ATM
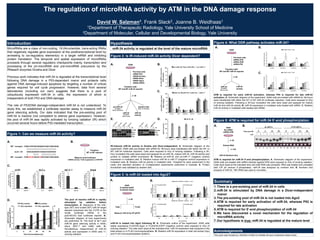
- 1. The regulation of microRNA activity by ATM in the DNA damage response David W. Salzman1, Frank Slack2, Joanne B. Weidhaas1 1Department of Therapeutic Radiology, Yale University School of Medicine 2Department of Molecular, Cellular and Developmental Biology, Yale University MicroRNAs are a class of non-coding, 18-24nucleotide, trans-acting RNAs that negatively regulate gene expression at the posttranscriptional level by annealing to cis-regulatory element(s) in a target mRNA and inhibiting protein translation. The temporal and spatial expression of microRNAs proceeds through several regulatory checkpoints mainly, transcription and processing of the pri-microRNA and pre-microRNA precursors by the RNaseIII enzymes Drosha and Dicer. Previous work indicates that miR-34 is regulated at the transcriptional level following DNA damage in a P53-dependent manor and protects cells against DNA damage-induced apoptosis by targeting a number of critical genes required for cell cycle progression. However, data from several laboratories (including our own) suggests that there is a pool of ubiquitously expressed miR-34 in cells, the expression of which is independent of both P53 and DNA damage. The role of P53/DNA damage-independent miR-34 is not understood. To study this, we established a luciferase reporter assay to measure miR-34 gene silencing activity. Our data indicated that the pre-existing pool of miR-34 is inactive (not competent to silence gene expression). However, the pool of miR-34 was rapidly activated by ionizing radiation (IR) which occurred several hours before P53-mediated transcription. Introduction 0 2 4 6 8 10 12 14 16 let-7 miR-34 microRNAactivity (folddifferenceMT/WT) MCF-7 0 2 4 6 8 10 12 14 16 18 let-7 miR-34 microRNAactivity (folddifferenceMT/WT) H460 0 2 4 6 8 10 12 14 let-7 miR-34 microRNAactivity (folddifferenceMT/WT) A549 0 1 2 3 4 5 6 7 8 9 10 let-7 miR-34 microRNAactivity (folddifferenceMT/WT) HeLa Figure 1: Can we measure miR-34 activity? Hypothesis WT target CGACCGUCACAGAAUCGACCAACAGA |||||||||||||||||||||| miR-34 UGGCAGUGUCUUAGCUGGUUGU | |||| ||||||||||| MT target CGAGGCACACACUAUCGACCAACAGA A. B. Transfect: WT or MT reporter Measure dual-luciferase miRNA activity = fold suppression of MT/WT C. D. The pool of inactive miR-34 is rapidly stimulated by radiation before transcription. A. Sequence of the wild- type (WT) and mutant (MT) miR-34 target element subcloned into the 3’UTR of the renilla luciferase mRNA in the psiCHECK2 dual luciferase reporter. B. Schematic diagram of how experiments are preformed. C. The level of let-7 and miR-34 gene silencing activity in A549, HeLa, MCF-7 and H460 cells, D. Simultaneous measurement of miR-34 activity and expression in A549 cells +/- ionizing radiation. miR-34 activity is regulated at the level of the mature microRNA Figure 2: Is IR-induced miR-34 activity Dicer dependent? 0 2 4 6 8 10 No siRNA siP53 siDROSHA siDICER siAGO2 siGAPD pri-microRNAexpression (foldchange) pri-miR-34a pri-miR-34b/c pri-miR-17 0 2 4 6 8 10 No siRNA siP53 siDROSHA siDICER siAGO2 siGAPD microRNAexpression (foldchange) miR-34a miR-34b miR-34c miR-17 0 5 10 15 20 No siRNA siP53 siDROSHA siDICER siAGO2 siGAPD miR-34activity -IR +IR Pre-treat cells: siRNA-P53 siRNA-Drosha siRNA-Dicer siRNA-Ago2 siRNA-GAPD Transfect with: WT and MT miR-34 reporter A. A549 -/+ IR pri-miR-34 RT-qPCR Norm. to B-Actin miR-34 RT-qPCR Norm. to U6 miR-34 activity western blot Norm. to Vimentin