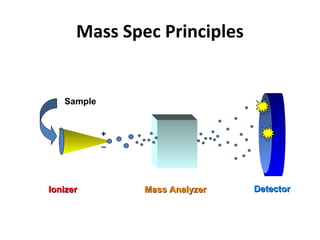
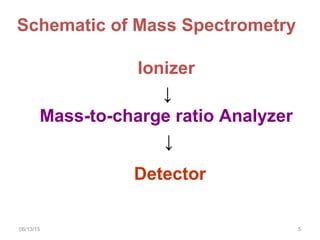
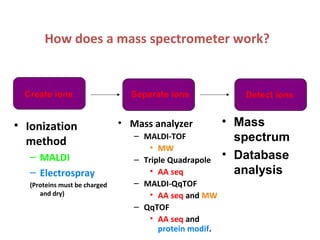
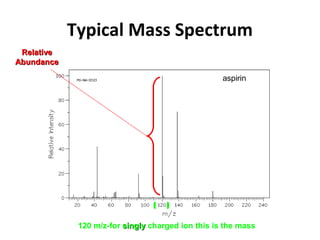
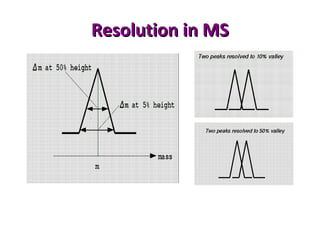
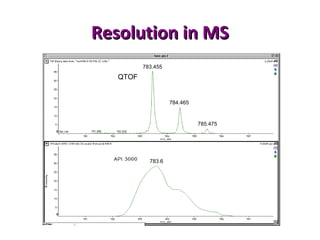
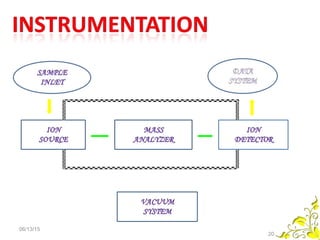
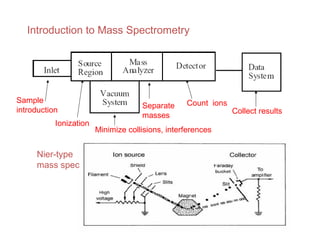
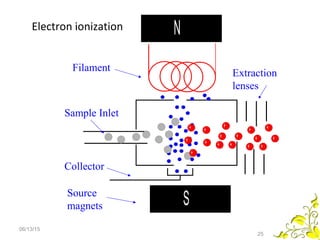
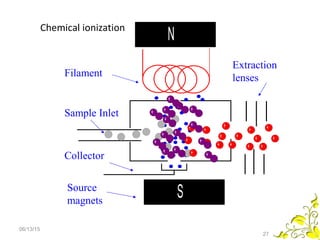
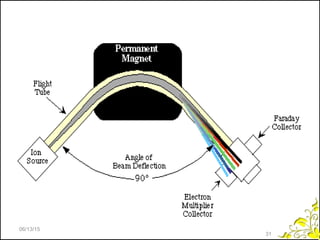
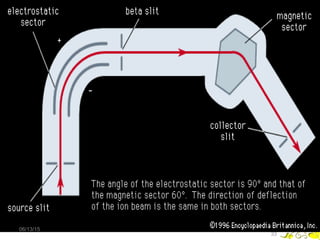
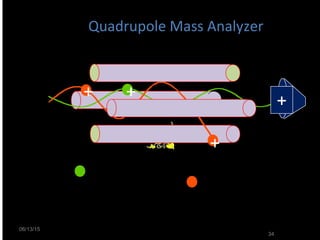
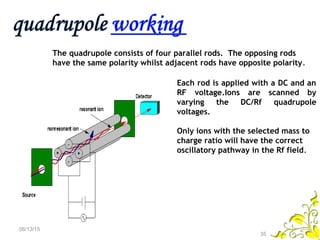
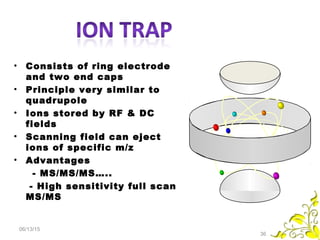
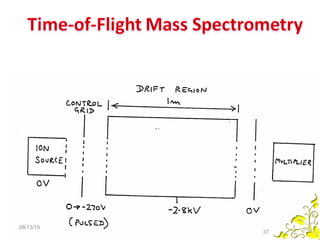
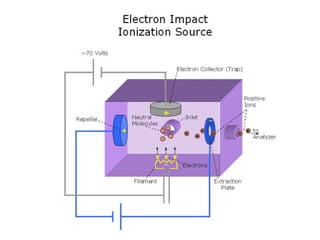
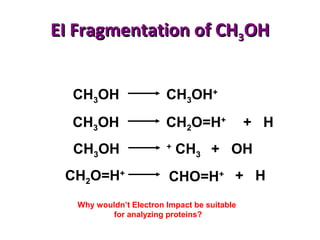
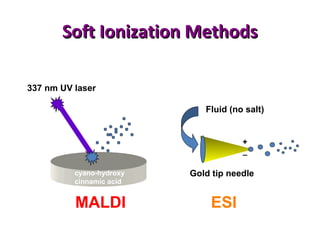
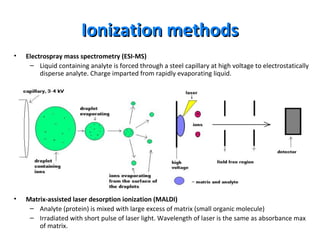
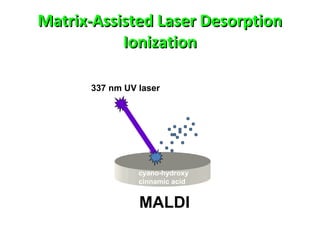
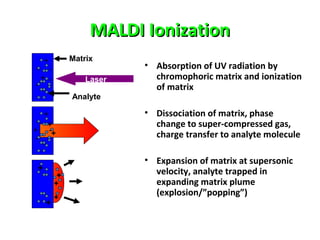
