Professor Yoon Sup Choi discusses digital health and the future of healthcare centered around changes in the pharmaceutical industry. He notes that three key steps in implementing digital medicine are: 1) measuring data through devices like smartphones, wearables, and genetic analysis; 2) collecting the data; and 3) gaining insights from the data using artificial intelligence. Choi also provides an overview of the digital health industry landscape and increasing investment in digital health startups from pharmaceutical companies and other investors.































































![Fig 1. What can consumer wearables do? Heart rate can be measured with an oximeter built into a ring [3], muscle activity with an electromyographi
sensor embedded into clothing [4], stress with an electodermal sensor incorporated into a wristband [5], and physical activity or sleep patterns via an
accelerometer in a watch [6,7]. In addition, a female’s most fertile period can be identified with detailed body temperature tracking [8], while levels of me
attention can be monitored with a small number of non-gelled electroencephalogram (EEG) electrodes [9]. Levels of social interaction (also known to a
PLOS Medicine 2016](https://image.slidesharecdn.com/merck1hrv16digitalmedicine180115-180115023755/85/slide-65-320.jpg)







































































































































![DeepFace: Closing the Gap to Human-Level
Performance in FaceVerification
Taigman,Y. et al. (2014). DeepFace: Closing the Gap to Human-Level Performance in FaceVerification, CVPR’14.
Figure 2. Outline of the DeepFace architecture. A front-end of a single convolution-pooling-convolution filtering on the rectified input, followed by three
locally-connected layers and two fully-connected layers. Colors illustrate feature maps produced at each layer. The net includes more than 120 million
parameters, where more than 95% come from the local and fully connected layers.
very few parameters. These layers merely expand the input
into a set of simple local features.
The subsequent layers (L4, L5 and L6) are instead lo-
cally connected [13, 16], like a convolutional layer they ap-
ply a filter bank, but every location in the feature map learns
a different set of filters. Since different regions of an aligned
image have different local statistics, the spatial stationarity
The goal of training is to maximize the probability of
the correct class (face id). We achieve this by minimiz-
ing the cross-entropy loss for each training sample. If k
is the index of the true label for a given input, the loss is:
L = log pk. The loss is minimized over the parameters
by computing the gradient of L w.r.t. the parameters and
Human: 95% vs. DeepFace in Facebook: 97.35%
Recognition Accuracy for Labeled Faces in the Wild (LFW) dataset (13,233 images, 5,749 people)](https://image.slidesharecdn.com/merck1hrv16digitalmedicine180115-180115023755/85/slide-204-320.jpg)
![FaceNet:A Unified Embedding for Face
Recognition and Clustering
Schroff, F. et al. (2015). FaceNet:A Unified Embedding for Face Recognition and Clustering
Human: 95% vs. FaceNet of Google: 99.63%
Recognition Accuracy for Labeled Faces in the Wild (LFW) dataset (13,233 images, 5,749 people)
False accept
False reject
s. This shows all pairs of images that were
on LFW. Only eight of the 13 errors shown
he other four are mislabeled in LFW.
on Youtube Faces DB
ge similarity of all pairs of the first one
our face detector detects in each video.
False accept
False reject
Figure 6. LFW errors. This shows all pairs of images that were
incorrectly classified on LFW. Only eight of the 13 errors shown
here are actual errors the other four are mislabeled in LFW.
5.7. Performance on Youtube Faces DB
We use the average similarity of all pairs of the first one
hundred frames that our face detector detects in each video.
This gives us a classification accuracy of 95.12%±0.39.
