Fossilized WSSV-Like Element DNAV-1_LVa in Genome of Original SPF Shrimp
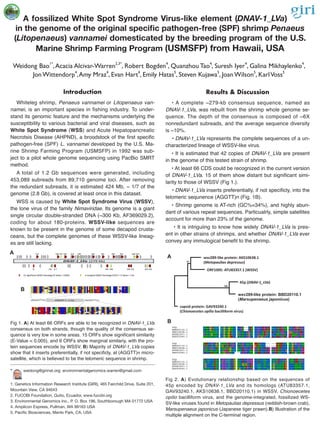
- 1. A fossilized White Spot Syndrome Virus-like element (DNAV-1_LVa) in the genome of the original specific pathogen-free (SPF) shrimp Penaeus (Litopenaeus) vannamei domesticated by the breeding program of the U.S. Marine Shrimp Farming Program (USMSFP) from Hawaii, USA Weidong Bao1* ,Acacia Alcivar-Warren2,3* , Robert Bogden4 , Quanzhou Tao4 , Suresh Iyer4 , Galina Mikhaylenko4 , Jon Wittendorp4 ,Amy Mraz4 , Evan Hart4 , Emily Hatas5 , Steven Kujawa5 , Joan Wilson5 , KarlVoss5 * weidong@girinst.org; environmentalgenomics.warren@gmail.com 1. Genetics Information Research Institute (GIRI), 465 Fairchild Drive, Suite 201, Mountain View, CA 94043 2. FUCOBI Foundation, Quito, Ecuador, www.fucobi.org 3. Environmental Genomics Inc., P. O. Box 196, Southborough MA 01772 USA 4. Amplicon Express, Pullman, WA 99163 USA 5. Pacific Biosciences, Menlo Park, CA, USA Introduction Whiteleg shrimp, Penaeus vannamei or Litopenaeus van- namei, is an important species in fishing industry. To under- stand its genomic feature and the mechanisms underlying the susceptibility to various bacterial and viral diseases, such as White Spot Syndrome (WSS) and Acute Hepatopancreatic Necrotsis Disease (AHPND), a broodstock of the first specific pathogen-free (SPF) L. vannamei developed by the U.S. Ma- rine Shrimp Farming Program (USMSFP) in 1992 was sub- ject to a pilot whole genome sequencing using PacBio SMRT method. A total of 1.2 Gb sequences were generated, including 453,089 subreads from 89,710 genome loci. After removing the redundant subreads, it is estimated 424 Mb, ~ 1/7 of the genome (2.8 Gb), is covered at least once in this dataset. WSS is caused by White Spot Syndrome Virus (WSSV), the lone virus of the family Nimaviridae. Its genome is a giant single circular double-stranded DNA (~300 Kb, AF369029.2), coding for about 180-proteins. WSSV-like sequences are known to be present in the genome of some decapod crusta- ceans, but the complete genomes of these WSSV-like lineag- es are still lacking. Results & Discussion • A complete ~279-kb consensus sequence, named as DNAV-1_LVa, was rebuilt from the shrimp whole genome se- quence. The depth of the consensus is composed of ~6X nonredundant subreads, and the average sequence diversity is ~10%. • DNAV-1_LVa represents the complete sequences of a un- characterized lineage of WSSV-like virus. • It is estimated that 42 copies of DNAV-1_LVa are present in the genome of this tested strain of shrimp. • At least 66 CDS could be recognized in the current version of DNAV-1_LVa. 15 of them show distant but significant simi- larity to those of WSSV (Fig 1.). • DNAV-1_LVa inserts preferentially, if not specificly, into the telomeric