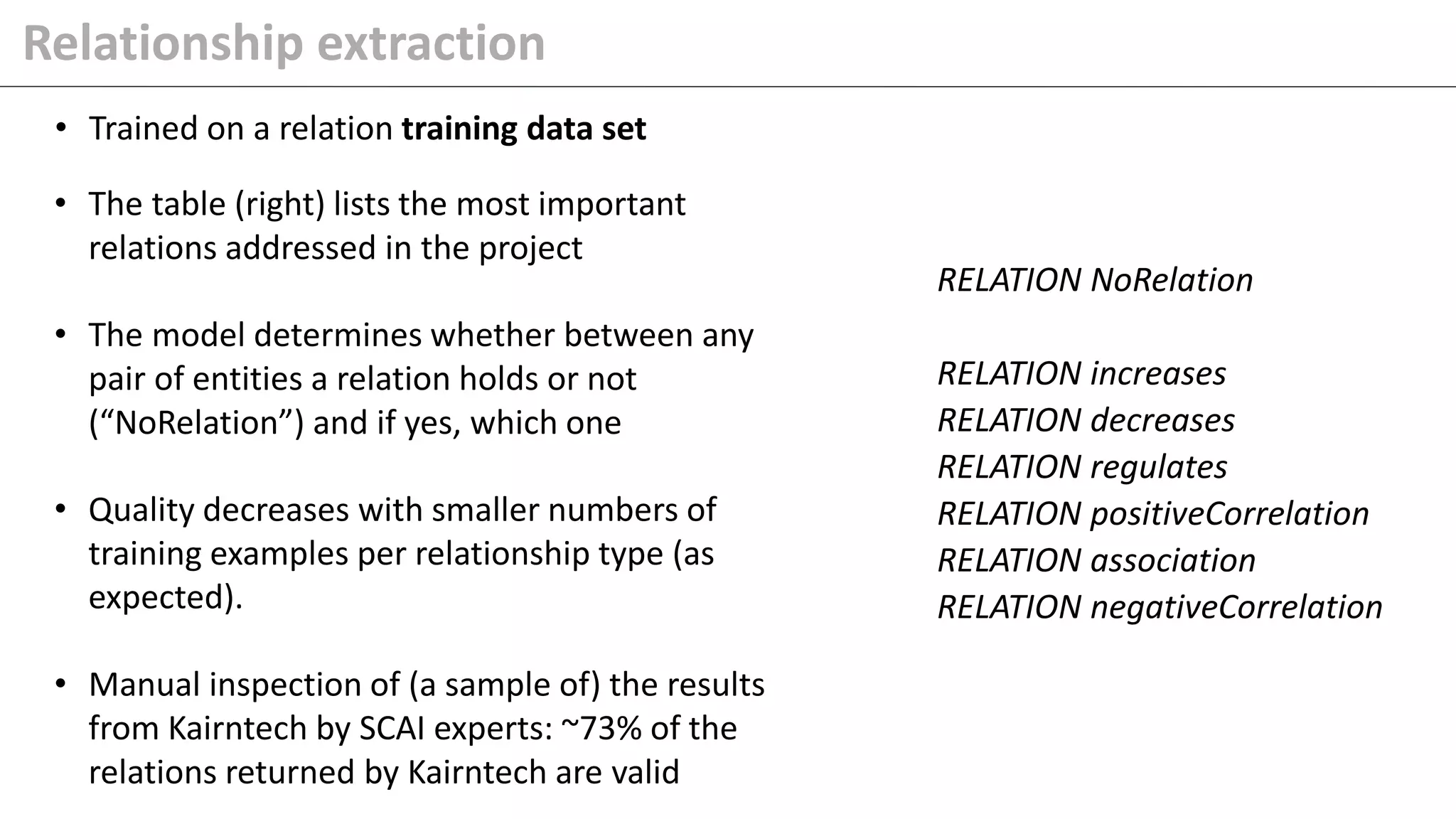
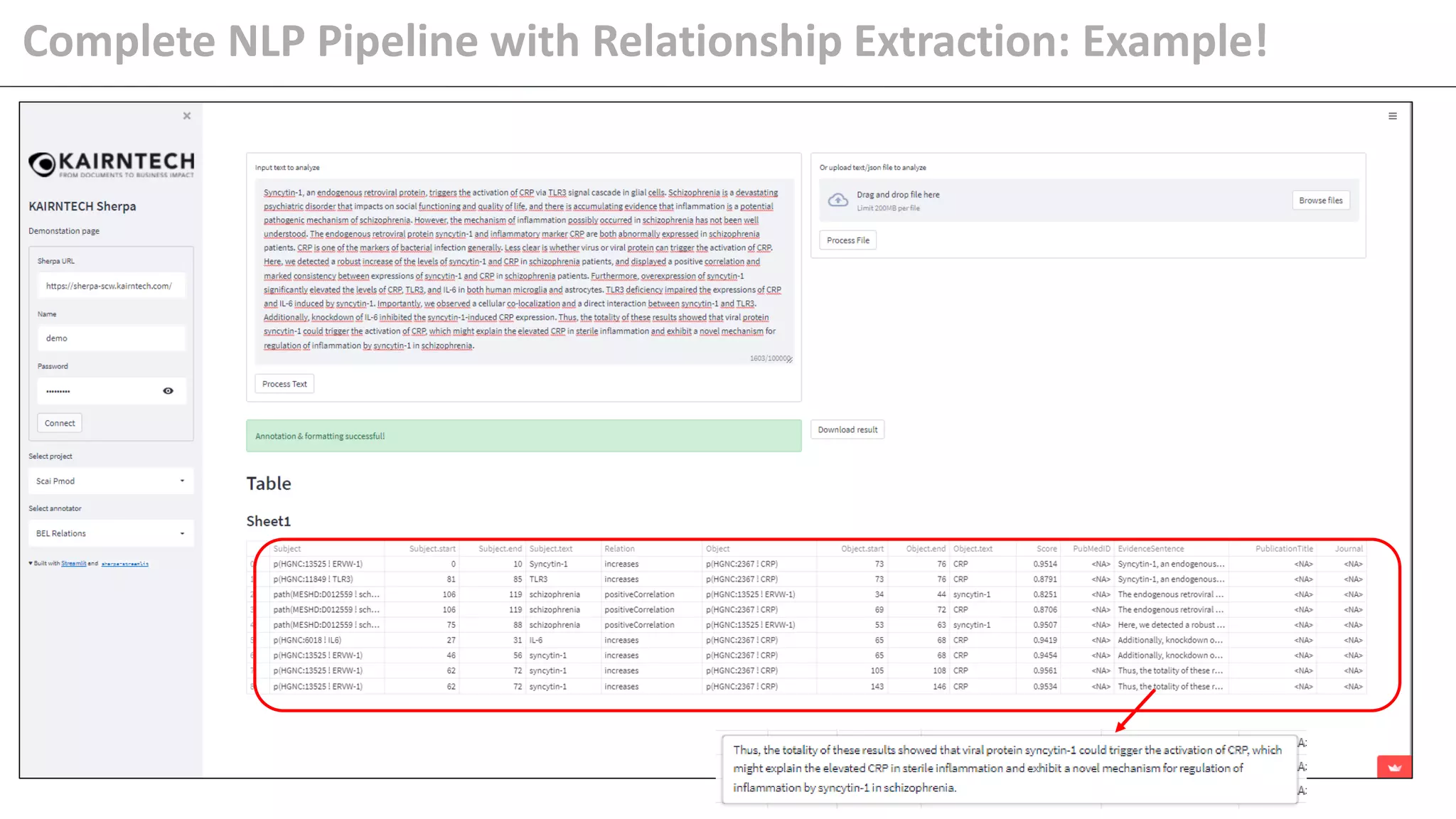
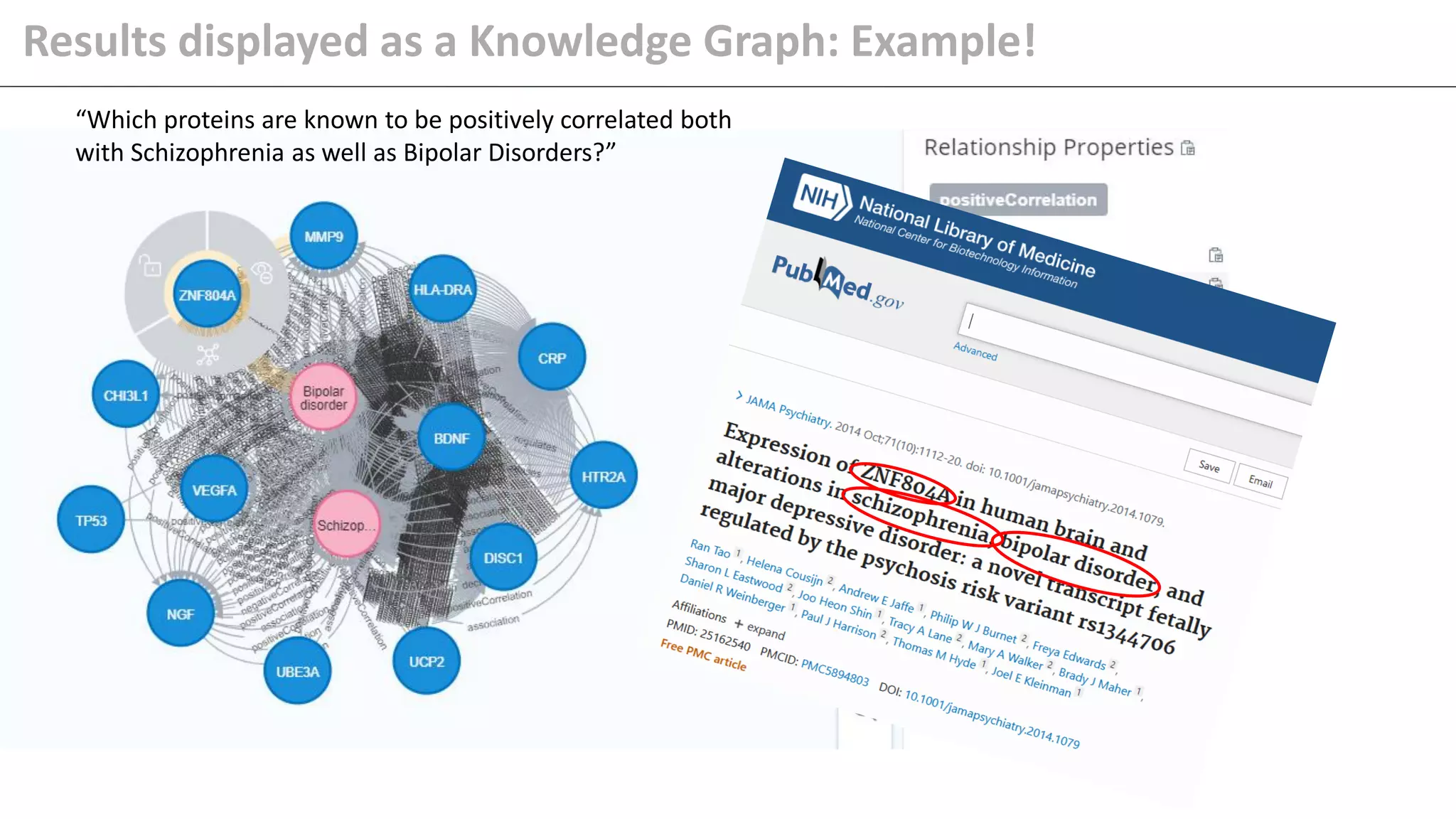
The document details Kairntech's efforts to automate relation extraction from biomedical literature to enhance knowledge graph creation, particularly focusing on psychiatric disorders. It discusses the challenges of keeping knowledge graphs updated and outlines Kairntech's no-code NLP platform designed to streamline the extraction process using machine learning. The results demonstrate an accuracy of 70-80% in relation extraction, with ongoing plans to extend the methodology to broader therapeutic areas and improve processing speeds.