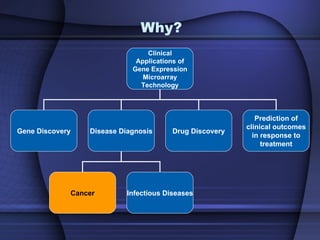
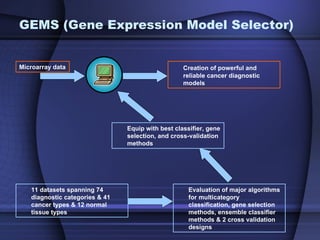
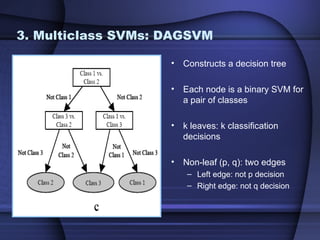
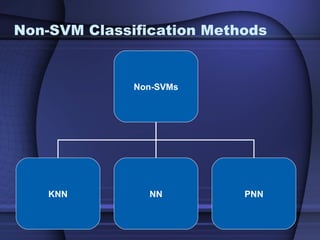
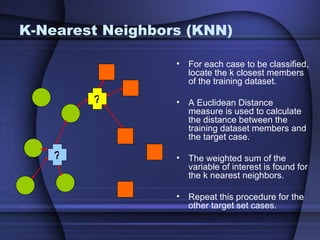
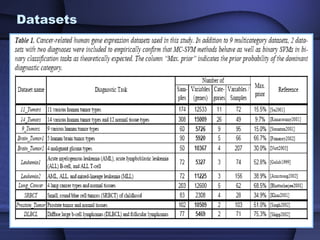
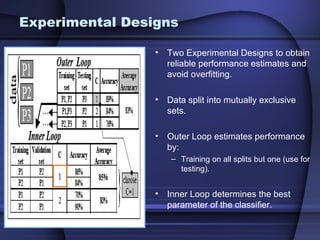
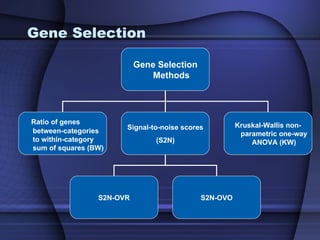
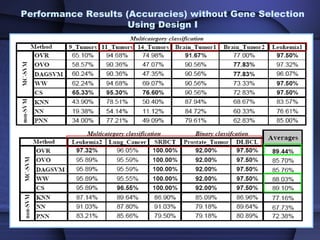
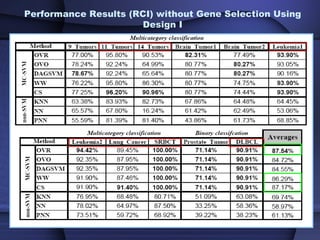
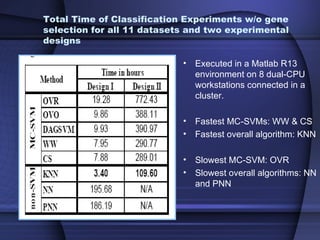
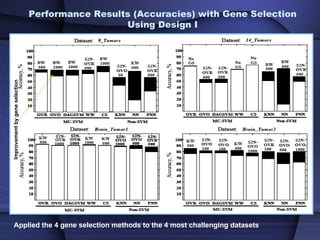
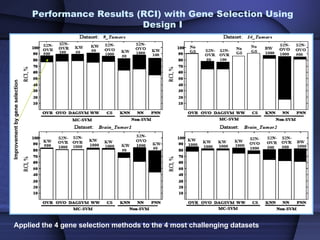
This document summarizes a study that evaluated multiple classification methods for cancer diagnosis using microarray gene expression data. It tested support vector machines (SVMs), other classifiers, and ensemble methods on 11 cancer datasets. Gene selection improved the performance of some methods. Overall, multiclass SVMs like one-versus-rest, Weston-Watkins, and Crammer-Singer performed best for cancer diagnosis from microarray gene expression data.