sequnence (AGGTT)n (Fig. 1B). • Shrimp genome is AT-rich (GC%=34%), and highly abun- dant of various repeat sequences. Particualrly, simple satellites account for more than 23% of the genome. • It is intriguiing to know how widely DNAV-1_LVa is pres- ent in other strains of shrimps, and whether DNAV-1_LVa ever convey any immulogical benefit to the shrimp. Fig 1. A) At least 66 ORFs are able to be recognized in DNAV-1_LVa consensus on both strands, though the quality of the consensus se- quence is very low in some areas. 15 ORFs show significant similarity (E-Value < 0.005), and 6 ORFs show marginal similariy, with the pro- tein sequences encode by WSSV. B) Majority of DNAV-1_LVa copies show that it inserts preferentially, if not specificly, at (AGGTT)n micro- satellite, which is believed to be the telomeric sequence in shrimp. 9 16 10 6517 18 19 20 24 28 40 45 46 48 49 55 58 59 60 61 64 DNAV-1_LVa (279 Kb) 15 significant WSSV homologs (E-Value < 0.005) 6 marginal WSSV homologs (0.017 < E-Value < 1.9) A B 45p (DNAV-1_LVa) 100 91 wsv289-like protein: BBD20110.1 (Marsupenaeus japonicus) capsid protein: GAV93240.1 (Chionoecetes opilio bacilliform virus) wsv289-like protein: AKS10638.1 (Metopaulias depressus) ORF1005: ATU83357.1 (WSSV) 45p LTNMPDNLYI YVPIFPTSFL MAEQ---IKS VLEIAKDKIK HVVKHREYQS TVIRDCNDEI BBD20110.1 LMNMPDNLHI YVPTLPTSFL MAEQ---IKS ILDIAKDKTK HLIANKKEQS DLIEKHKNEI AKS10638.1 SVNFSSSQYI YVPINPKHIL THMQWLECMS VIEAATRDSA KAVRHFEEAA GGTLEELKKL ATU83357.1 DLNFSPAQNL FVPVNPRHIL TDMQWLNCIS IIETATRDSA IVMQSFQEQA DKTTTQLEEL GAV93240.1 SQNFSSAQHL FVPINPRRLL LNAQWLECVA LIGVGTEHFV KALKKLETRI KDNNEELKTL 45p QKRFVDIWGS YNSQDKIQEA INLDILSVQT RKQDIAI--- ---KNKKLLS SLPVCLDIST BBD20110.1 NEQFMTIWGN SGRLNKLEEA FRQDFLSVLK GKQNIAI--- ---KNKKLFF NLPFLLDLVQ AKS10638.1 LEQWDDIVKQ VTSTESPAVL SLLKLEWMDK EATRIAKLRD EAERSRVALA VQGKVVNIEK ATU83357.1 LSQWNNIVSQ VTDEKSPAYV SSVKLEWLNN EASRIAAIRE NSEKSKIVMG VQGKIVNIDE GAV93240.1 LKLWKQRTNM STGTDDTVF- --LELDWVQQ QAKRLIKLRE HLERTYAAIV VRSIPINIDK 45p YRLHLAIKSM MDIDYYIKVP NFWRLMDIEE MLQFAVSVML ITLQKLVDVG LNTARRYSEV BBD20110.1 YQLHLAIRSV TDIDYYIKVP NFWRMMDIDE MLKFAISLML ITLEKLVNVG LNTAKRFSEI AKS10638.1 YGIIAVSRSL IDVDFYVKLP NSWPSGNWRE LVYSAVSLAS IPLQVNISKG IMAASQATMV ATU83357.1 LGIVAVARSI VDVDFYIKMP NVWASRDWKN LIYYAVNIAA TPLINNISRG IMAASQTSVL GAV93240.1 YSITAIARSL SDVDYYLKVP NPWINMDWHD VMYSALLLTI LPLNLNISRG IILASNASVL Fig 2. A) Evolutionary relationship based on the sequences of 45p encoded by DNAV-1_LVa and its homologs (ATU83357.1, GAV93240.1, AKS10638.1, BBD20110.1) in WSSV, Chionoecetes opilio bacilliform virus, and the genome-integrated, fossilized WS- SV-like viruses found in Metopaulias depressus (reddish-brown crab), Marsupenaeus japonicus (Japanese tiger prawn).B) Illustration of the multiple alignment on the C-terminal region. A B
