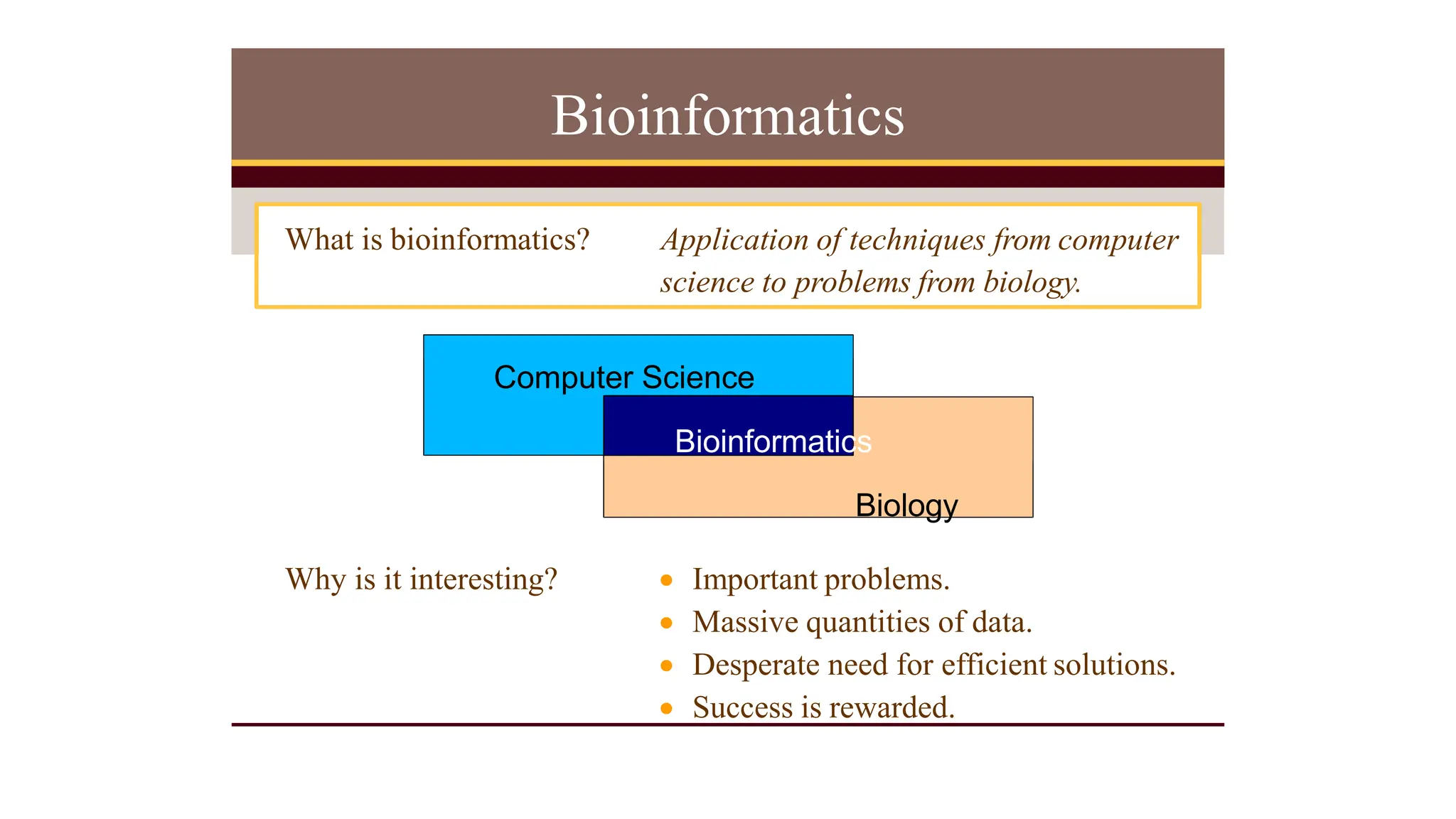
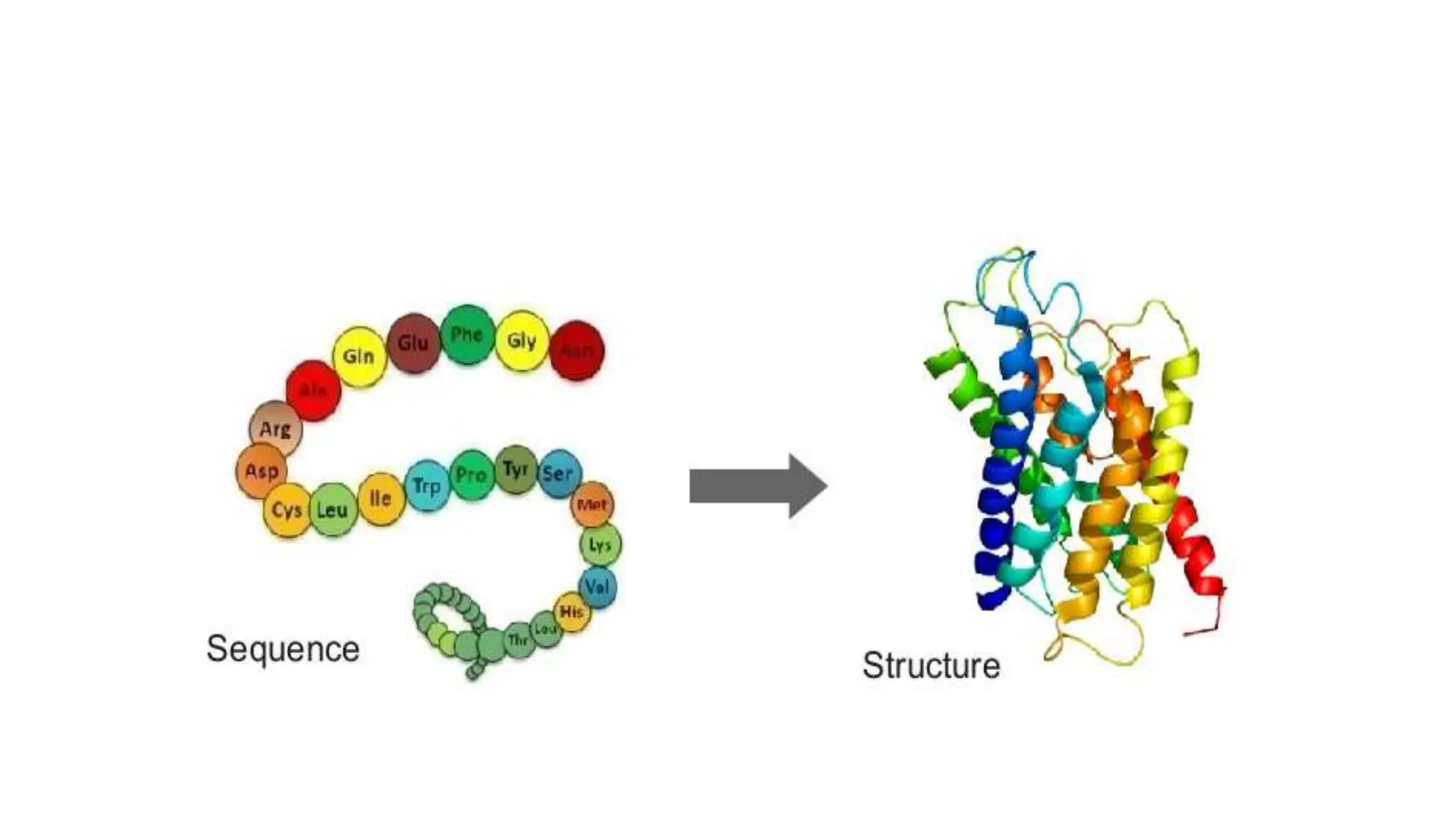
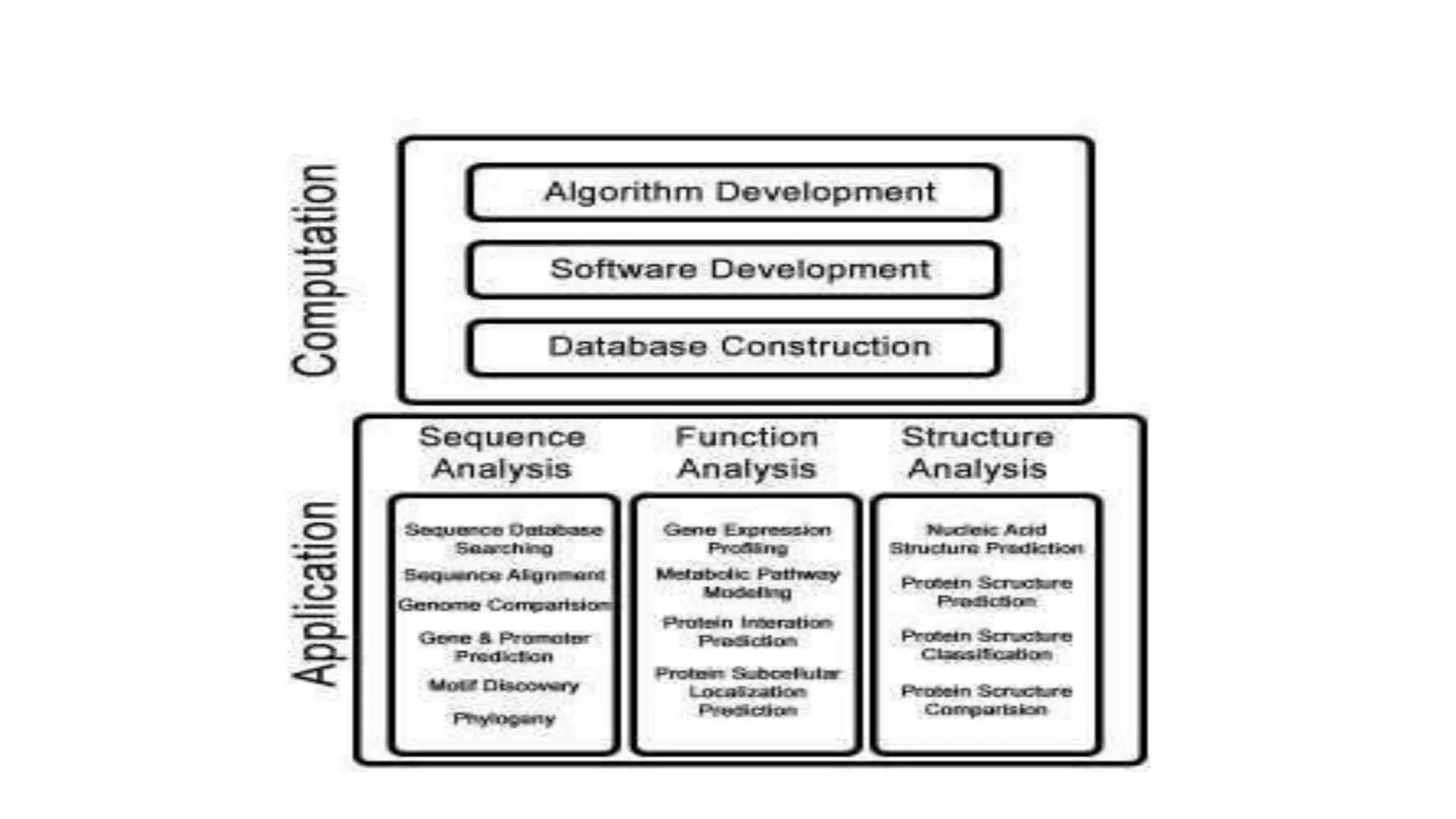
Bioinformatics is the application of computer science and information technology to the field of biology and healthcare. It involves storing, organizing, and analyzing large amounts of biological data, such as DNA sequences and protein structures. Some key goals of bioinformatics include understanding normal biological processes, diseases, and improving drug discovery through computational analysis of various types of biological data, including sequences, structures, and scientific literature.