Using the first one thousand frames results in 95.18%.
Compared to [17] 91.4% who also evaluate one hundred
frames per video we reduce the error rate by almost half.
DeepId2+ [15] achieved 93.2% and our method reduces this
error by 30%, comparable to our improvement on LFW.
5.8. Face Clustering
Our compact embedding lends itself to be used in order
to cluster a users personal photos into groups of people with
the same identity. The constraints in assignment imposed
by clustering faces, compared to the pure verification task,
lead to truly amazing results. Figure 7 shows one cluster in
a users personal photo collection, generated using agglom-
erative clustering. It is a clear showcase of the incredible
invariance to occlusion, lighting, pose and even age.
Figure 7. Face Clustering. Shown is an exemplar cluster for one
user. All these images in the users personal photo collection were
clustered together.
6. Summary
We provide a method to directly learn an embedding into
an Euclidean space for face verification. This sets it apart
from other methods [15, 17] who use the CNN bottleneck
layer, or require additional post-processing such as concate-
nation of multiple models and PCA, as well as SVM clas-
sification. Our end-to-end training both simplifies the setup
and shows that directly optimizing a loss relevant to the task
at hand improves performance.
Another strength of our model is that it only requires
False accept
False reject
Figure 6. LFW errors. This shows all pairs of images that were
incorrectly classified on LFW. Only eight of the 13 errors shown
here are actual errors the other four are mislabeled in LFW.
5.7. Performance on Youtube Faces DB
We use the average similarity of all pairs of the first one
hundred frames that our face detector detects in each video.
This gives us a classification accuracy of 95.12%±0.39.
Using the first one thousand frames results in 95.18%.
Compared to [17] 91.4% who also evaluate one hundred
frames per video we reduce the error rate by almost half.
DeepId2+ [15] achieved 93.2% and our method reduces this
error by 30%, comparable to our improvement on LFW.
5.8. Face Clustering
Our compact embedding lends itself to be used in order
to cluster a users personal photos into groups of people with
the same identity. The constraints in assignment imposed
by clustering faces, compared to the pure verification task,
Figure 7. Face Clustering. Shown is an exemplar cluster for one
user. All these images in the users personal photo collection were
clustered together.
6. Summary
We provide a method to directly learn an embedding into
an Euclidean space for face verification. This sets it apart
from other methods [15, 17] who use the CNN bottleneck
layer, or require additional post-processing such as concate-
nation of multiple models and PCA, as well as SVM clas-](https://image.slidesharecdn.com/merck1hrv16digitalmedicine180115-180115023755/85/slide-205-320.jpg)
![Targeting Ultimate Accuracy: Face
Recognition via Deep Embedding
Jingtuo Liu (2015) Targeting Ultimate Accuracy: Face Recognition via Deep Embedding
Human: 95% vs.Baidu: 99.77%
Recognition Accuracy for Labeled Faces in the Wild (LFW) dataset (13,233 images, 5,749 people)
3
Although several algorithms have achieved nearly perfect
accuracy in the 6000-pair verification task, a more practical
can achieve 95.8% identification rate, relatively reducing the
error rate by about 77%.
