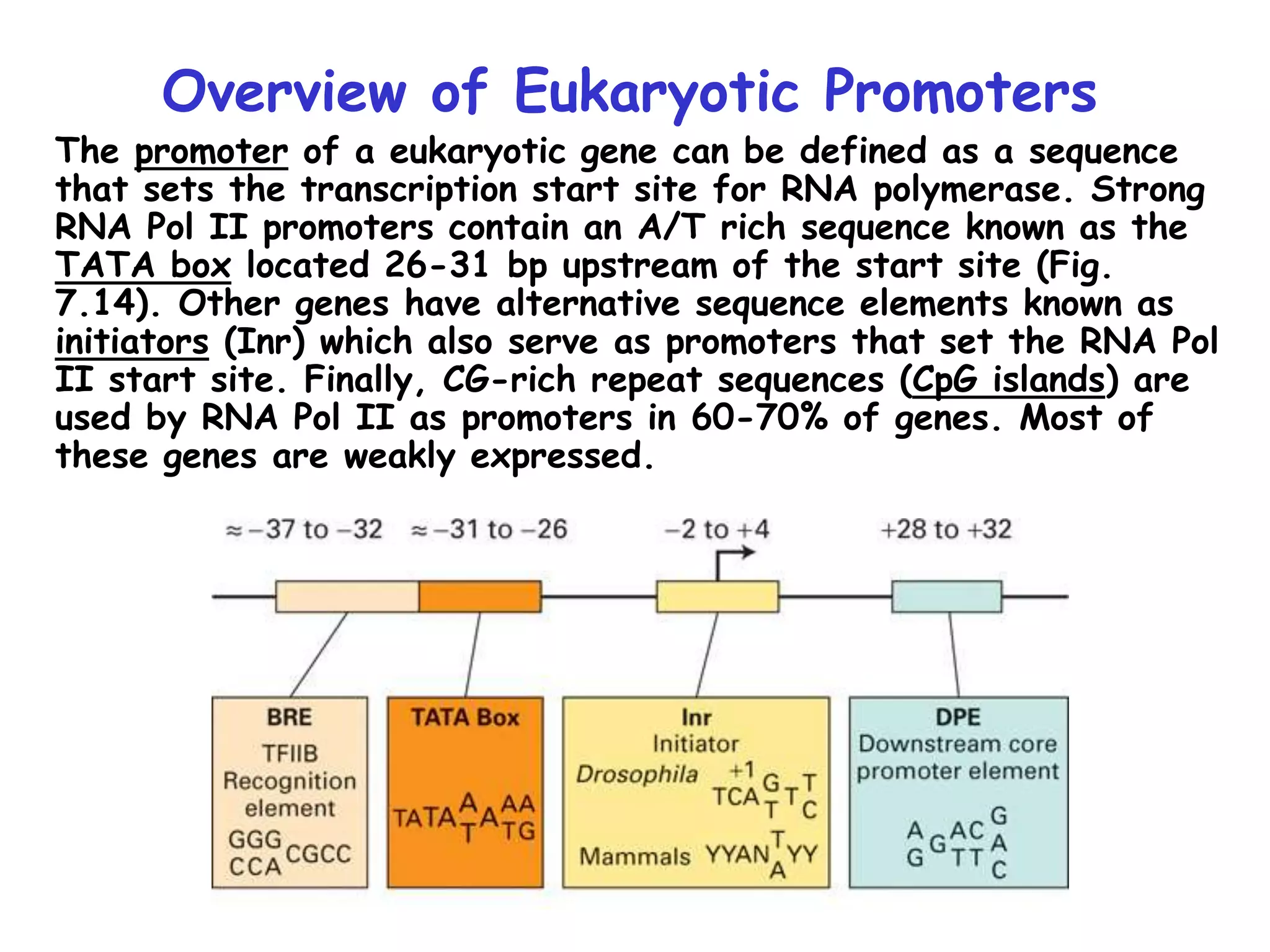