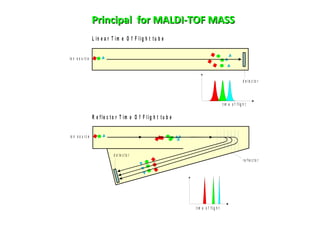
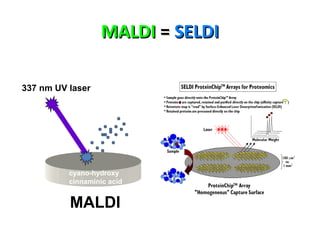
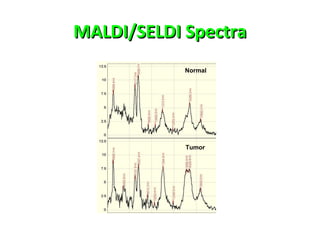
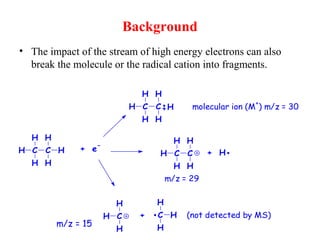
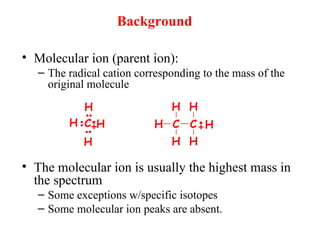
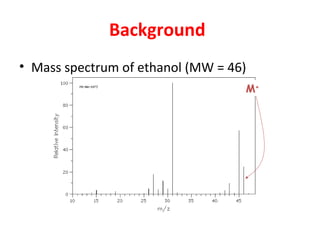
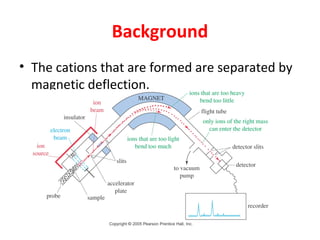
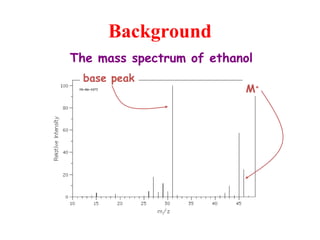
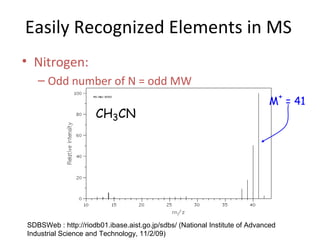
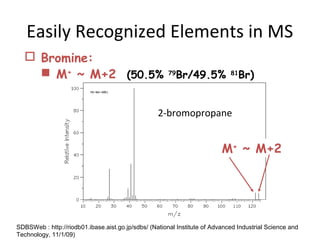
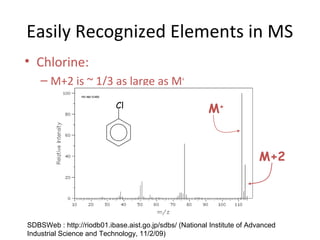
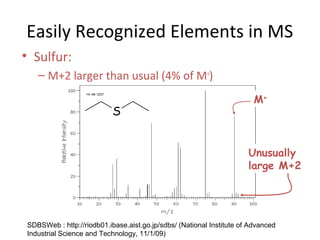
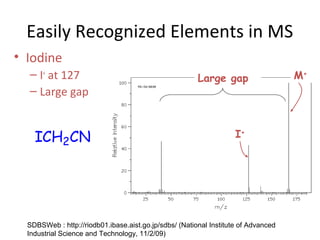
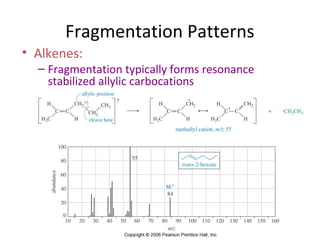
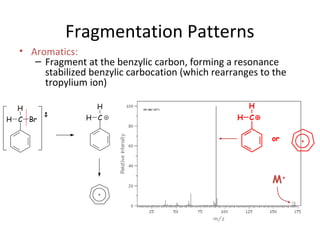
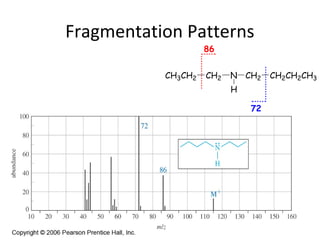
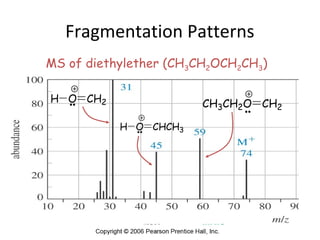
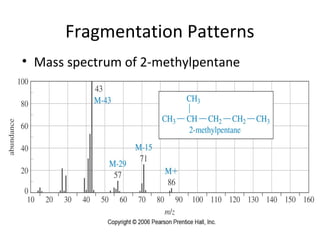
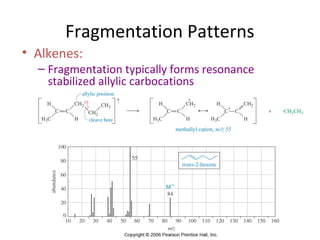
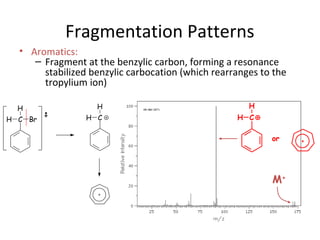
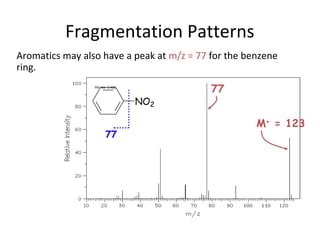
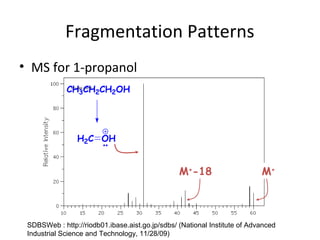
