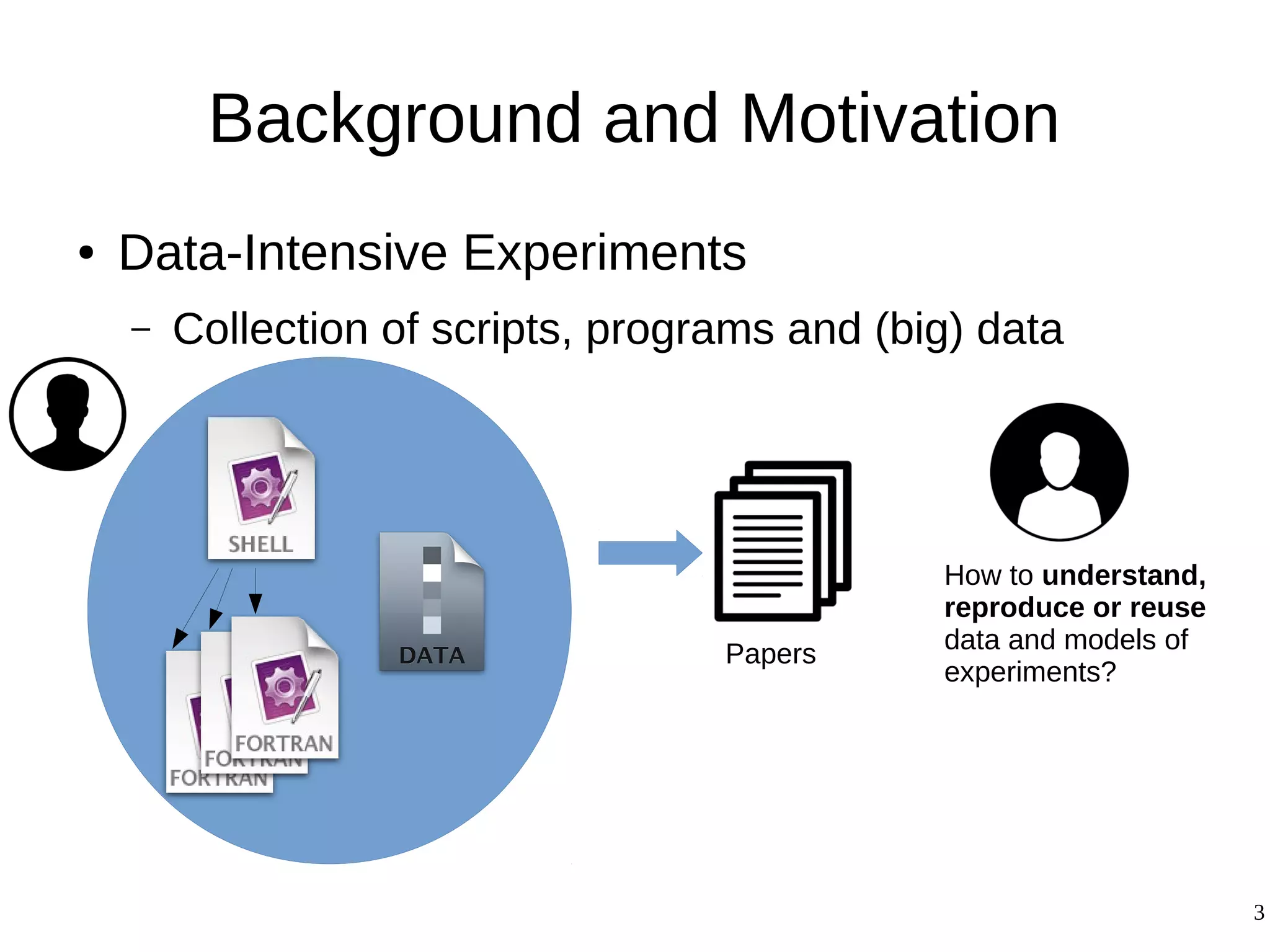
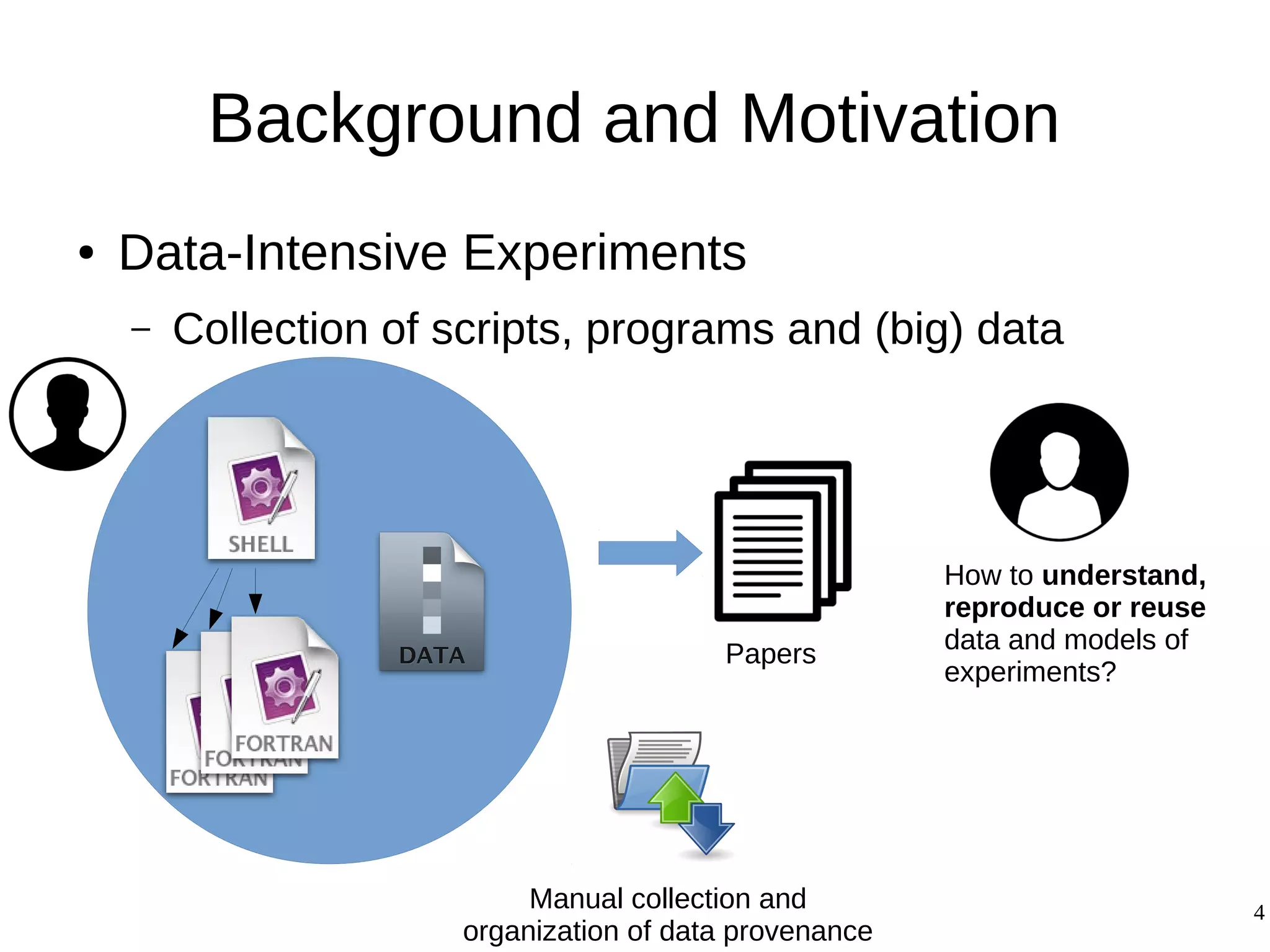
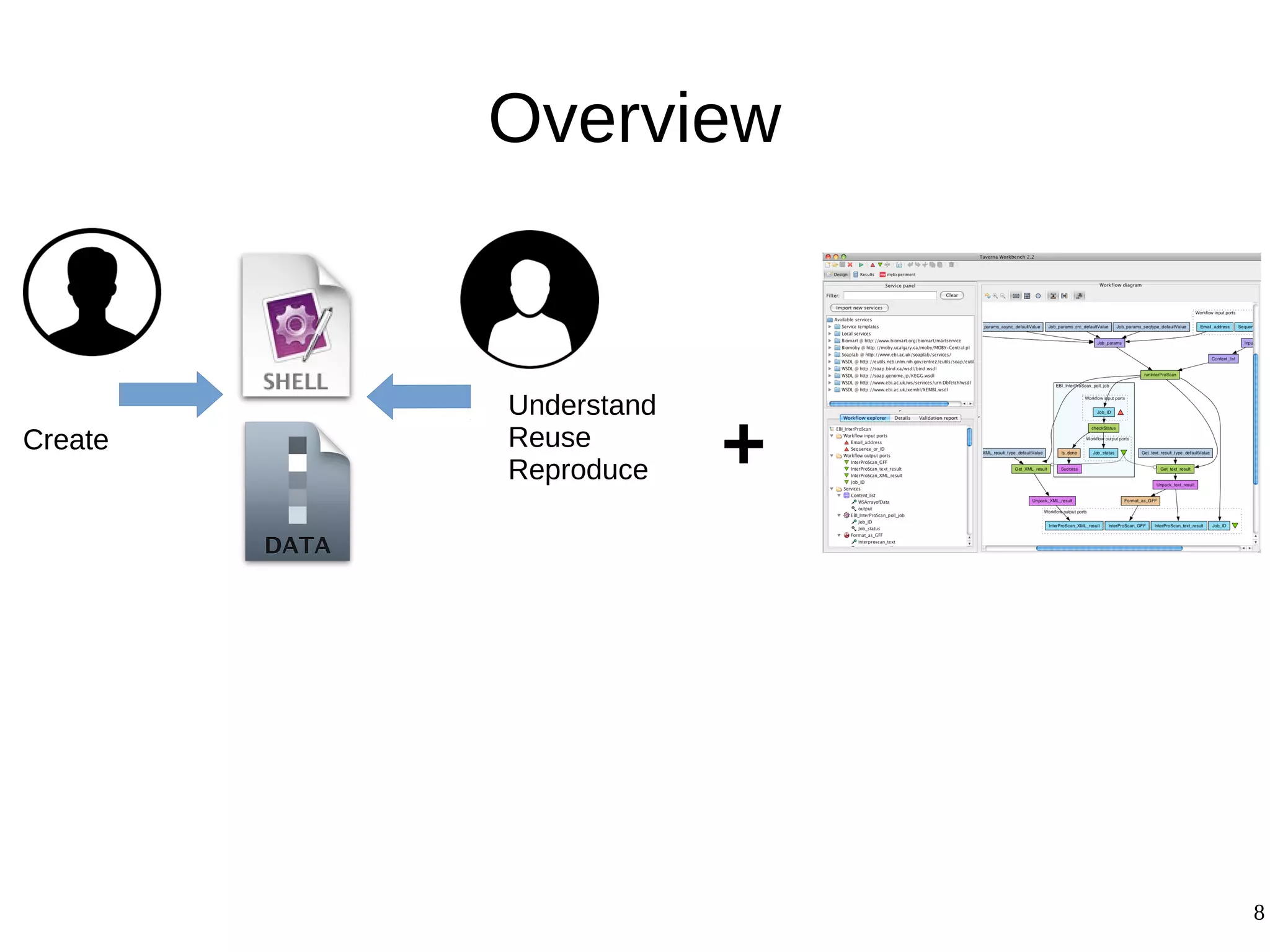
1) The document presents a methodology to convert script-based experiments into reproducible workflow research objects (WROs). This addresses issues of understanding, reusing, and reproducing experiments conducted through scripts.
