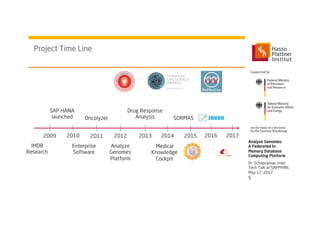
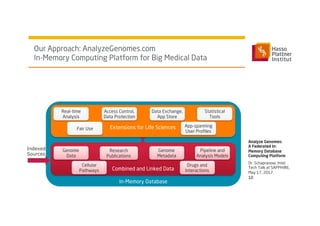
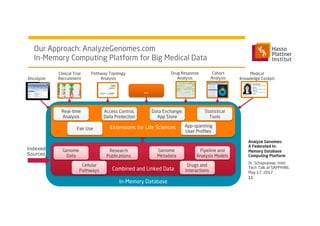
1) Dr. Schapranow presents a federated in-memory database computing platform called AnalyzeGenomes.com to enable real-time analysis of big medical data.
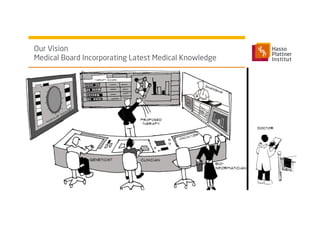
2) The platform aims to incorporate all available patient data, reference latest lab results and medical knowledge, and support interactive analysis to help clinicians make treatment decisions.
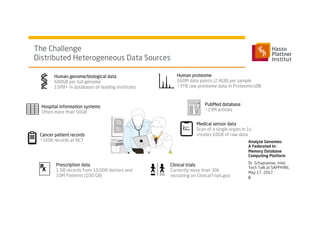
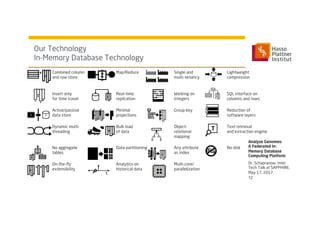
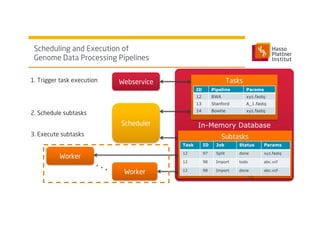
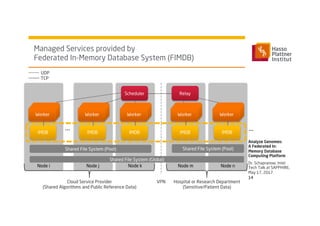
3) It uses a distributed in-memory database across nodes to combine and link heterogeneous medical data sources while addressing challenges of data privacy, locality, and volume.