Ismb2012_poster_cwu
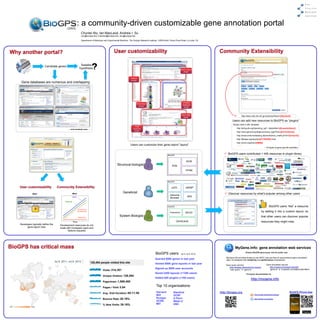
- 1. : a community-driven customizable gene annotation portal Chunlei Wu, Ian MacLeod, Andrew I. Su cwu@scripps.edu, imacleod@scripps.edu, asu@scripps.edu Department of Molecular and Experimental Medicine, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA Candidate genes Testable hypothesis Gene databases are numerous and overlapping http://www.ncbi.nlm.nih.gov/pubmed?term={{Symbol}} Users can add new resources to BioGPS as “plugins” Simply need a URL template: http://string-db.org/newstring_cgi?...&identifier={{EnsemblGene}} … and hundreds more … http://www.genome.jp/dbget-bin/www_bget?hsa:{{EntrezGene}} http://smart.embl-heidelberg.de/smart/show_motifs.pl?ID={{Uniprot}} http://flybase.org/reports/{{FLYBASE}}.html http://omim.org/entry/{{MIM}} Users can customize their gene-report ”layout” …… ~ 30 types of gene-specific identifiers BioGPS BioGPS users contributed > 400 resources in plugin library NCBI Structural biologist PDB PFAM BioGPS eQTL dbSNP Why? Why? Geneticist Discover resources by what’s popular among other users Genome Users MGI Browser Requests BioGPS BioGPS users “like” a resource Community development Expression KEGG by adding it into a custom layout, so Resources System Biologist that other users can discover popular Time GeneCards resources they might miss. Developers typically define the Development resources do not gene-report view scale with increased users and feature-requests MyGene.Info: gene annotation web services (Powers BioGPS gene query, now for public use) BioGPS users: (up to Jul 8, 2012) MyGene.Info provides simple-to-use REST web services to query/retrieve gene annotation Queried 680k genes in last year data. It's designed with simplicity and performance emphasized. Viewed 800k gene-reports in last year Gene query service: Gene annotation service http://mygene.info/query?q=<query> http://mygene.info/gene/<geneid> Signed up 5600 user accounts “user query” “gene id” “gene id” “a specific annotation/identifiers” Saved 2200 layouts (>1300 users) Full query documentation at: Added 400 plugins (>100 users) http://mygene.info Top 10 organizations: Harvard Stanford http://biogps.org BioGPS iPhone App NIH UCSF http://sulab.org/category/biogps Scripps U Penn http://twitter.com/biogps UCSD Wash U MIT UNC