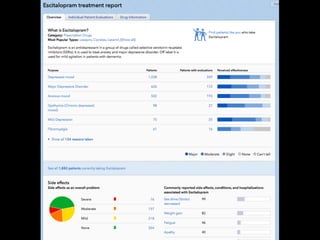
TABLE 3. COMPARISONS WITH OTHER METHODS ON SEVERAL EVALUATION TASKS
Score = -0.060 (pair #113) Score = -0.022 (pair #202) Score = -0.034 (pair #656)
Score = -0.031 (pair #1230) Score = -0.073 (pair #1862) Score = -0.091(pair #2499)
Score = -0.024 (pair #2551) Score = -0.036 (pair #2552) Score = -0.089 (pair #2610)
Method
Performance on tasks
Pair-wise
Accuracy(%)
Rank-1(%)
DIR(%) @
FAR =1%
Verification(%
)@ FAR=0.1%
Open-set
Identification(%
)@ Rank =
1,FAR = 0.1%
IDL Ensemble
Model
99.77 98.03 95.8 99.41 92.09
IDL Single Model 99.68 97.60 94.12 99.11 89.08
FaceNet[12] 99.63 NA NA NA NA
DeepID3[9] 99.53 96.00 81.40 NA NA
Face++[2] 99.50 NA NA NA NA
Facebook[15] 98.37 82.5 61.9 NA NA
Learning from
Scratch[4]
97.73 NA NA 80.26 28.90
HighDimLBP[10] 95.17 NA NA
41.66(reported
in [4])
18.07(reported
in [4])
• 6,000쌍의 얼굴 사진 중에 바이두의 인공지능은 불과 14쌍만을 잘못 판단
• 알고 보니 이 14쌍 중의 5쌍의 사진은 오히려 정답에 오류가 있었고,
실제로는 인공지능이 정확 (red box)](https://image.slidesharecdn.com/merck1hrv16digitalmedicine180115-180115023755/85/slide-206-320.jpg)
















![AtomNet: A Deep Convolutional Neural Network for
Bioactivity Prediction in Structure-based Drug
Discovery
Izhar Wallach
Atomwise, Inc.
izhar@atomwise.com
Michael Dzamba
Atomwise, Inc.
misko@atomwise.com
Abraham Heifets
Atomwise, Inc.
abe@atomwise.com
Abstract
Deep convolutional neural networks comprise a subclass of deep neural networks
(DNN) with a constrained architecture that leverages the spatial and temporal
structure of the domain they model. Convolutional networks achieve the best pre-
dictive performance in areas such as speech and image recognition by hierarchi-
cally composing simple local features into complex models. Although DNNs have
been used in drug discovery for QSAR and ligand-based bioactivity predictions,
none of these models have benefited from this powerful convolutional architec-
ture. This paper introduces AtomNet, the first structure-based, deep convolutional
neural network designed to predict the bioactivity of small molecules for drug dis-
covery applications. We demonstrate how to apply the convolutional concepts of
feature locality and hierarchical composition to the modeling of bioactivity and
chemical interactions. In further contrast to existing DNN techniques, we show
that AtomNet’s application of local convolutional filters to structural target infor-
mation successfully predicts new active molecules for targets with no previously
known modulators. Finally, we show that AtomNet outperforms previous docking
approaches on a diverse set of benchmarks by a large margin, achieving an AUC
greater than 0.9 on 57.8% of the targets in the DUDE benchmark.
1 Introduction
Fundamentally, biological systems operate through the physical interaction of molecules. The ability
to determine when molecular binding occurs is therefore critical for the discovery of new medicines
and for furthering of our understanding of biology. Unfortunately, despite thirty years of compu-
tational efforts, computer tools remain too inaccurate for routine binding prediction, and physical
experiments remain the state of the art for binding determination. The ability to accurately pre-
dict molecular binding would reduce the time-to-discovery of new treatments, help eliminate toxic
molecules early in development, and guide medicinal chemistry efforts [1, 2].
In this paper, we introduce a new predictive architecture, AtomNet, to help address these challenges.
AtomNet is novel in two regards: AtomNet is the first deep convolutional neural network for molec-
ular binding affinity prediction. It is also the first deep learning system that incorporates structural
information about the target to make its predictions.
Deep convolutional neural networks (DCNN) are currently the best performing predictive models
for speech and vision [3, 4, 5, 6]. DCNN is a class of deep neural network that constrains its model
architecture to leverage the spatial and temporal structure of its domain. For example, a low-level
image feature, such as an edge, can be described within a small spatially-proximate patch of pixels.