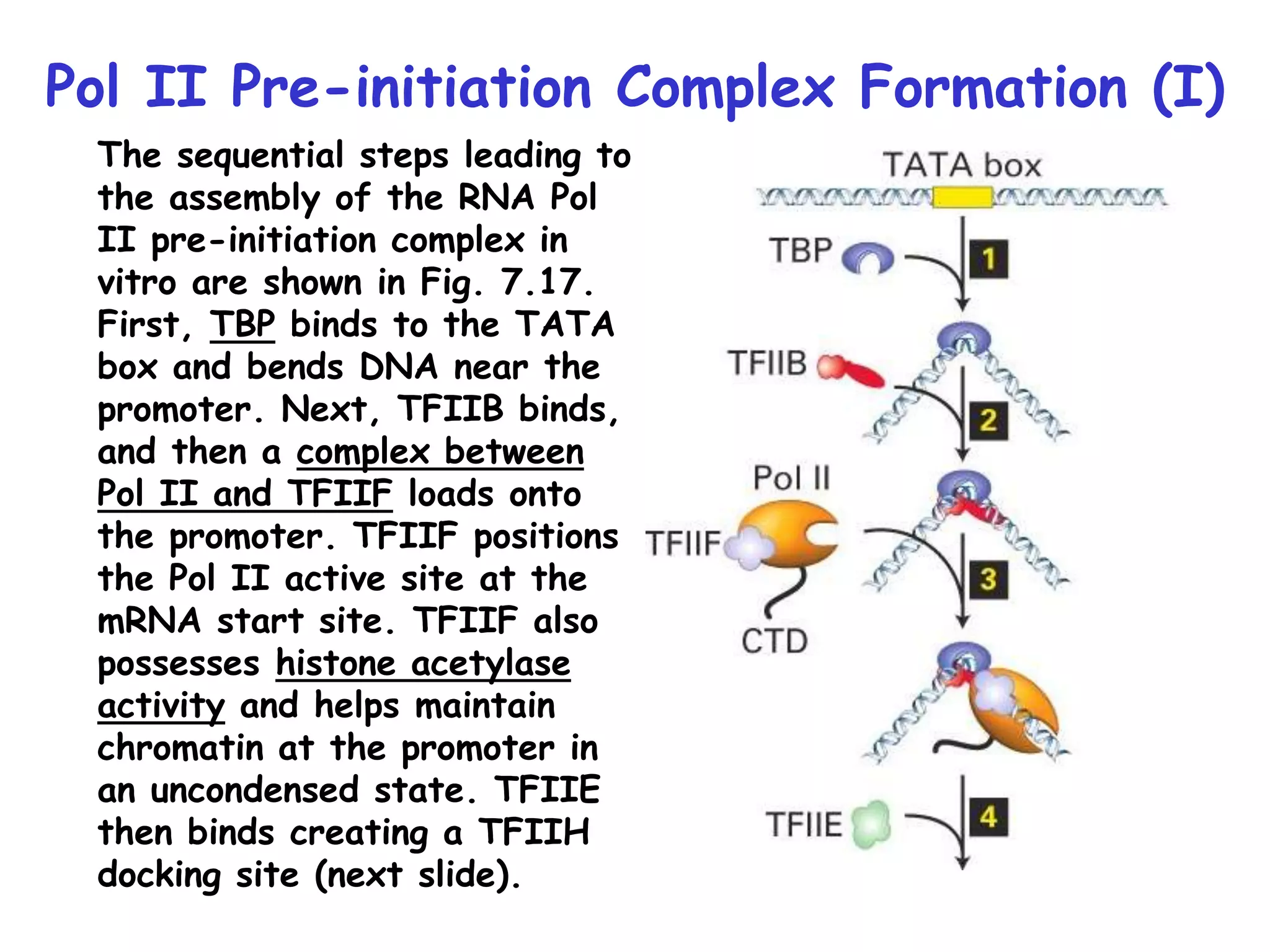