B. E. VIM Drosha P53 _ + _ + _ + _ + Vim Dicer Ago2 IR GAPD _ + _ + _ + IR IR-induced miR-34 activity is Drosha and Dicer-independent. A. Schematic diagram of the experiment. A549 cells pre-treated with siRNA for 36-hours were transfected with either the WT or MT miR-34 luciferase reporters. Cells were exposed to 4Gy of ionizing radiation. Following a 30- hour incubation the cells were lysed and assayed for pri-miR-34, mature miR-34, miR-34 activity and protein to validate siRNA knockdown. B. Relative pri-miR-34 and pri-miR-17 (negative control) expression in irradiated cells. C. Relative mature miR-34 or miR-17 (negative control) expression in irradiated cells. D. Relative miR-34 activity in irradiated cells. *Graphed for each experiment is the mean and standard deviation of 2-independent experiments preformed in triplicate. E. Protein knockdown was validated by western blot. C. D. Summary 1. There is a pre-existing pool of miR-34 in cells 2. miR-34 is stimulated by DNA damage in a Dicer-independent manor 3. The pre-existing pool of miR-34 is not loaded into Ago2 4. ATM is required for early activation of miR-34, whereas P53 is required for late activation 5. ATM is required for 5’-end phosphorylation of miR-34 6. We have discovered a novel mechanism for the regulation of microRNA activity 7. Our hypothesis is true: miR-34 is regulated at the mature level Figure 4: What DDR pathway activates miR-34? Pre-treat cells: siRNA-ATR siRNA-ATM siRNA-P53 Transfect with: WT and MT miR-34 reporter A549 -/+ IR miR-34 RT-qPCR Norm. to U6 miR-34 activity 0 1 2 3 4 5 6 7 no siRNA ATR siRNA ATM siRNA P53 siRNA RelativemiR-34activity 0hr 4hr 12hr 36hr 0 1 2 3 4 5 no siRNA ATR siRNA ATM siRNA P53 siRNA RelativemiR-34aexpression 0hr 4hr 12hr 36hr B.A. ATM is required for early miR-34 activation, whereas P53 is required for late miR-34 activation. A. Schematic diagram of the experiment. A549 cells pre-treated with siRNA for 36-hours were transfected with either the WT or MT miR-34 luciferase reporters. Cells were exposed to 4Gy of ionizing radiation. Following a 30-hour incubation the cells were lysed and assayed for mature miR-34 and miR-34 activity. B. miR-34 expression in irradiated cells treated with siRNA. C. Relative miR-34 activity in irradiated cells treated with siRNA. C. Figure 3: Is miR-34 loaded into Ago2? Measure miR-34 by RT-qPCR A549 FLAG/HA-EGFP FLAG/HA-AGO2 -/+ IR FLAGTotal A. miR-34 is loaded into Ago2 following IR. A. Schematic outline of the experiment. A549 cells stably expressing FLAG/HA-Ago2 or FLAG/HA-EGFP (negative control) were exposed to 4Gy of ionizing radiation. The cells were lysed at the indicated time. miR-34 expression was analyzed in the total extract or in FLAG immunoprecipitates. B. Relative miR-34 expression in total cell extract (top) and FLAG immunoprecipitates (bottom). B. Acknoledgements This work was funded by: NIH/NCI 5 K08 CA124484-05 and a Kalimeris Seed Grant. Figure 5: ATM is required for miR-34 5’-end phosphorylation miR-34 Northern blot A549 No siRNA siRNA-ATM -/+ IR Extract RNA -/+ CIP A. B. ATM is required for miR-34 5’-end phosphorylation. A. Schematic diagram of the experiment. A549 cells pre-treated with siRNA directed against ATM were exposed to 2Gy of ionizing radiation. Cells were lysed at the indicated time and total RNA was extracted. RNA was treated and untreated with CIP (to remove 5’-phosphates). miR-34 was analyzed by northern blot. B. Northern blot analysis of miR-34. 18S rRNA was used to normalize.
