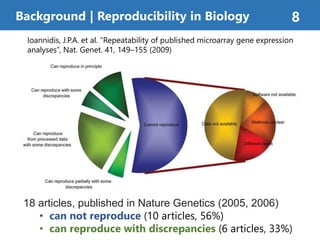
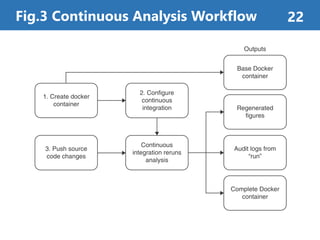
The document summarizes a proposed method called "Continuous Analysis" that aims to improve reproducibility in computational research. Continuous Analysis combines continuous integration (CI) practices with computational research to automatically verify reproducibility. It proposes using Docker containers to capture computational environments and CI tools like Drone to automatically rebuild images and rerun analyses on code changes, flagging any differences in results. An experiment applying Continuous Analysis to two genomic analyses demonstrated it could more easily detect changes between versions.



![Target Problem
Reproducibility of computational research
Proposed Method
Continuous Integration + Computational Research
= Continuous Analysis
Continuous Analysis can automatically
verify the research reproducibility
• Easy to reproduce, review, and cooperate
What is the value of this research ? 4
[GitHub] https://greenelab.github.io/continuous_analysis/](https://image.slidesharecdn.com/20170420journalseminaraoyama-170421064457/85/Reproducibility-of-computational-workflows-is-automated-using-continuous-analysis-4-320.jpg)

![Research reproducibility is crucial for science
But 90% of researchers acknowledged reproducibility crisis[1]
Background | Reproducibility Crisis 6
[1] Baker, M. 1,500 scientists lift the lid on reproducibility. Nature 533, 452–454 (2016).](https://image.slidesharecdn.com/20170420journalseminaraoyama-170421064457/85/Reproducibility-of-computational-workflows-is-automated-using-continuous-analysis-6-320.jpg)


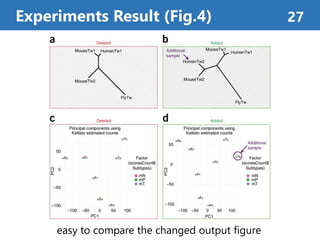
![Survey of Differential Gene Expression Research
• Probe information is necessary for reproduction
• probe, is the oligonucleotides of certain sequences,
is used to measure transcript expression levels
BrainArray Custom CDF [1]
• A popular source of probe set description files
• [Dai, M. et al.] published and maintains
• Version of Custom CDF can verify detailed information of probe set
Authors analyzed the 200 articles, which cited [Dai, M. et al.][1].
Reproducibility on RNA-Analysis 10
[1] Dai, M. et al. Evolving gene/transcript definitions significantly alter the
interpretation of GeneChip data. Nucleic Acids Res. 33, e175 (2005).](https://image.slidesharecdn.com/20170420journalseminaraoyama-170421064457/85/Reproducibility-of-computational-workflows-is-automated-using-continuous-analysis-10-320.jpg)
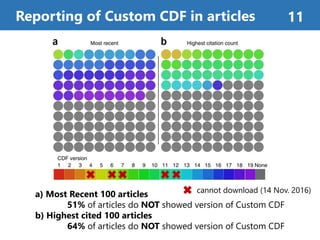

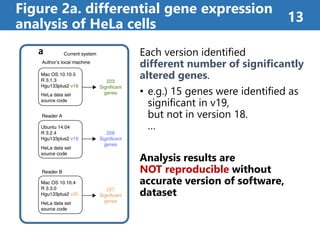
![Figure 2b. container-based approaches 14
Using Docker[1] containers
improves reproducibility
• Docker can create “image”
which contains software env.
• Docker allows users to run the
exact same apps in any env.
Using Docker container enabled
versions to be matched and
produced same result.
[1] https://www.docker.com](https://image.slidesharecdn.com/20170420journalseminaraoyama-170421064457/85/Reproducibility-of-computational-workflows-is-automated-using-continuous-analysis-14-320.jpg)