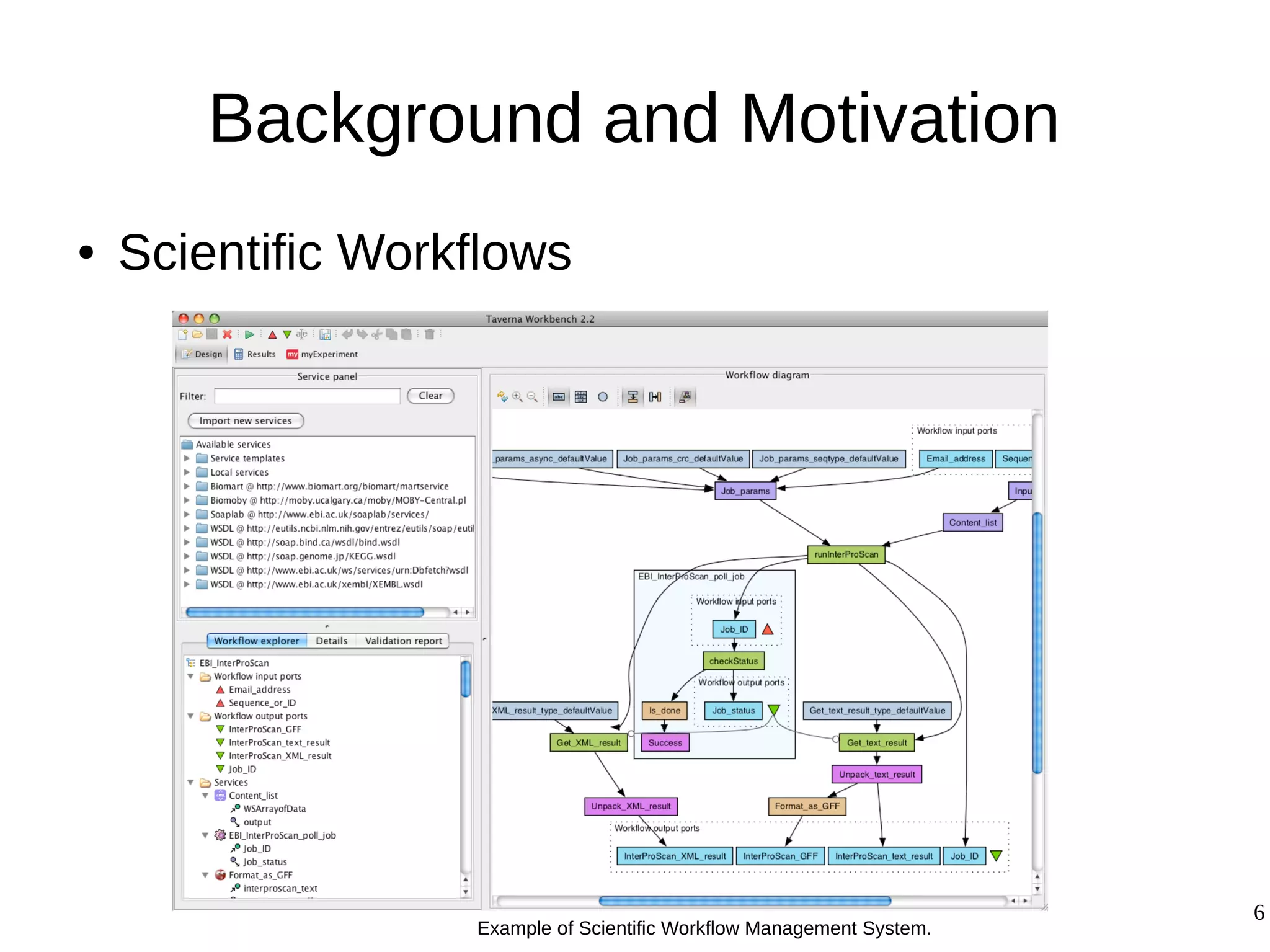
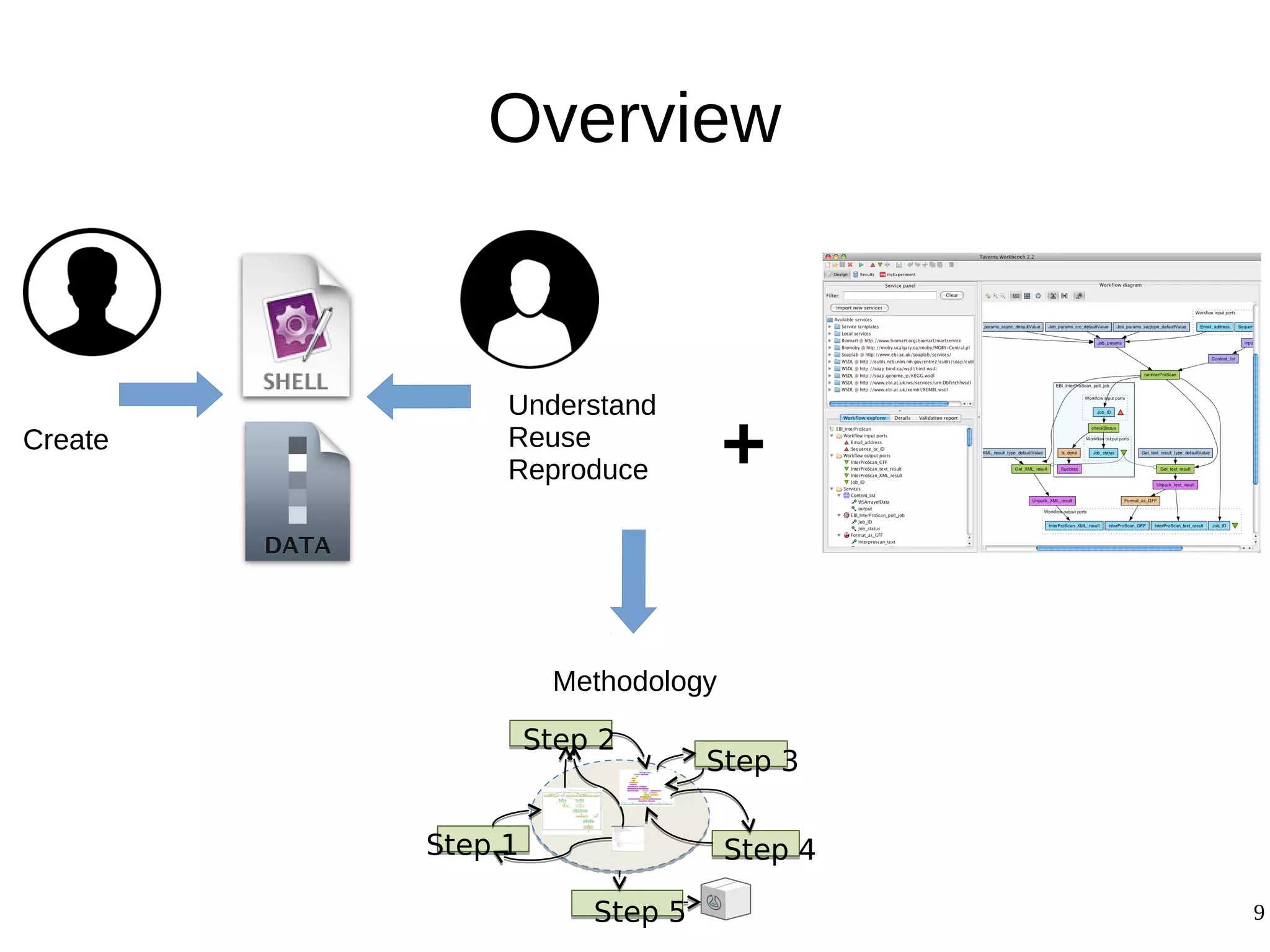
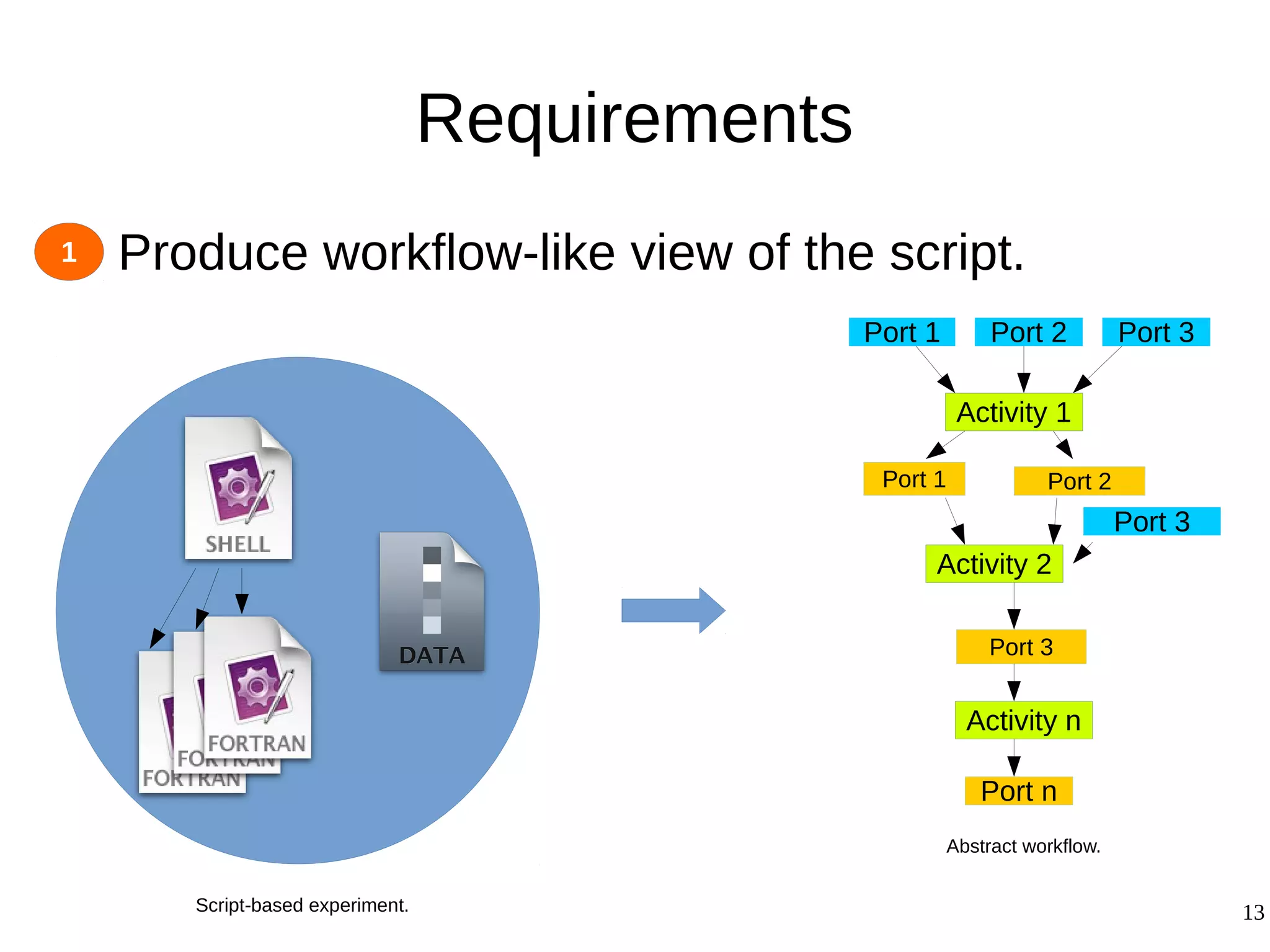
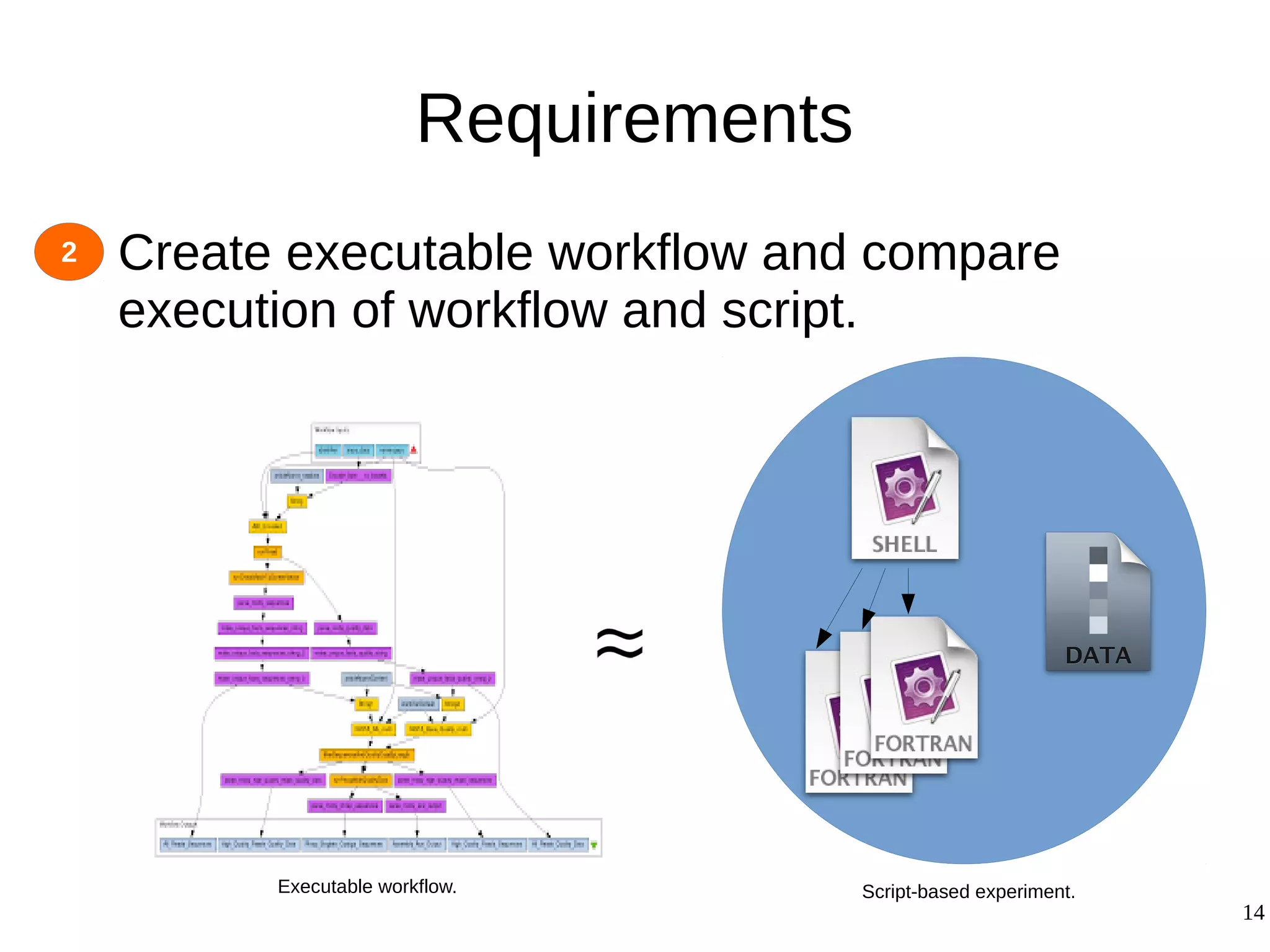
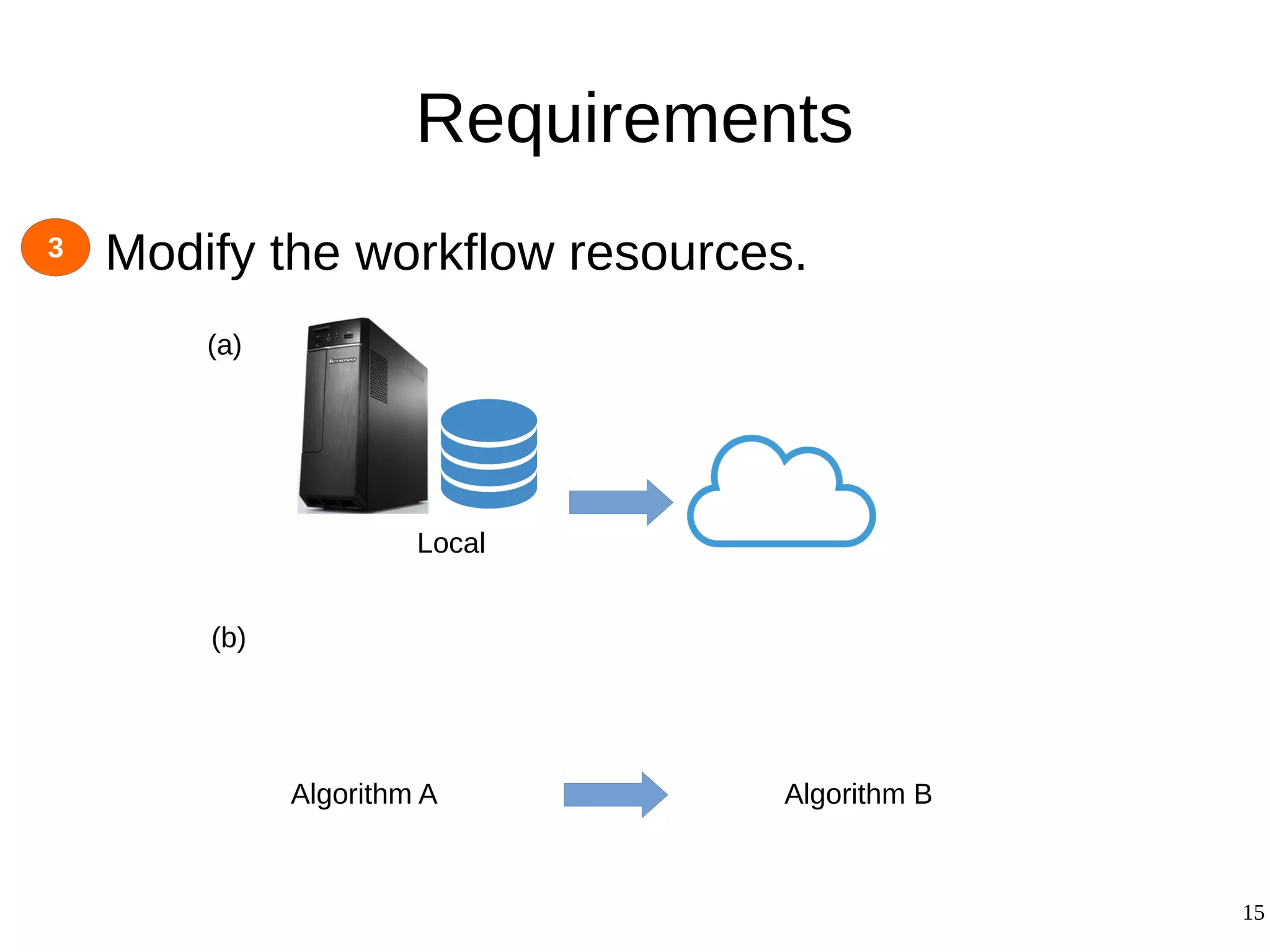
