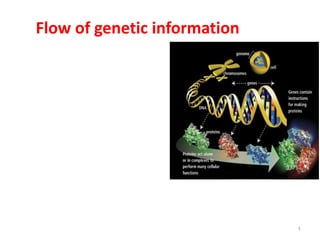
Flow of Genetic Information: DNA, RNA, Proteins
- 1. Flow of genetic information 1
- 2. Macromolecules • Macromolecules: Very large molecules that make most of the structure of the body made of smaller pieces called monomers that can be assembled to form a variety of structures. A large chain of monomers is called a polymer. 2
- 3. Proteins • Monomer: amino acid (20) • Polymer: protein or polypeptide • Structure central carbon atom with hydrogen, amine, carboxyl, & R groups • Uses: structure, emergency energy • Function expression of genetic information • Examples: skin, insulin, enzymes 3
- 4. Nucleic Acids • Monomer: nucleotide (5) • Polymer: nucleic acid • Structure: 5-carbon sugar attached to nitrogen base and phosphate group • Uses: stores genetic code • Examples: DNA and RNA • DNA store genetic information . • RNA process genetic information • Protein express of genetic information 4
- 5. Nucleic acids (DNA and RNA) • Nucleic acids, DNA and RNA are polymers of nucleotides which themselves consist of a nitrogenous base, a pentose sugar and phosphoric acid. Nucleic acids are involved in the storage and processing of genetic information. • Bases: The bases of DNA and RNA are aromatic rings (Fig. 1). Adenine (A) and guanine (G) are purines, bicyclic structures whereas cytosine (C), thymine (T) and uracil (U) are monocyclic pyrimidines. In RNA, the thymine base is replaced by uracil. 5
- 6. • Nucleosides A nucleoside consists of a base covalently bonded to the 1'-position of a pentose sugar molecule. In RNA the sugar is ribose and the compounds are ribonucleosides, or nucleosides, whereas in DNA it is 2'-deoxyribose and the nucleosides are named 2'-deoxyribonucleosides, or just deoxynucleosides. Base + sugar = nucleoside. 6
- 7. Nucleotides: • Nucleotides are nucleosides with one or more phosphate groups covalently bound to the 3'-, 5'- or (in ribonucleotides) the 2'-position. If the sugar is deoxyribose, then the compounds are termed deoxynucleotides (Fig. 3) In the case of the 5'-position, up to three phosphates may be attached, to form, for example, adenosine 5'- triphosphate, or deoxyguanosine 5'-triphosphate, commonly abbreviated to ATP and dGTP respectively.In the same way, we have dCTP, UTP and dTTP (equivalent to TTP). 5'-Mono and -diphosphates are abbreviated as, for example, AMP and dGDP. Nucleoside 5'- triphosphates (NTPs), or deoxynucleoside 5'-triphosphates (dNTPs) are the building blocks of the polymeric nucleic acids..The repeat unit of a DNA or RNA chain is hence a nucleotide. Base + sugar +phosphate = nucleotide 7
- 8. Properties of nucleic acids • Stability of nucleic acids Although it might seem obvious that DNA double strands and RNA structures are stabilized by hydrogen bonding, this is not the case. H-bonds determine the specificity of the base pairing, but the stability of a nucleic acid helix is the result of hydrophobic and dipole–dipole interactions between the stacked base pairs. • Effect of acid: Highly acidic conditions may hydrolyze nucleic acids to their components: bases, sugar and phosphate. Moderate acid causes the hydrolysis of the purine base glycosylic bonds to yield a purinic acid. More complex chemistry has been developed to remove particular bases, and is the basis of chemical DNA sequencing. Effect of alkali: High pH denatures DNA and RNA by altering the tautomeric state of the bases and disrupting specific hydrogen bonding. RNA is also susceptible to hydrolysis at high pH, by participation of the 2'-OH in intramolecular cleavage of the phosphodiester backbone. • Chemical denaturation : Some chemicals, such as urea and formamide, can denature DNA and RNA at neutral pH by disrupting the hydrophobic forces between the stacked bases. 8
- 9. Properties of nucleic acids • Viscosity: DNA is very long and thin, and DNA solutions have a high viscosity. Long DNA molecules are susceptible to cleavage by shearing in solution, this process can be used to generate DNA of a specific average length. • UV absorption The aromatic bases of nucleic acids absorb light with a max of 260 nm. • Hypochromicity: The extinction coefficient of nucleic acid bases depends on their environment.The absorbance of isolated nucleotides is greater than that of RNA and single stranded DNA, which is in turn greater than that of double-stranded DNA. Double-stranded DNA is hypochromic with respect to single-stranded DNA. • Quantitation of nucleic acids:The absorbance at 260 nm is used to determine the concentration of nucleic acids. At a concentration of 1 mg ml and 1 cm path length, double-stranded DNA has A260 = 20. RNA and single-stranded DNA have A260 = 25. The values for RNA and single- stranded DNA depend on base composition and secondary structure. • Purity of DNA The A260/A280 ratio of a double-stranded DNA sample can be used to assess its purity. For pure DNA, the value is 1.8. Values above 1.8 suggest RNA contamination and those below 1.8 suggest protein contamination. 9