Such a feature detector can share evidence across the entire receptive field by “tying the weights”
of the detector neurons, as the recognition of the edge does not depend on where it is found within
1
arXiv:1510.02855v1[cs.LG]10Oct2015](https://image.slidesharecdn.com/merck1hrv16digitalmedicine180115-180115023755/85/slide-223-320.jpg)
![AtomNet: A Deep Convolutional Neural Network for
Bioactivity Prediction in Structure-based Drug
Discovery
Izhar Wallach
Atomwise, Inc.
izhar@atomwise.com
Michael Dzamba
Atomwise, Inc.
misko@atomwise.com
Abraham Heifets
Atomwise, Inc.
abe@atomwise.com
Abstract
Deep convolutional neural networks comprise a subclass of deep neural networks
(DNN) with a constrained architecture that leverages the spatial and temporal
structure of the domain they model. Convolutional networks achieve the best pre-
dictive performance in areas such as speech and image recognition by hierarchi-
cally composing simple local features into complex models. Although DNNs have
been used in drug discovery for QSAR and ligand-based bioactivity predictions,
none of these models have benefited from this powerful convolutional architec-
ture. This paper introduces AtomNet, the first structure-based, deep convolutional
neural network designed to predict the bioactivity of small molecules for drug dis-
covery applications. We demonstrate how to apply the convolutional concepts of
feature locality and hierarchical composition to the modeling of bioactivity and
chemical interactions. In further contrast to existing DNN techniques, we show
that AtomNet’s application of local convolutional filters to structural target infor-
mation successfully predicts new active molecules for targets with no previously
known modulators. Finally, we show that AtomNet outperforms previous docking
approaches on a diverse set of benchmarks by a large margin, achieving an AUC
greater than 0.9 on 57.8% of the targets in the DUDE benchmark.
1 Introduction
Fundamentally, biological systems operate through the physical interaction of molecules. The ability
to determine when molecular binding occurs is therefore critical for the discovery of new medicines
and for furthering of our understanding of biology. Unfortunately, despite thirty years of compu-
tational efforts, computer tools remain too inaccurate for routine binding prediction, and physical
experiments remain the state of the art for binding determination. The ability to accurately pre-
dict molecular binding would reduce the time-to-discovery of new treatments, help eliminate toxic
molecules early in development, and guide medicinal chemistry efforts [1, 2].
In this paper, we introduce a new predictive architecture, AtomNet, to help address these challenges.
AtomNet is novel in two regards: AtomNet is the first deep convolutional neural network for molec-
ular binding affinity prediction. It is also the first deep learning system that incorporates structural
information about the target to make its predictions.
Deep convolutional neural networks (DCNN) are currently the best performing predictive models
for speech and vision [3, 4, 5, 6]. DCNN is a class of deep neural network that constrains its model
architecture to leverage the spatial and temporal structure of its domain. For example, a low-level
image feature, such as an edge, can be described within a small spatially-proximate patch of pixels.
Such a feature detector can share evidence across the entire receptive field by “tying the weights”
of the detector neurons, as the recognition of the edge does not depend on where it is found within
1
arXiv:1510.02855v1[cs.LG]10Oct2015
Smina 123 35 5 0 0
Table 3: The number of targets on which AtomNet and Smina exceed given adjusted-logAUC thresh-
olds. For example, on the CHEMBL-20 PMD set, AtomNet achieves an adjusted-logAUC of 0.3
or better for 27 targets (out of 50 possible targets). ChEMBL-20 PMD contains 50 targets, DUDE-
30 contains 30 targets, DUDE-102 contains 102 targets, and ChEMBL-20 inactives contains 149
targets.
To overcome these limitations we take an indirect approach. Instead of directly visualizing filters
in order to understand their specialization, we apply filters to input data and examine the location
where they maximally fire. Using this technique we were able to map filters to chemical functions.
For example, Figure 5 illustrate the 3D locations at which a particular filter from our first convo-
lutional layer fires. Visual inspection of the locations at which that filter is active reveals that this
filter specializes as a sulfonyl/sulfonamide detector. This demonstrates the ability of the model to
learn complex chemical features from simpler ones. In this case, the filter has inferred a meaningful
spatial arrangement of input atom types without any chemical prior knowledge.
Figure 5: Sulfonyl/sulfonamide detection with autonomously trained convolutional filters.