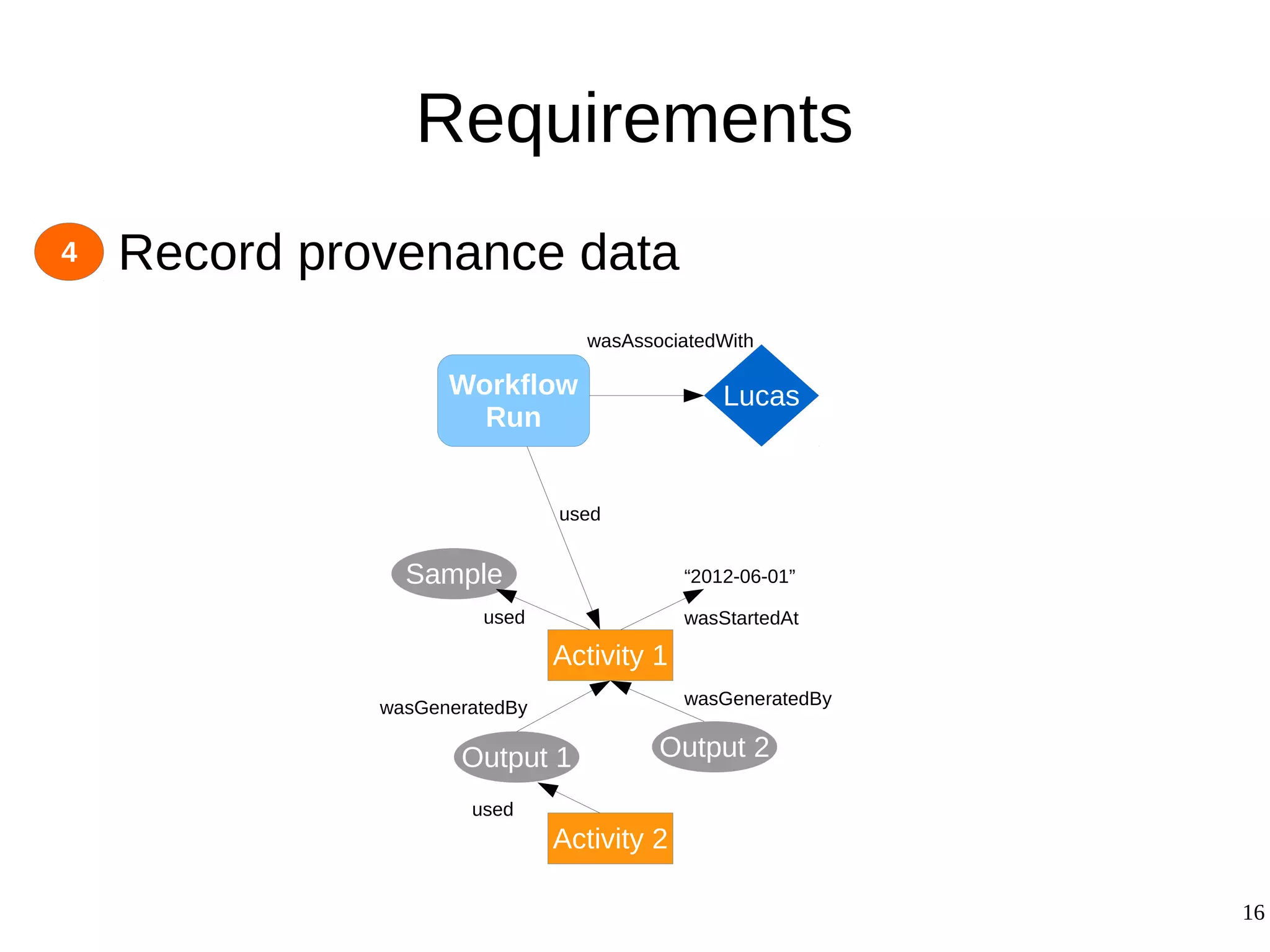
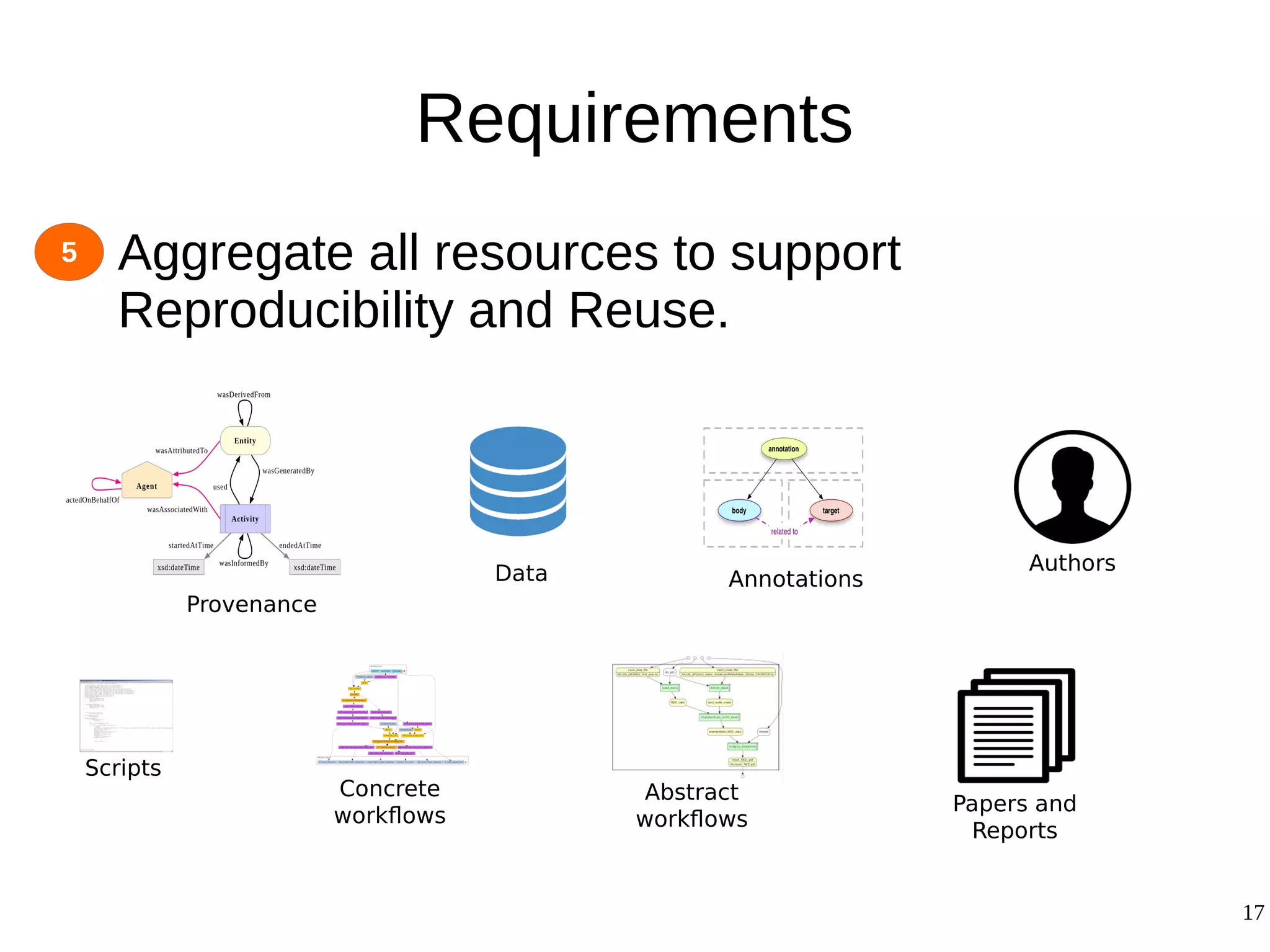
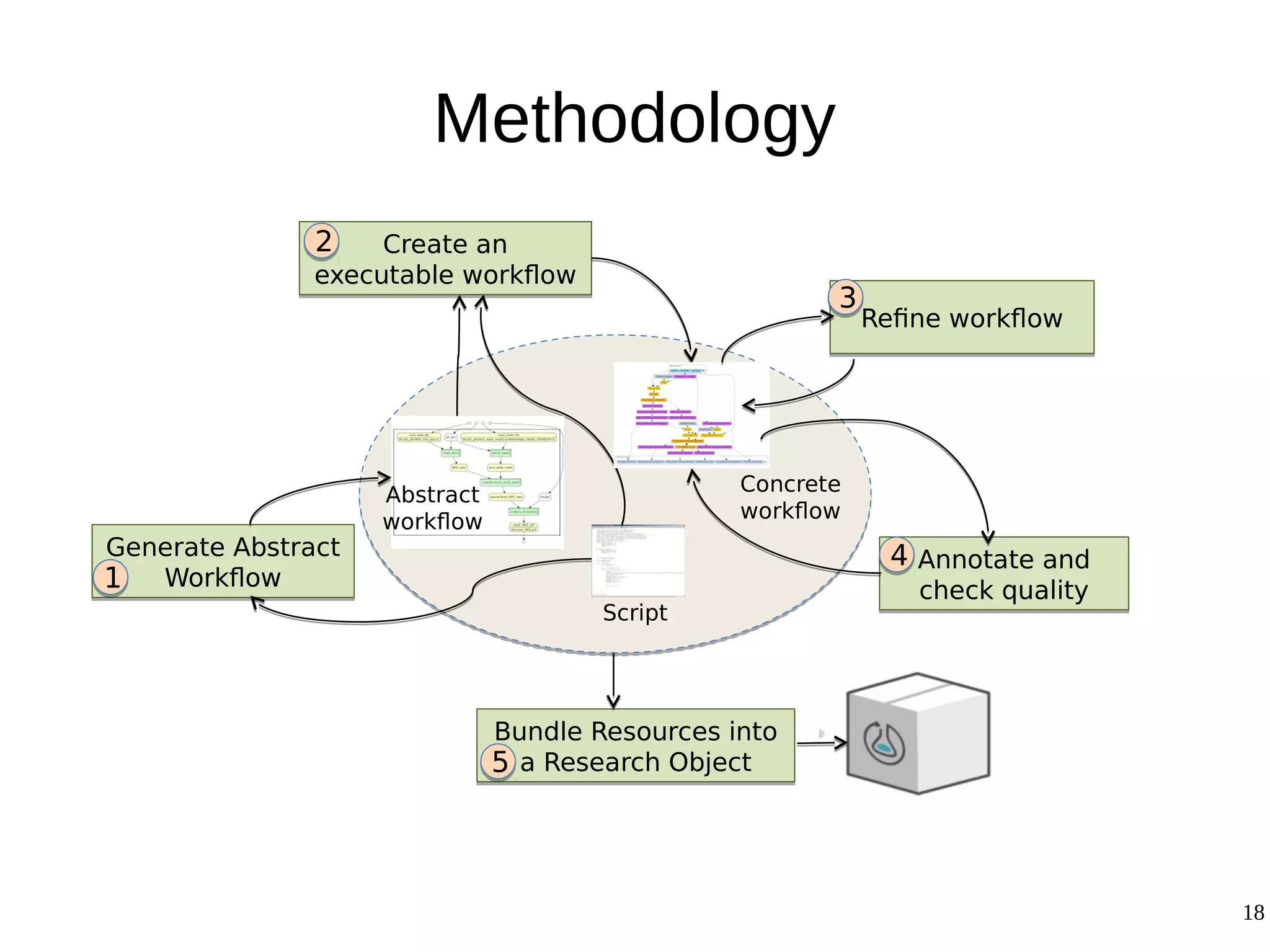
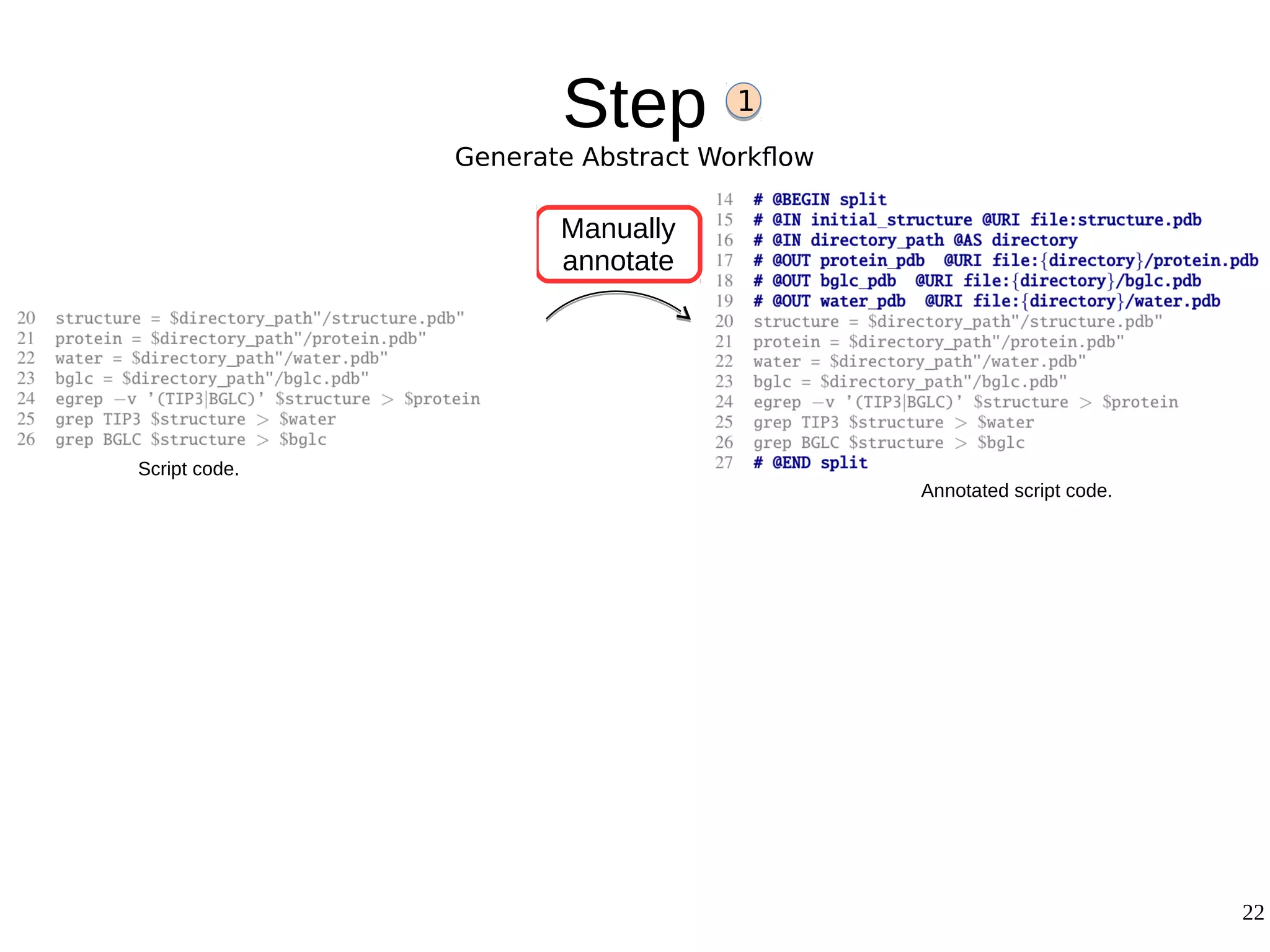
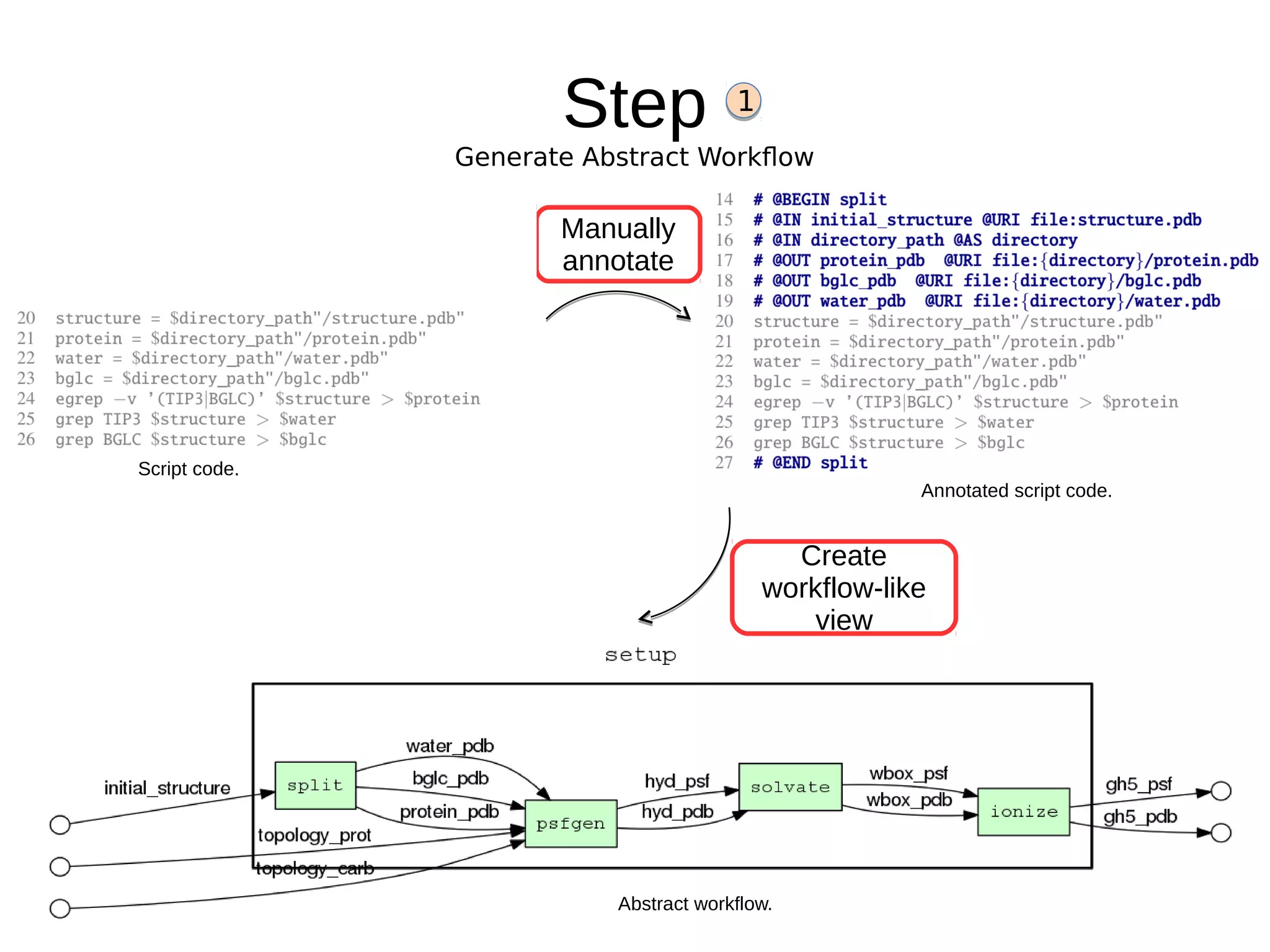