- 10. Properties of nucleic acids • Thermal denaturation: Increased temperature can bring about the denaturation of DNA and RNA. RNA denatures gradually on heating, but double-stranded DNA ‘melts’ cooperatively to give single strands at a defined temperature, Tm, which is a function of the G+C content of the DNA. Denaturation may be detected by the change in A260. • Renaturation DNA: renatures on cooling but will only form fully double- stranded native DNA if the cooling is sufficiently slow to allow the complementary strands to anneal. • Intercalators Intercalators, such as ethidium bromide, bind to DNA by inserting themselves between the base pairs (intercalation), resulting in the local untwisting of the DNA helix. If the DNA is closed-circular, then there will be a corresponding increase in writhe. • Closed-circular DNA: Many DNA molecules in cells consist of closed- circular double-stranded molecules, for example bacterial plasmids and chromosomes and many viral DNA molecules. This means that the two complementary single strands are each joined into circles, 5' to 3', and are twisted around one another by the helical path of the DNA. The molecule has no free ends, and the two single strands are linked together a number of times corresponding to the number of double-helical turns in the molecule. This number is known as the linking number (Lk). 10
- 11. Properties of nucleic acids • Supercoiling: Supercoiling is the coiling of the DNA axis upon itself, caused by a change in the linking number from Lk°, the value for a relaxed closed circle. Most natural DNA is negatively supercoiled, that is the DNA is deformed in the direction of unwinding of the double helix. • Supercoiling is biologically important for two reasons. First, a supercoiled DNA molecule has a more • compact shape than does its relaxed counterpart. Second, supercoiling may hinder or favor the capacity of the double helix to unwind and thereby affects the interactions between DNA and other molecules. 11
- 12. Chromatin structure • Chromatin The total length of DNA in a eukaryotic cell depends on the species, but it can be thousands of times as much as in a prokaryotic genome, and is made up of a number of discrete bodies called chromosomes (46 in humans). The DNA in each chromosome is believed to be a single linear molecule, which can be up to several centimeters long. All this DNA must be packaged into the nucleus, a space of approximately the same volume as a bacterial cell; in fact, in their most highly condensed forms, the chromosomes have an enormously high DNA concentration .This feat of packing is accomplished by the formation of a highly organized complex of DNA and protein, known as chromatin, a nucleoprotein complex. More than 50% of the mass of chromatin is protein. • Histones The major protein components of chromatin are the histones; small, basic (positively charged) proteins which bind tightly to DNA. There are four families of core histone, H2A, H2B, H3, H4, and a further family, H1, which has some different properties, and a distinct role. Individual species have a number of variants of the different histone proteins. • Nucleosomes The nucleosome core is the basic unit of chromosome structure, consisting of a protein octamer containing two each of the core histones, with 146 bp of DNA . 12
- 13. Chromatin structure • The role of H1: A single molecule of H1 stabilizes the DNA at the point at which it enters and leaves the nucleosome core, and organizes the DNA between nucleosomes. • Higher order structure The organization of chromatin at the highest level seems rather similar to that of prokaryotic DNA. Electron micrographs of chromosomes which have been stripped of their histone proteins show a looped domain structure.. Even the size of the loops is approximately the same, up to around 100 kb of DNA, although there are many more loops in a eukaryotic chromosome. The loops are constrained by interaction with a protein complex known as the nuclear matrix. The DNA in the loops is in13