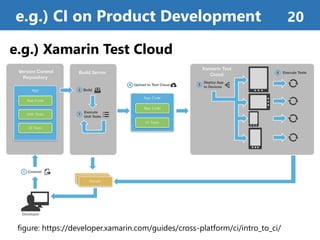
![Continuous Integration (CI)[1]
• is a software engineering practice for fast development
• automatically build, run tests, and make analytics
which triggered by version control system (e.g. git)
About Continuous Integration 18
[1] Grady (1991). Object Oriented Design: With Applications. Benjamin
Cummings. p. 209. ISBN 9780805300918. Retrieved 2014-08-18.
[2] Travis CI, https://travis-ci.org/
e.g.) Travis CI[2] badge](https://image.slidesharecdn.com/20170420journalseminaraoyama-170421064457/85/Reproducibility-of-computational-workflows-is-automated-using-continuous-analysis-18-320.jpg)








![Linux Container
• virtualizes the host resource as containers
• Filesystem, hostname, IPC, PID, Network, User, etc.
• can be used like Virtual Machines
Linux Kernel Features
• Containers are sharing same host kernel
• namespace[1], chroot, cgroup, SELinux, etc.
Container-based Virtualization 30
[1] E. W. Biederman. “Multiple instances of the global Linux namespaces.”,
In Proceedings of the 2006 Ottawa Linux Symposium, 2006.
Machine
Linux Kernel Space
Container
Process
Process
Container
Process
Process](https://image.slidesharecdn.com/20170420journalseminaraoyama-170421064457/85/Reproducibility-of-computational-workflows-is-automated-using-continuous-analysis-30-320.jpg)
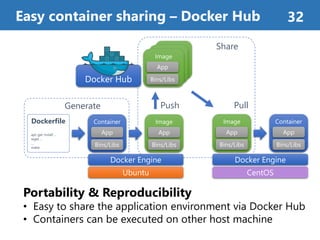
![Docker [1]
• Most popular Linux Container management platform
• Many useful components and services
Linux Container Management Tools 31
[1] Solomon Hykes and others. “What is Docker?” - https://www.docker.com/what-docker
[2] W. Bhimji, S. Canon, D. Jacobsen, L. Gerhardt, M. Mustafa, and J. Porter, “Shifter : Containers for
HPC,” Cray User Group, pp. 1–12, 2016.
[3] “Singularity” - http://singularity.lbl.gov/
[1]
[2] [3]](https://image.slidesharecdn.com/20170420journalseminaraoyama-170421064457/85/Reproducibility-of-computational-workflows-is-automated-using-continuous-analysis-31-320.jpg)
![AUFS (Advanced multi layered unification filesystem) [1]
• Docker default filesystem as AUFS
• Layers can be reused in other container image
• AUFS helps software Reproducibility
Docker - Filesystem 33
[1] Advanced multi layered unification filesystem. http://aufs.sourceforge.net, 2014.
Docker Container (image)
f49eec89601e 129.5 MB ubuntu:16.04 (base image)
366a03547595 39.85 MB
ef122501292c 3.6 MB
e50c89716342 15.4 KB
tag: beta
tag: version-1.0
tag: version-1.0.2
tag: version-1.15aec9aa5462c 1.17 MB
tag: latest0d3cccd04bdb 1.07 MB](https://image.slidesharecdn.com/20170420journalseminaraoyama-170421064457/85/Reproducibility-of-computational-workflows-is-automated-using-continuous-analysis-33-320.jpg)
![Linux Container – Performance [1] 34
[1] W. Felter, A. Ferreira, R. Rajamony, and J. Rubio, “An updated performance comparison of virtual
machines and Linux containers,” IEEE International Symposium on Performance Analysis of Systems and
Software, pp.171-172, 2015. (IBM Research Report, RC25482 (AUS1407-001), 2014.)
0.96 1.00 0.98
0.78
0.83
0.99
0.82
0.98
0.00
0.20
0.40
0.60
0.80
1.00
PXZ [MB/s] Linpack [GFLOPS] Random Access [GUPS]
PerformanceRatio
[basedNative]
Native Docker KVM KVM-tuned](https://image.slidesharecdn.com/20170420journalseminaraoyama-170421064457/85/Reproducibility-of-computational-workflows-is-automated-using-continuous-analysis-34-320.jpg)