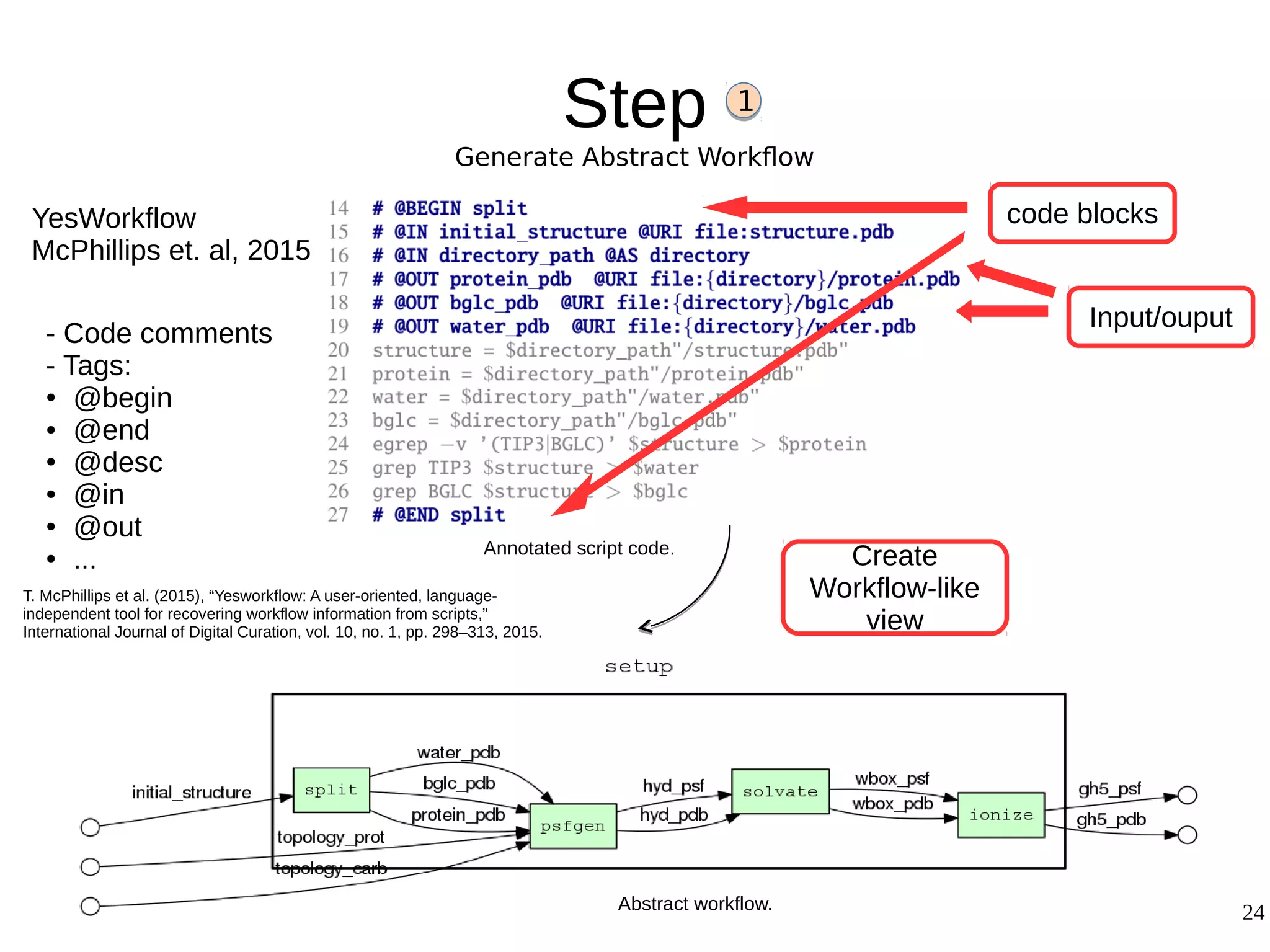
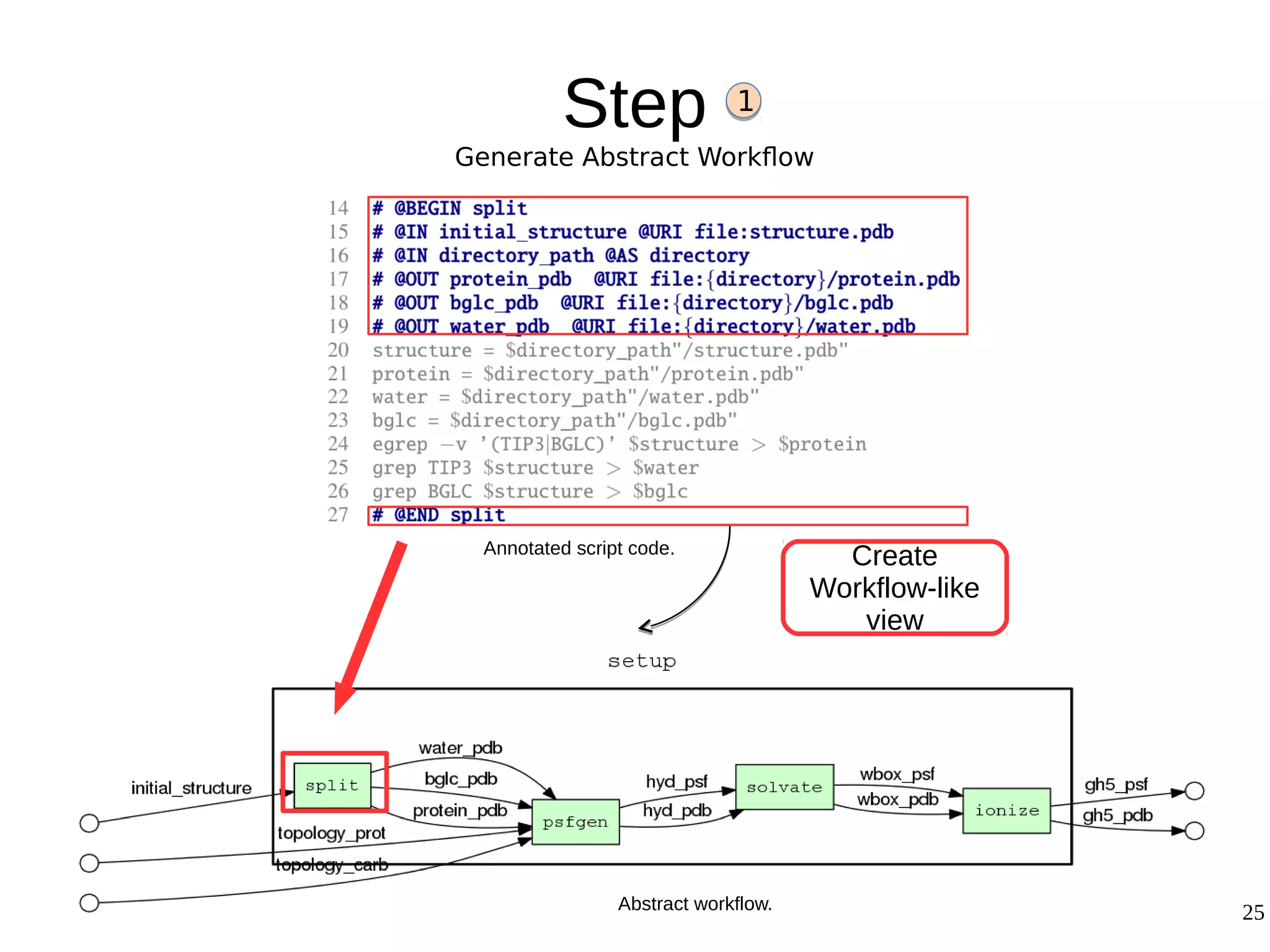
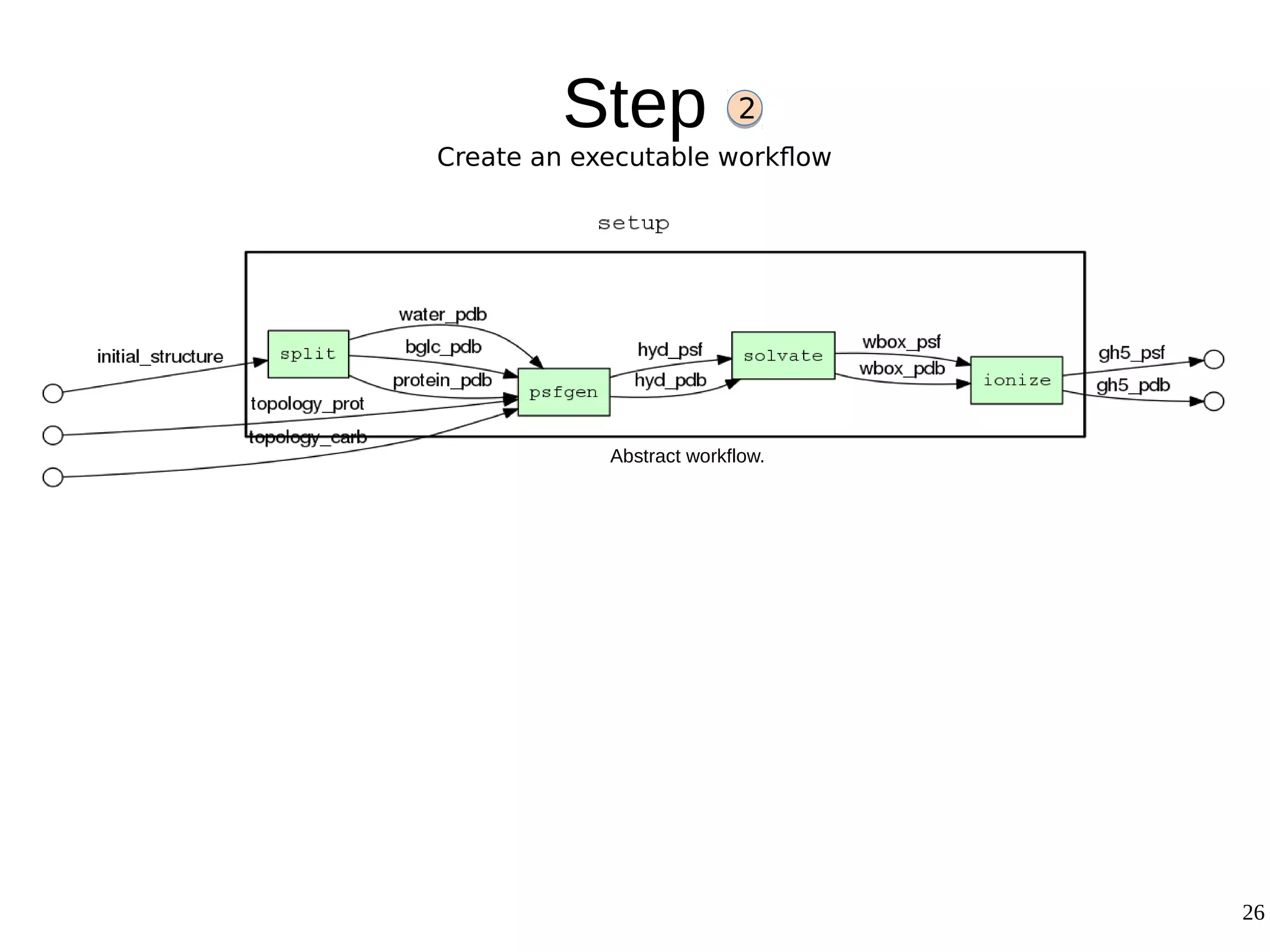
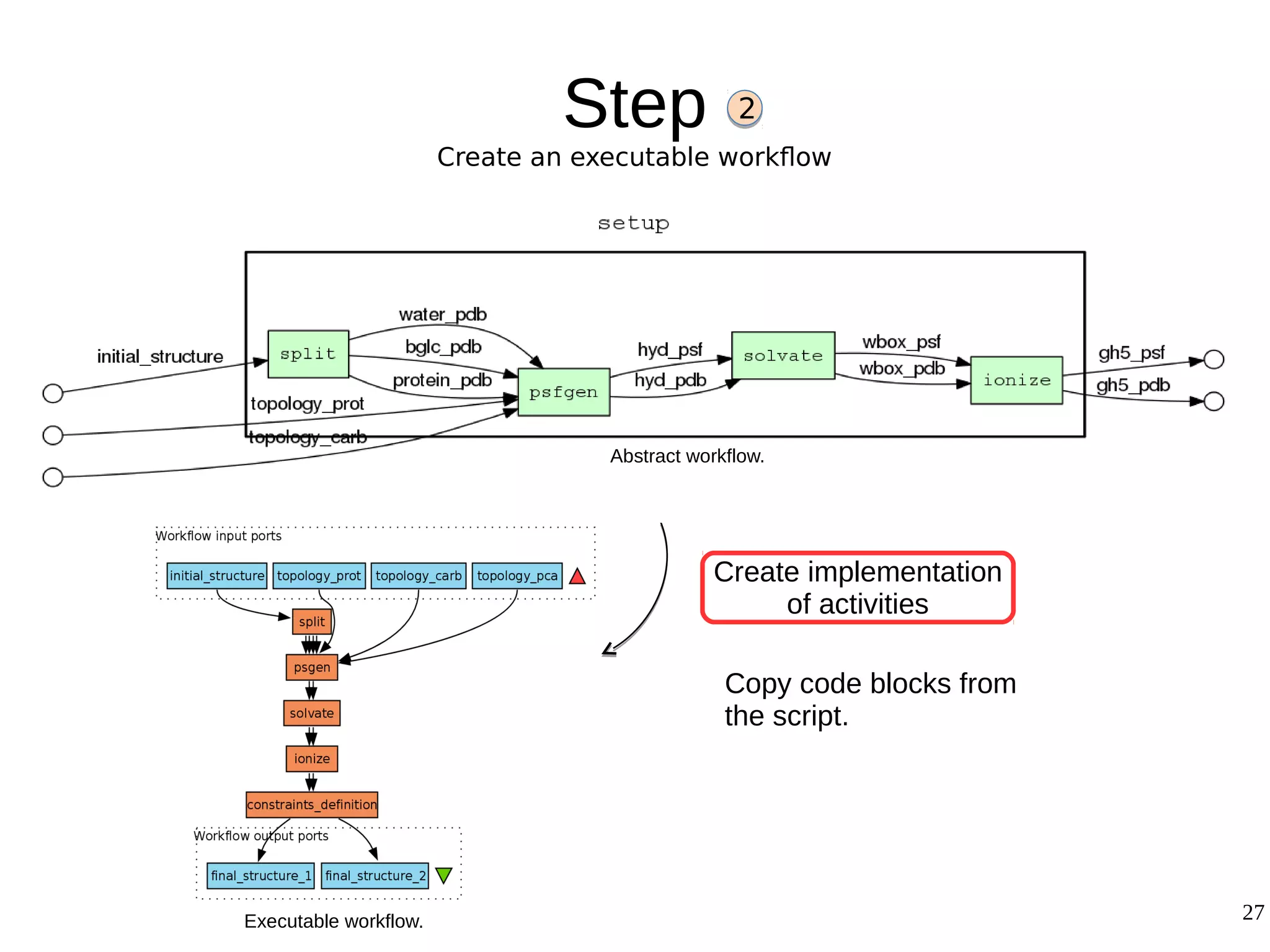
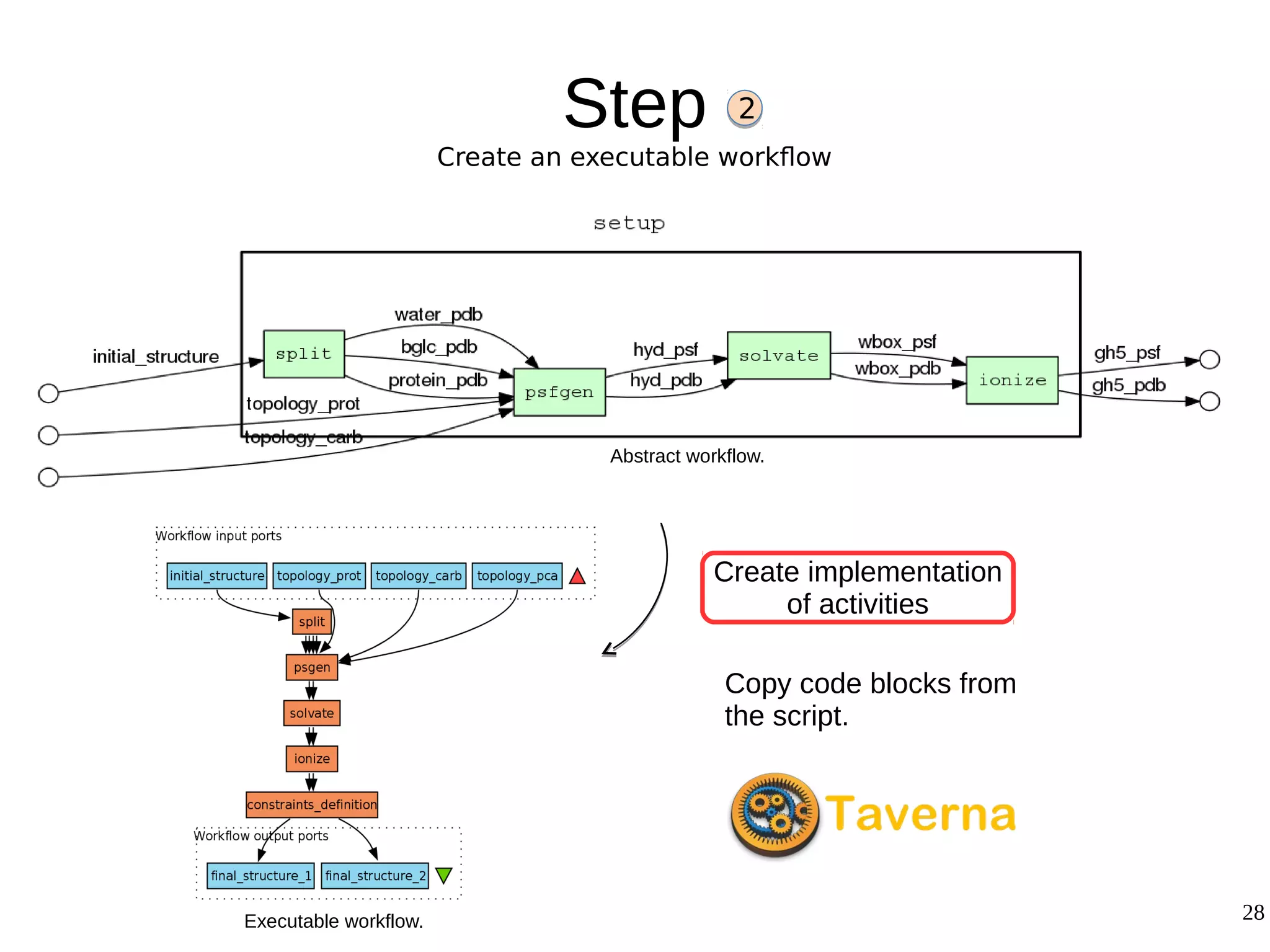