- 14. Eukaryotic chromosome structure • The mitotic chromosome: The familiar picture of a chromosome (Fig. 1) is actually that of its most highly condensed state at mitosis. As the daughter chromosomes are pulled apart by the mitotic spindle at cell division, the fragile centimeters-long chromosomal DNA would certainly be shared by the forces generated, were it not in this highly compact state. The structure in Fig. 1 actually illustrates two identical sister chromatids, the products of replication of a single chromosome, joined at their centromeres. The tips of the chromosomes are the telomeres, which are also the ends of the DNA molecule; the DNA maps in a linear fashion along the length of the chromosome, albeit in a very convoluted path. The structure of a section of a mitotic chromatid is shown in Fig. 1. The chromosomal loops fan out from a central scaffold or nuclear matrix region consisting of protein. The centromere The centromere is the region where the two chromatids are joined and is also the site of attachment to the mitotic spindle, which pulls apart the sister chromatids at anaphase. Centromeres are characterized by specific short DNA sequences although, in mammalian cells, there may be an involvement of satellite DNA. 14
- 15. Eukaryotic chromosome structure • Telomeres The ends of the linear chromosomal DNA are protected from degradation and gradual shortening by the telomeres, which are short repeating sequences synthesized by a specific enzyme, telomerase, independently of normal DNA replication. • • Histone variants and modification The control of the degree of condensation of chromatin operates, at least in part, through the chemical modification of histone proteins, which changes their charge during the cell cycle, or through the use of histone variants in particular cell types or during development. • Function of histone Coil the DNA into a smaller volume for fitting in the nucleus of the cell in form of a nucleosome Regulation of gene expression DNA damage repair DNA replication and recombination heritable epigenetic regulation 15
- 16. 16
- 17. Gene, Genome and Proteome • Gene: A specific sequence of nucleotides in DNA (in cells and certain viruses), or in RNA (in other viruses), that encodes a given polypeptide or an RNA molecule with a biosynthetic or control function. • Genome: total DNA complement or the sum of all the genes of an organism. It may comprise a single chromosome or be spread across multiple chromosomes. In eukaryotes, it can be subdivided into nuclear, mitochondrial and chloroplast genomes. • Transcriptome: sum of all the RNA transcripts (mRNA) of an organism, produced from the genome at any given time. • Proteome: sum of all the proteins of an organism. Is used both to describe the total set of proteins encoded in the genome of a cell and the various subsets that are expressed from the transcriptome at any one time. The proteome includes all the various products encoded by a single gene that may result from multiple transcripts • Metabolome: sum of all small molecules – amino acids, nucleotides, sugars etc. – and all the intermediates that exist within a cell during their synthesis and degradation. 17
- 18. The flow of genetic information 18 DNA makes RNA makes protein =protein synthesis DNA makes DNA = DNA replication DNA makes RNA = transcription RNA makes protein= Translation RNA makes DNA= Reverse transcription RNA makes RNA= RNA replication biological information moves from the genome to ecologies (DNA to RNA to protein to biomodules or networks to cells to organs to individuals to populations of individuals to ecologies)
- 19. Genome of Prokaryotes • Their genome is a single, circular DNA molecule. • Their genome size is in the order of a few million base pairs [0.6–8]. • Their gene density (the number of genes per base pairs in the genome) is approximately one gene per 1,000 base pairs. • Their genome is containing few useless parts (70 percent is coding for proteins). • Their genes are transcribed (copied into messenger RNA) right after a control region called a promoter. • Their Messenger RNA (mRNA) is collinear with the genome sequence. In other words, genes are in a single piece, not interrupted by noncoding DNA segment (called introns). mRNA doesn’t need processing (splicing, polyadenylation and capping ). • Gene often exhibit one mRNA (and protein) form. 19
- 20. Genome of Eukaryotes • Their genome consists of multiple linear pieces of DNA called chromosomes (up to a hundred million base pairs long). • Their genome size (10–670,000 million base pairs), especially for animals, plants is much bigger than in prokaryotes. • Their gene density is much lower than that for prokaryotes (one human gene per 100,000 base pairs). • Their genome is containing many useless parts (less than 5 percent of the human genome code for proteins). • Their genes are transcribed right after a control region called a promoter, but sequence elements located far away can have a strong influence on this process. • Gene sequences are not collinear with the final messenger RNA (mRNA) and protein sequences. Only small bits (the exons) are retained in the mature mRNA that encodes the final product. In other words, mRNA need processing (splicing, polyadenylation and capping ) • Genes often exhibit more than one mRNA (and protein) form. 20
