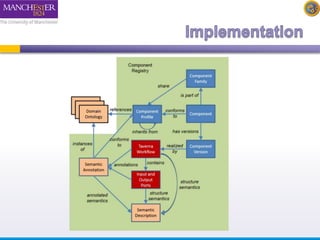
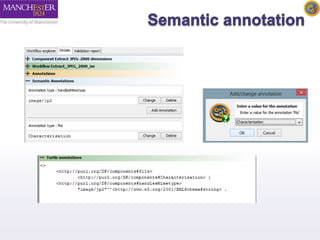
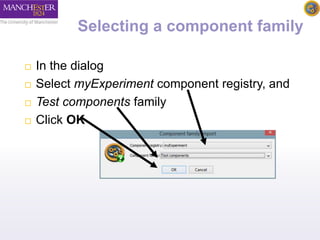
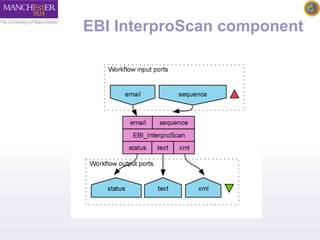
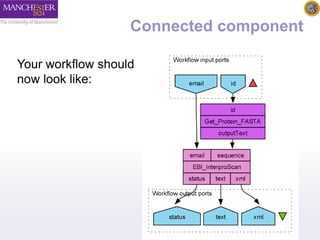
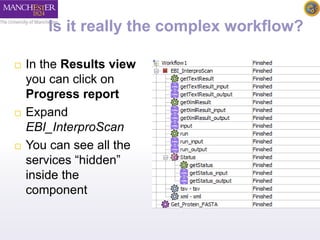
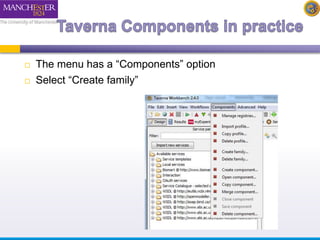
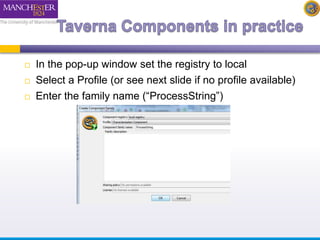
The document introduces Taverna components and their functionalities within workflows. It explains the characteristics of components, how they are created, annotated, and managed in component families, and their integration into workflows. The text outlines steps for creating components, adding them to workflows, and connecting different services efficiently.