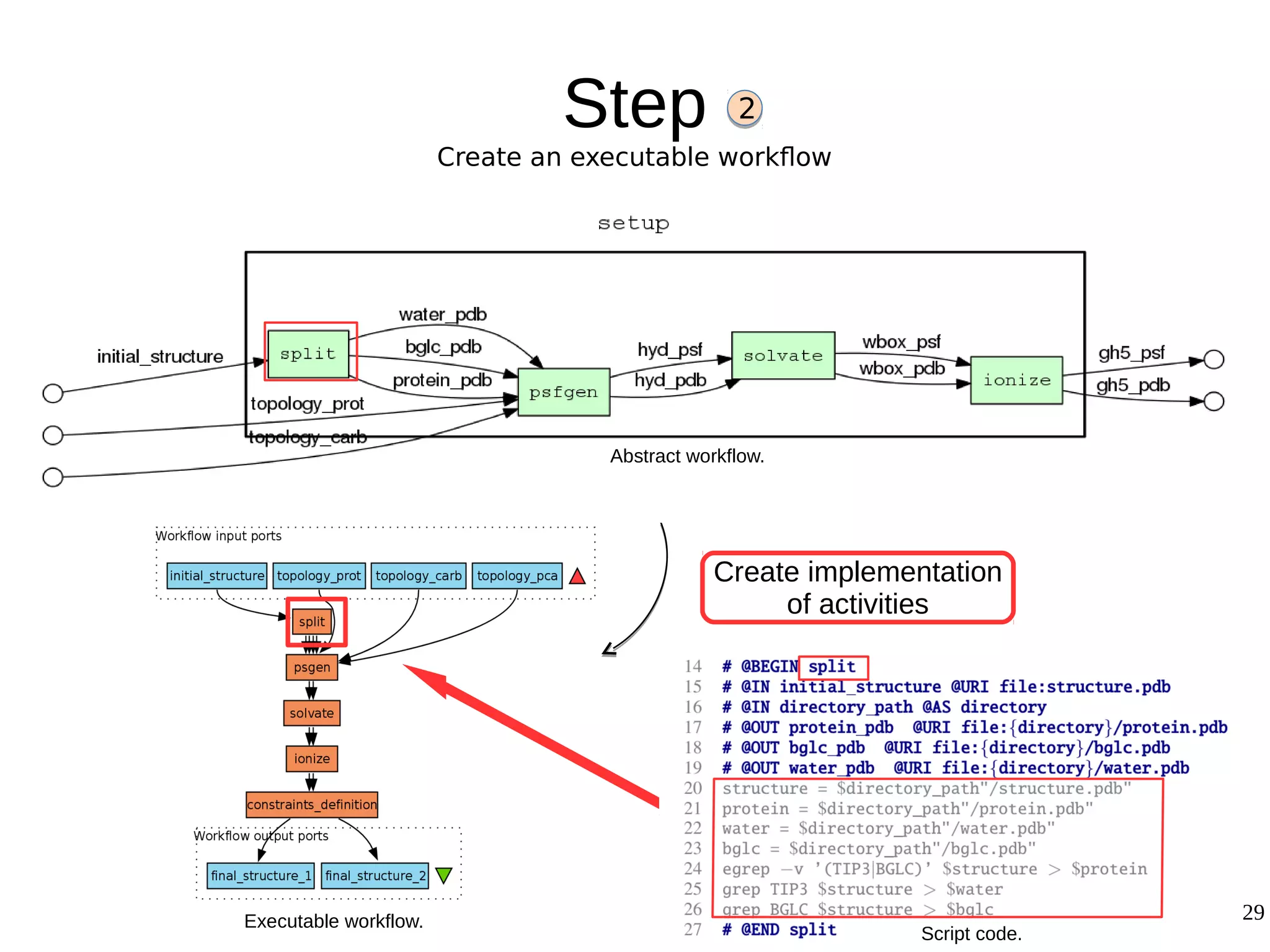
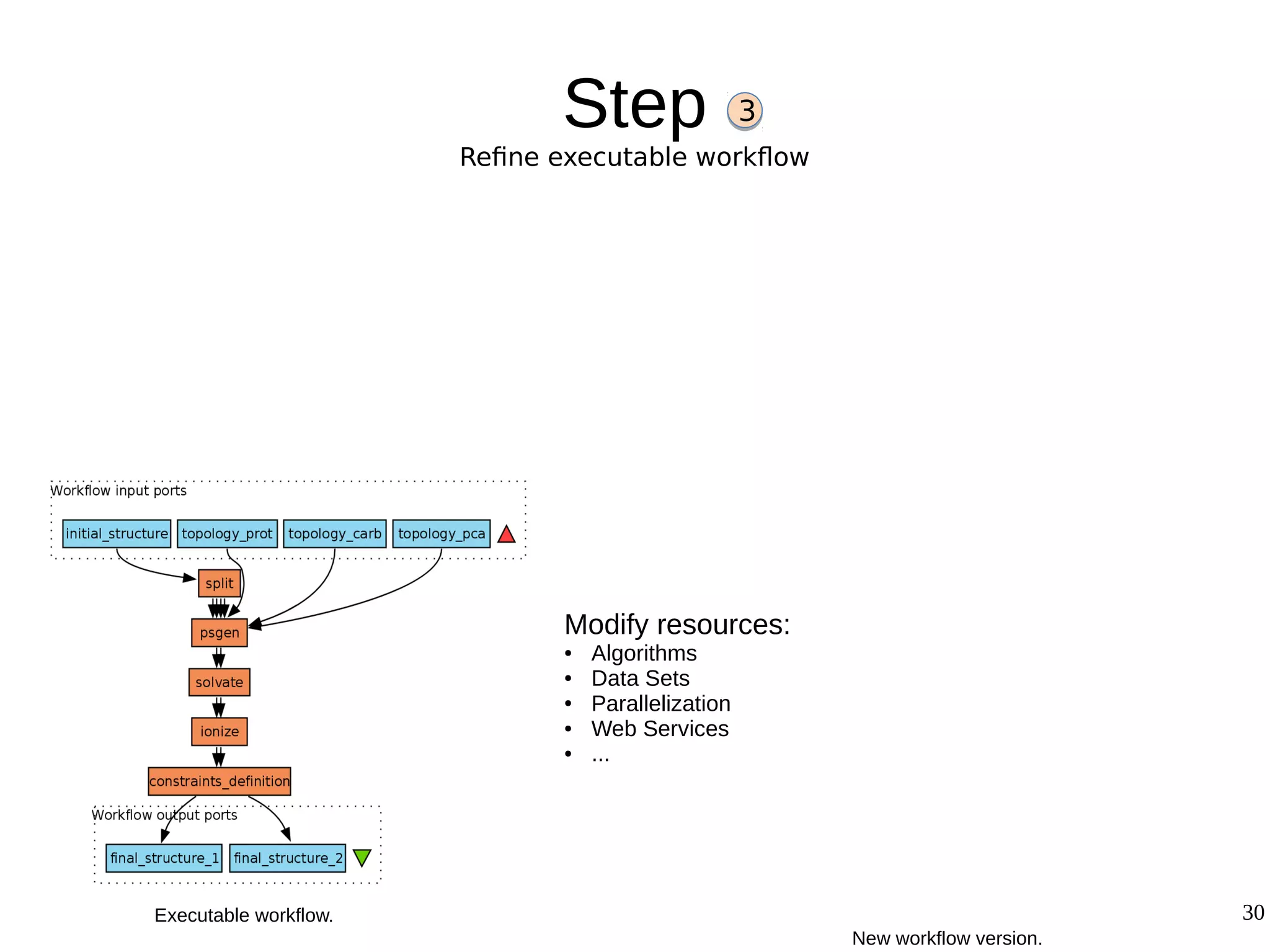
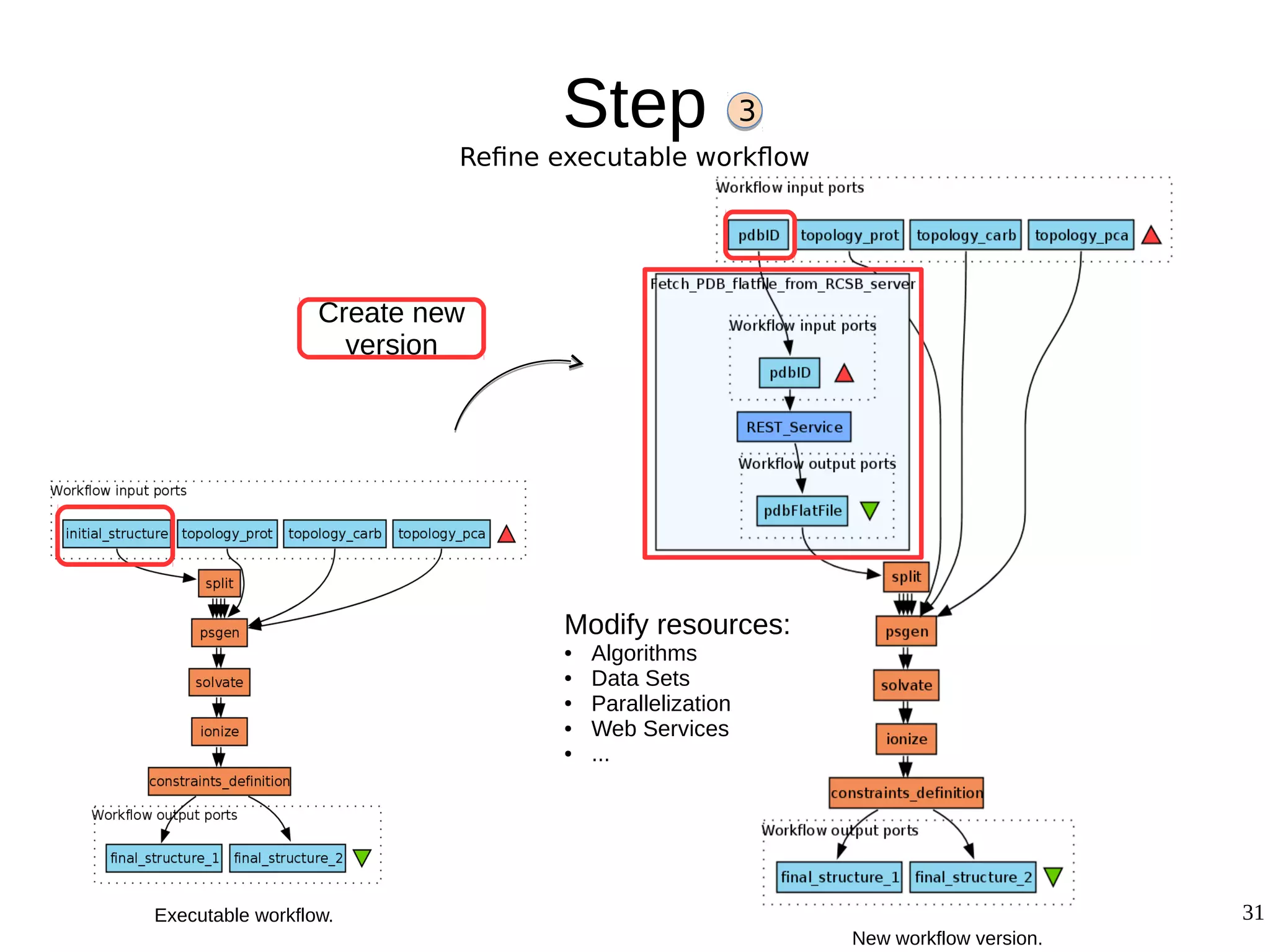
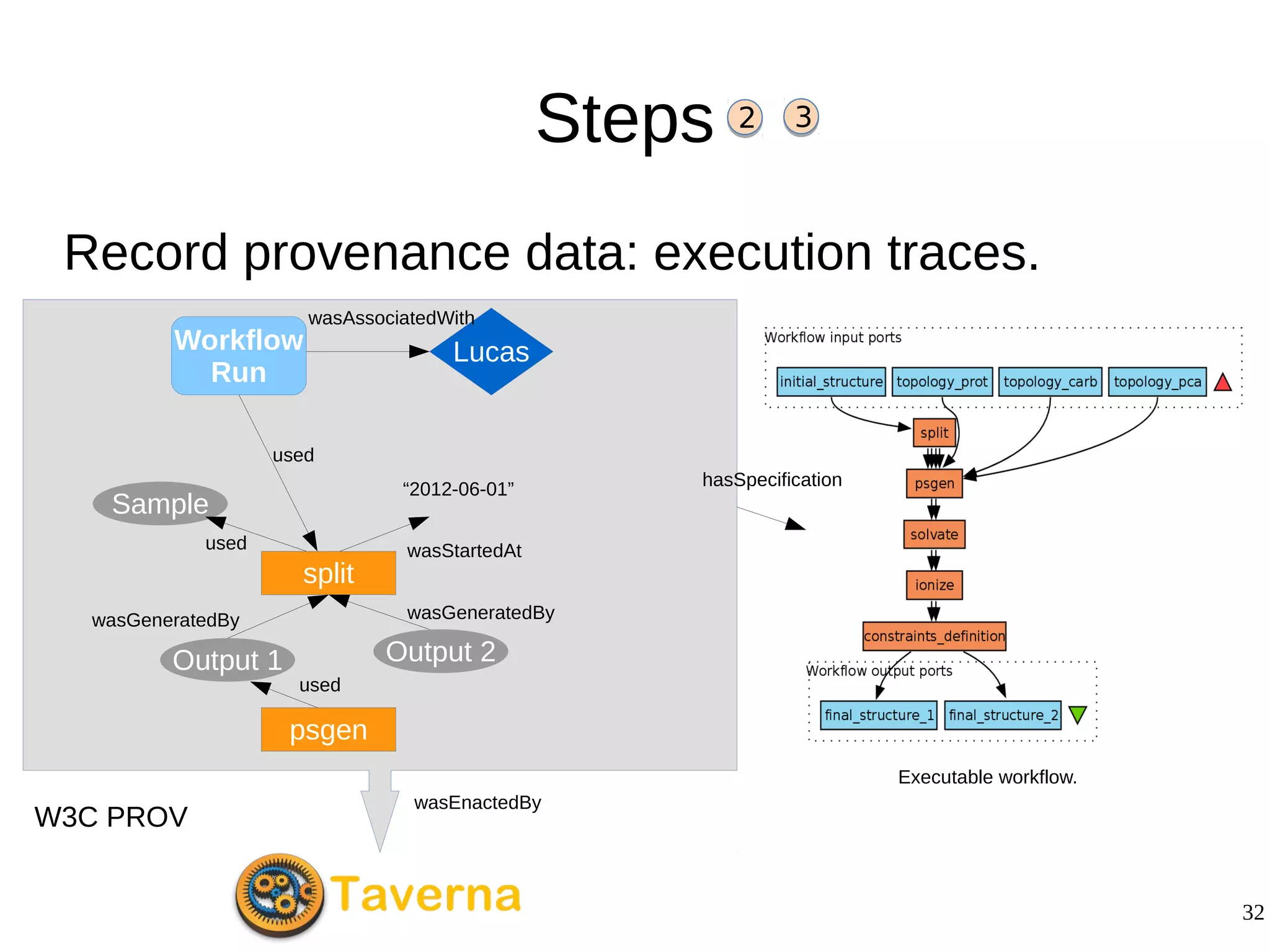
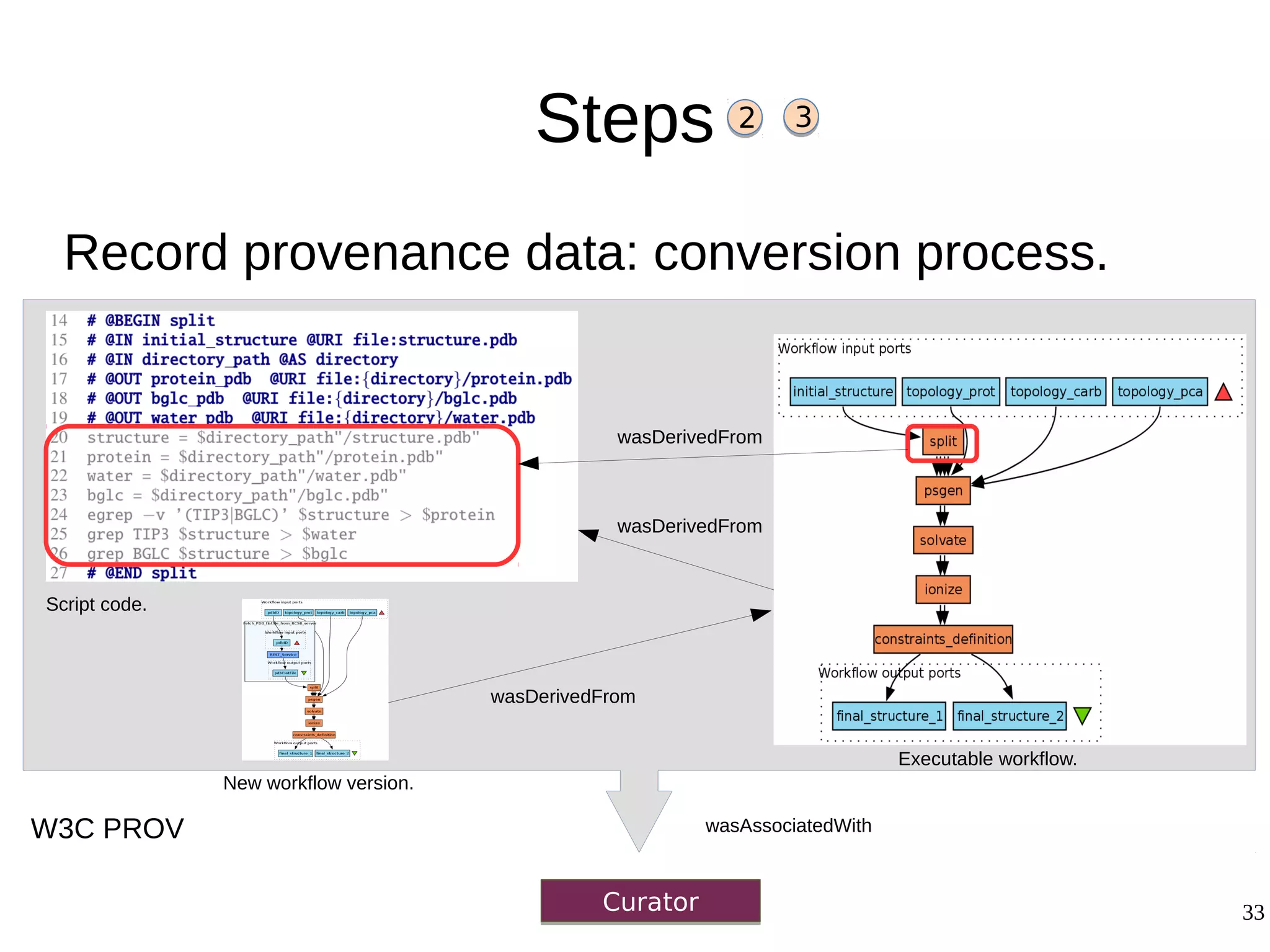