8
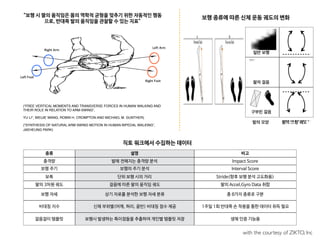
• 이미 알려진 단백질-리간드 3차원 결합 구조를 딥러닝(CNN)으로 학습
• 화학 결합 등에 대한 계산 없이도, 단백질-리간드 결합 여부를 계산
• 기존의 구조기반 예측 등 대비, 딥러닝으로 더 정확히 예측하였음](https://image.slidesharecdn.com/merck1hrv16digitalmedicine180115-180115023755/85/slide-224-320.jpg)
![AtomNet: A Deep Convolutional Neural Network for
Bioactivity Prediction in Structure-based Drug
Discovery
Izhar Wallach
Atomwise, Inc.
izhar@atomwise.com
Michael Dzamba
Atomwise, Inc.
misko@atomwise.com
Abraham Heifets
Atomwise, Inc.
abe@atomwise.com
Abstract
Deep convolutional neural networks comprise a subclass of deep neural networks
(DNN) with a constrained architecture that leverages the spatial and temporal
structure of the domain they model. Convolutional networks achieve the best pre-
dictive performance in areas such as speech and image recognition by hierarchi-
cally composing simple local features into complex models. Although DNNs have
been used in drug discovery for QSAR and ligand-based bioactivity predictions,
none of these models have benefited from this powerful convolutional architec-
ture. This paper introduces AtomNet, the first structure-based, deep convolutional
neural network designed to predict the bioactivity of small molecules for drug dis-
covery applications. We demonstrate how to apply the convolutional concepts of
feature locality and hierarchical composition to the modeling of bioactivity and
chemical interactions. In further contrast to existing DNN techniques, we show
that AtomNet’s application of local convolutional filters to structural target infor-
mation successfully predicts new active molecules for targets with no previously
known modulators. Finally, we show that AtomNet outperforms previous docking
approaches on a diverse set of benchmarks by a large margin, achieving an AUC
greater than 0.9 on 57.8% of the targets in the DUDE benchmark.
1 Introduction
Fundamentally, biological systems operate through the physical interaction of molecules. The ability
to determine when molecular binding occurs is therefore critical for the discovery of new medicines
and for furthering of our understanding of biology. Unfortunately, despite thirty years of compu-
tational efforts, computer tools remain too inaccurate for routine binding prediction, and physical
experiments remain the state of the art for binding determination. The ability to accurately pre-
dict molecular binding would reduce the time-to-discovery of new treatments, help eliminate toxic
molecules early in development, and guide medicinal chemistry efforts [1, 2].
In this paper, we introduce a new predictive architecture, AtomNet, to help address these challenges.
AtomNet is novel in two regards: AtomNet is the first deep convolutional neural network for molec-
ular binding affinity prediction. It is also the first deep learning system that incorporates structural
information about the target to make its predictions.
Deep convolutional neural networks (DCNN) are currently the best performing predictive models
for speech and vision [3, 4, 5, 6]. DCNN is a class of deep neural network that constrains its model
architecture to leverage the spatial and temporal structure of its domain. For example, a low-level
image feature, such as an edge, can be described within a small spatially-proximate patch of pixels.
Such a feature detector can share evidence across the entire receptive field by “tying the weights”
of the detector neurons, as the recognition of the edge does not depend on where it is found within
1
arXiv:1510.02855v1[cs.LG]10Oct2015
• 이미 알려진 단백질-리간드 3차원 결합 구조를 딥러닝(CNN)으로 학습
• 화학 결합 등에 대한 계산 없이도, 단백질-리간드 결합 여부를 계산
• 기존의 구조기반 예측 등 대비, 딥러닝으로 더 정확히 예측하였음](https://image.slidesharecdn.com/merck1hrv16digitalmedicine180115-180115023755/85/slide-225-320.jpg)




