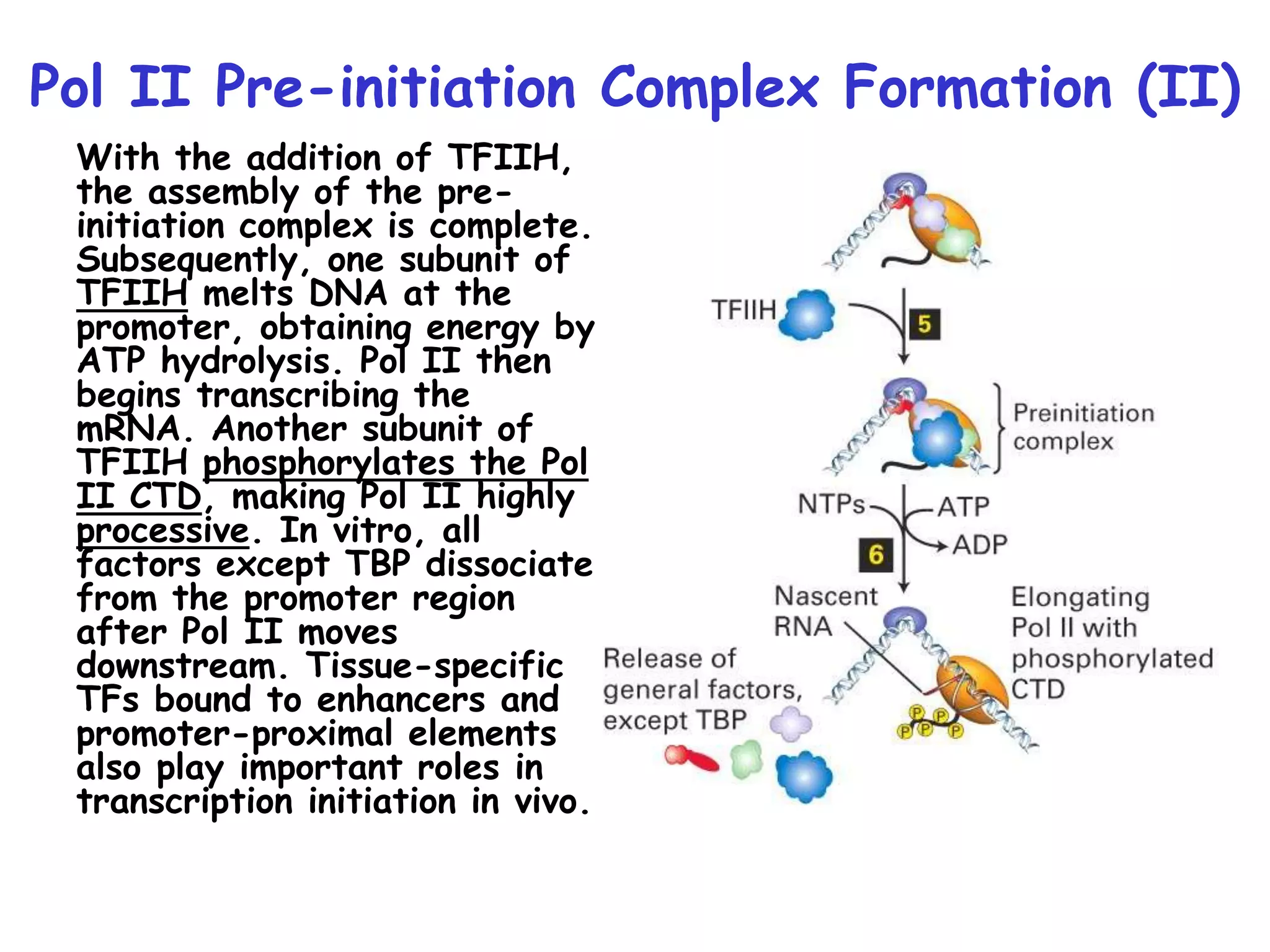