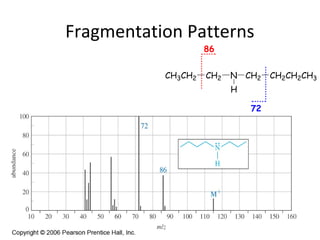
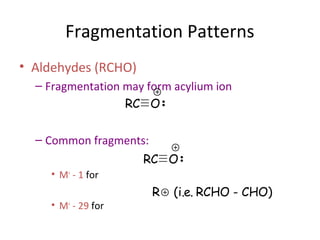
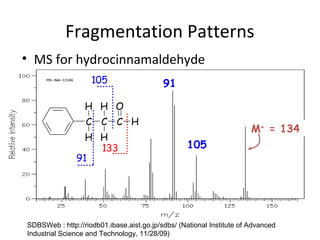
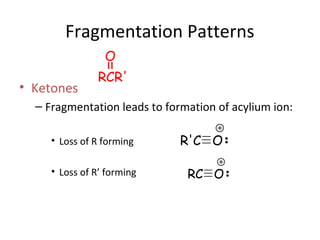
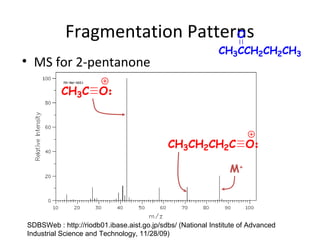
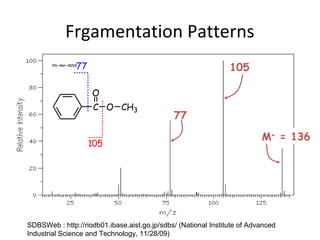
Mass spectrometry is a technique that uses high energy electrons to break molecules into fragments. It then measures the masses of the fragments to reveal information about the molecular structure. Key aspects of mass spectrometry include the ionization source, mass analyzer, and detector. Common ionization methods are electron impact, electrospray, and MALDI, with softer methods like electrospray and MALDI used for larger molecules like proteins. Mass analyzers separate the ions by mass to charge ratio and include quadrupoles, time-of-flight, and magnetic sectors. The detector then counts the ions to produce a mass spectrum.