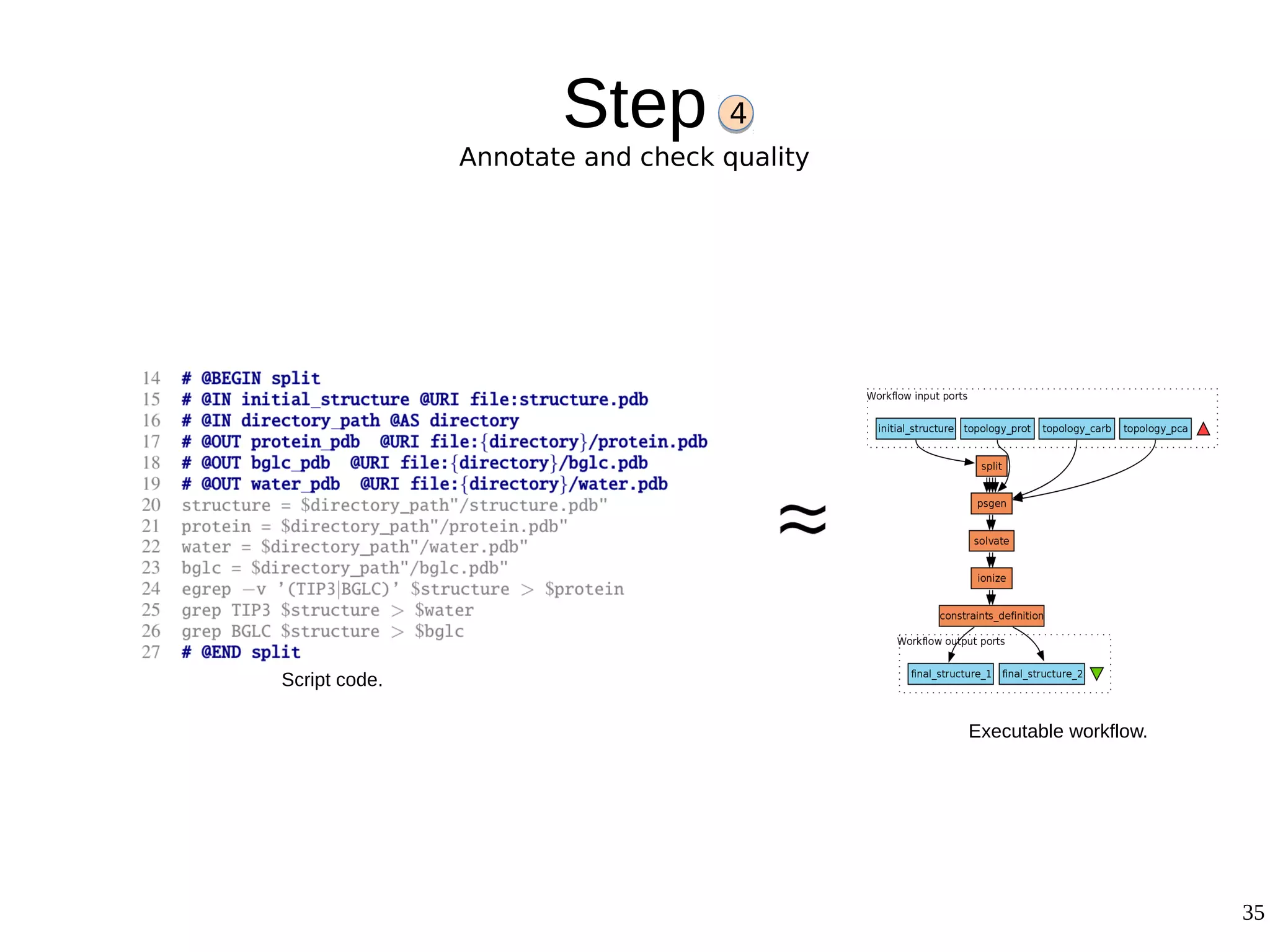
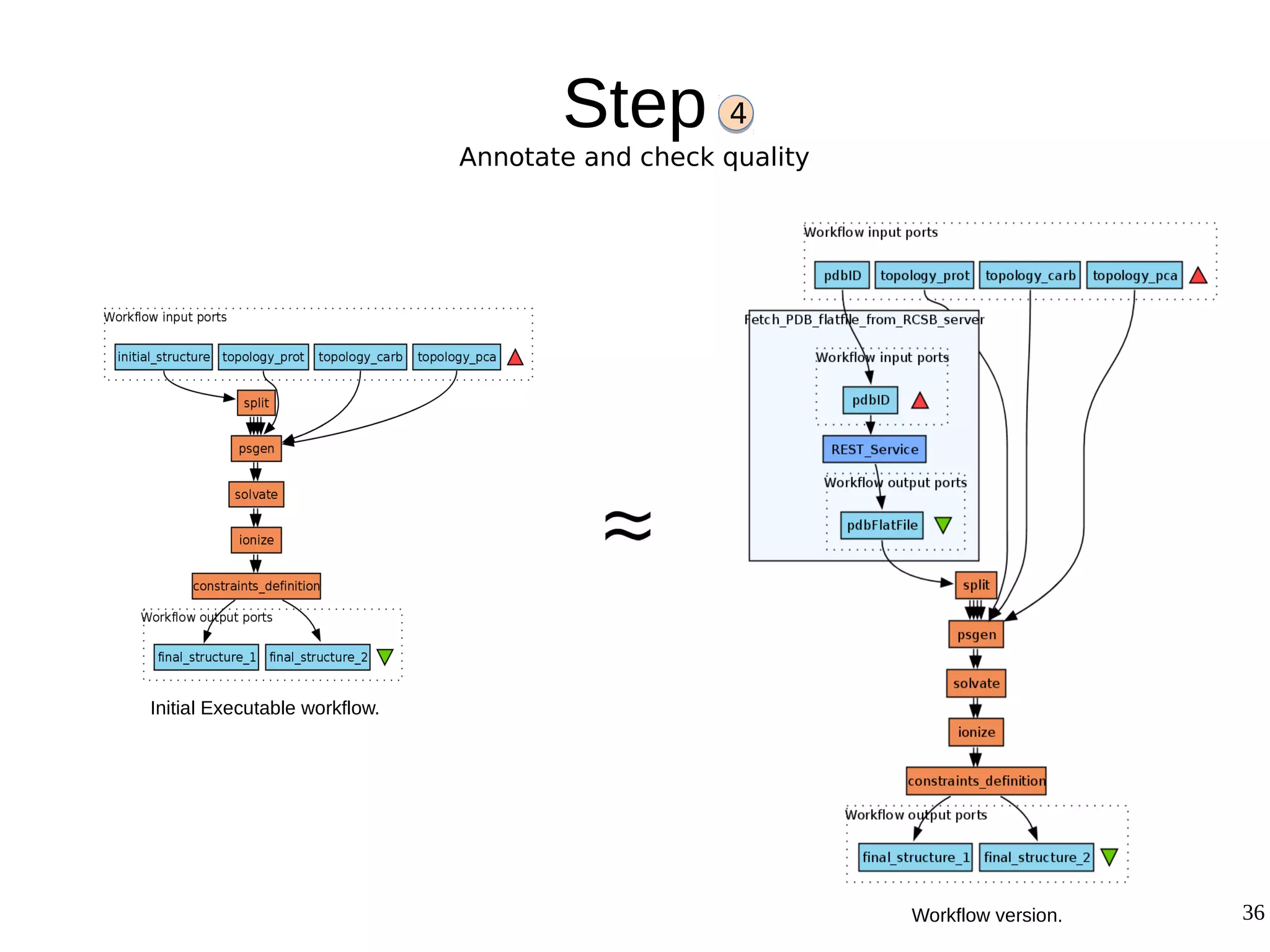
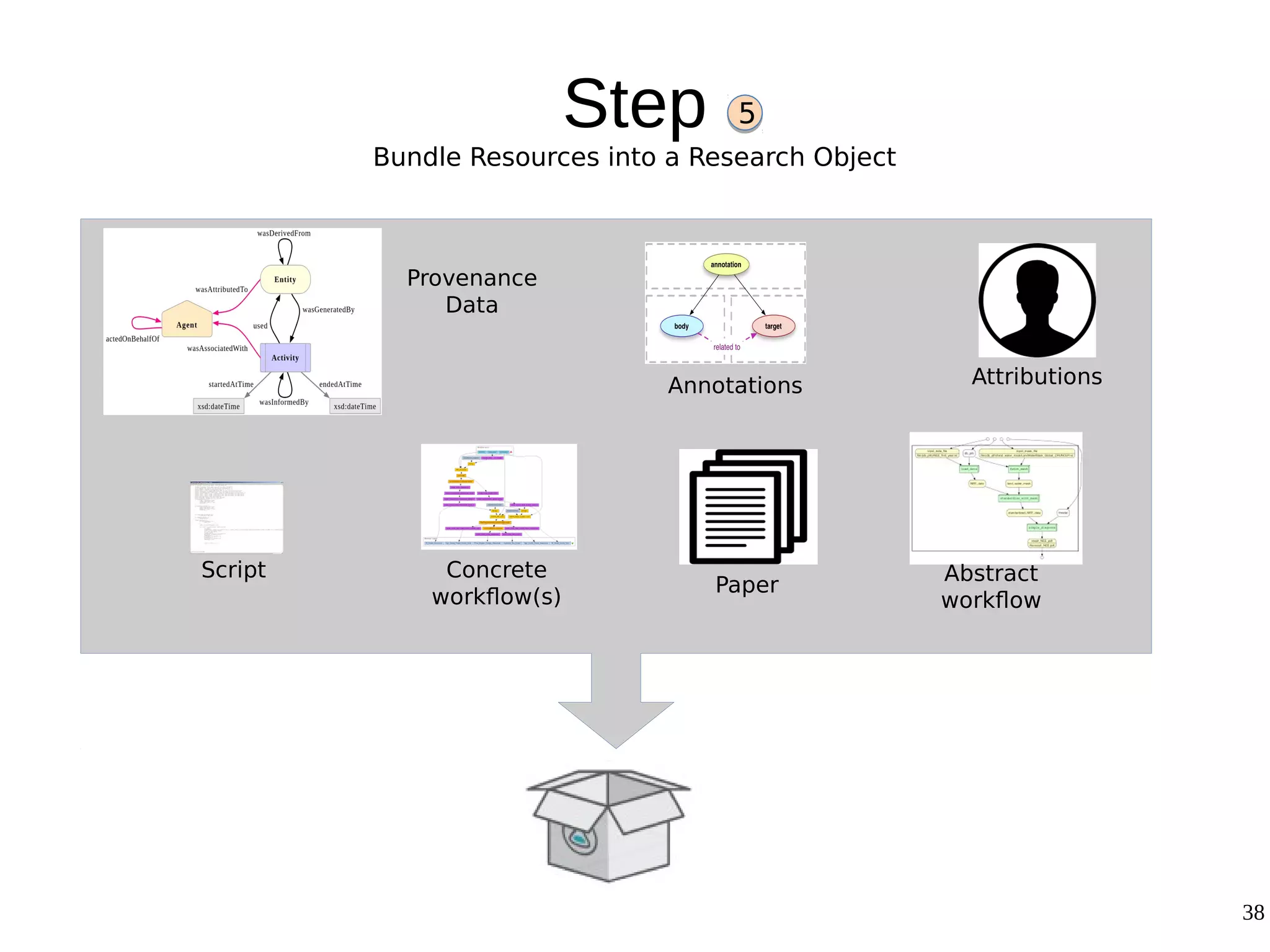
2) The methodology involves 5 steps: generate an abstract workflow, create an executable