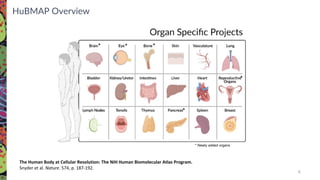
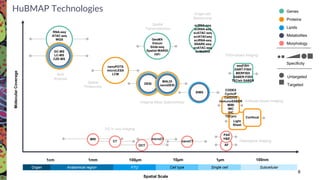
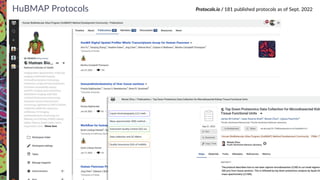
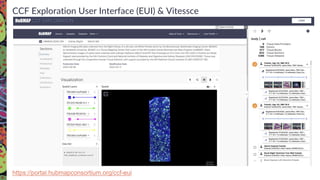
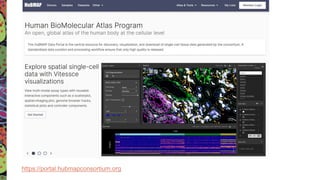
The Human Biomolecular Atlas Program (HuBMAP) aims to create an open global framework for mapping the human body at cellular resolution, focusing on tools and techniques for high-resolution spatial tissue maps. It promotes collaboration with the biomedical research community and supports various pilot projects to study tissue variations throughout the human lifespan. The program also encourages the development of an open data platform for effective data integration and visualization, enhancing the accessibility and usability of the resulting datasets for research purposes.













![Production Phase (2022 - 2026)
Demo and Collaborative Projects
- Integration with MatrisomeDB (Naba)
- Identification of Organotypic Vasculature (Gupta)
- Reverse Engineering Ovarian Organization (Laronda)
- Identification of mitochondria variants (Pei)
- Multi-tissue analysis from single donors (Pryhuber)
Outreach and Collaboration
- Continue Junior Investigators meetings, Jumpstart, Summer Internships, Kaggles…
- Organization joint workshops / meetings (e.g., HPAP)
- Plans for Enhancing Diverse Perspectives (PEDPs)
- Open Working Groups - ASCT+B, Affinity Reagents, Data Visualization [new]](https://image.slidesharecdn.com/dknetwebinarhubmap10142022-221019215258-7041a543/85/dkNET-Webinar-The-Human-BioMolecular-Atlas-Program-HuBMAP-10-14-2022-40-320.jpg)