workflow, refine the workflow, record provenance data, and annotate and bundle resources into a WRO.
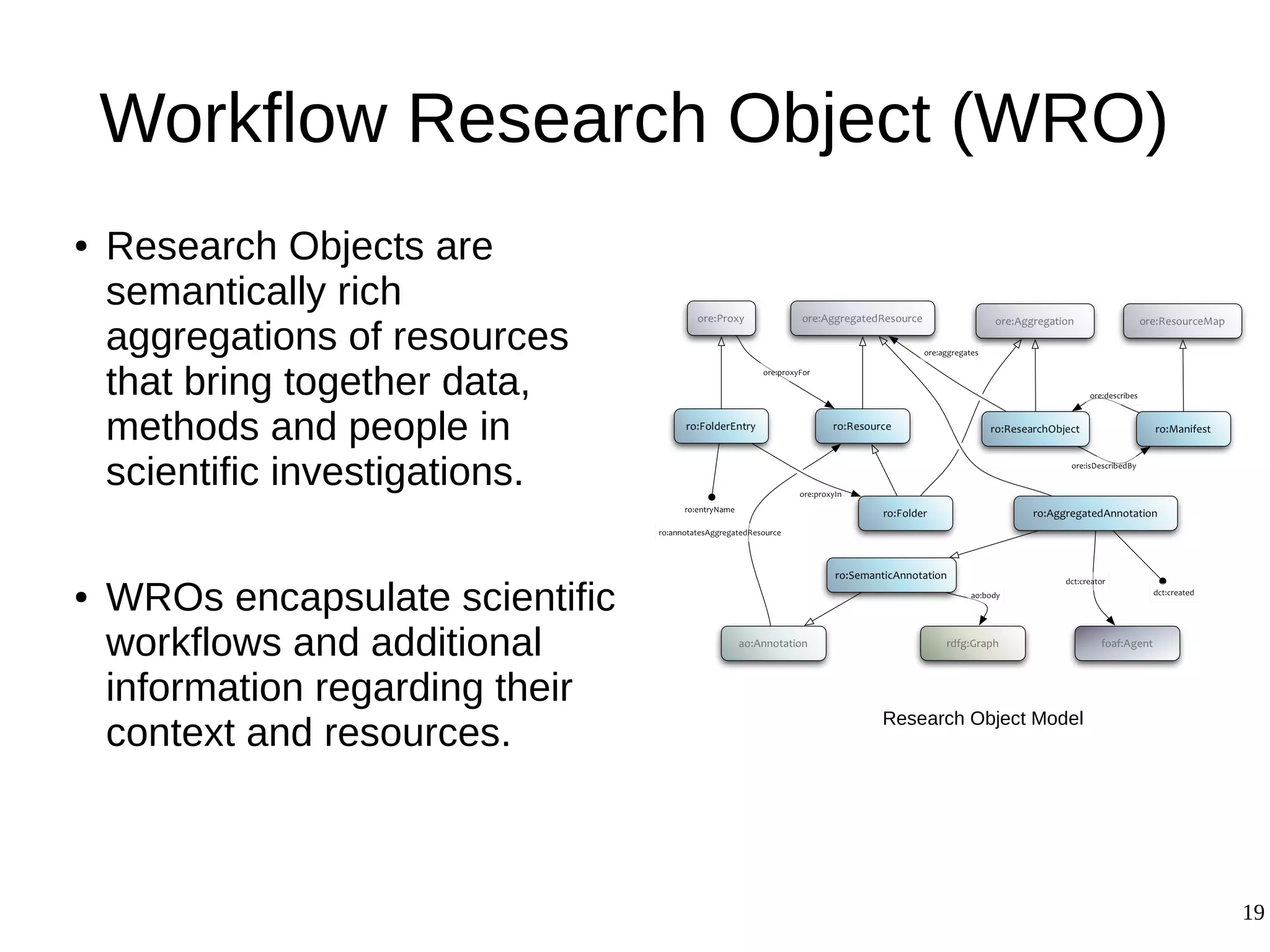
3) The methodology was demonstrated on a molecular dynamics simulation case study. It aims to produce executable workflows, record provenance of the conversion process, and bundle resources like scripts, workflows, annotations, and papers to support reproducibility and reuse.