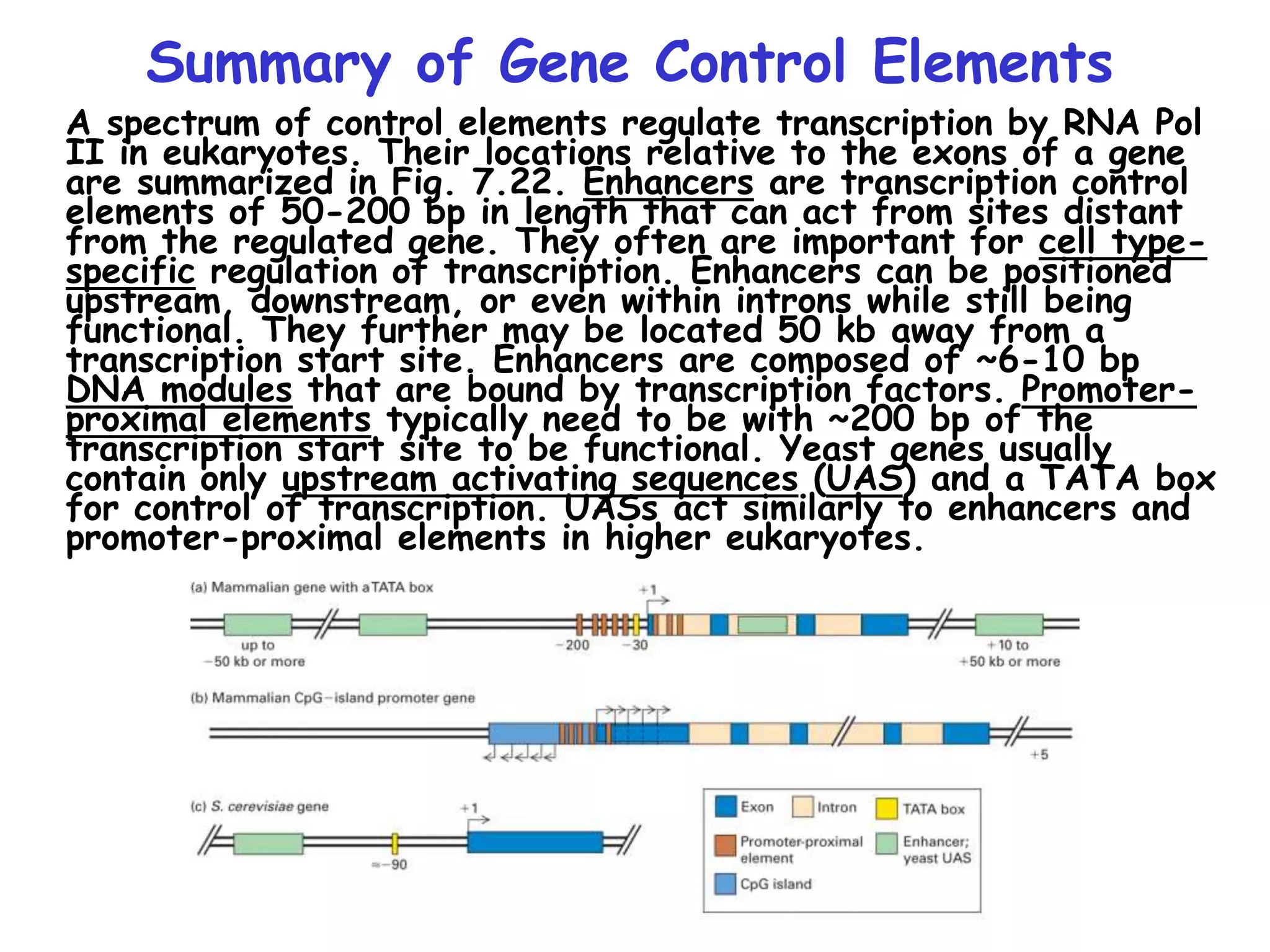
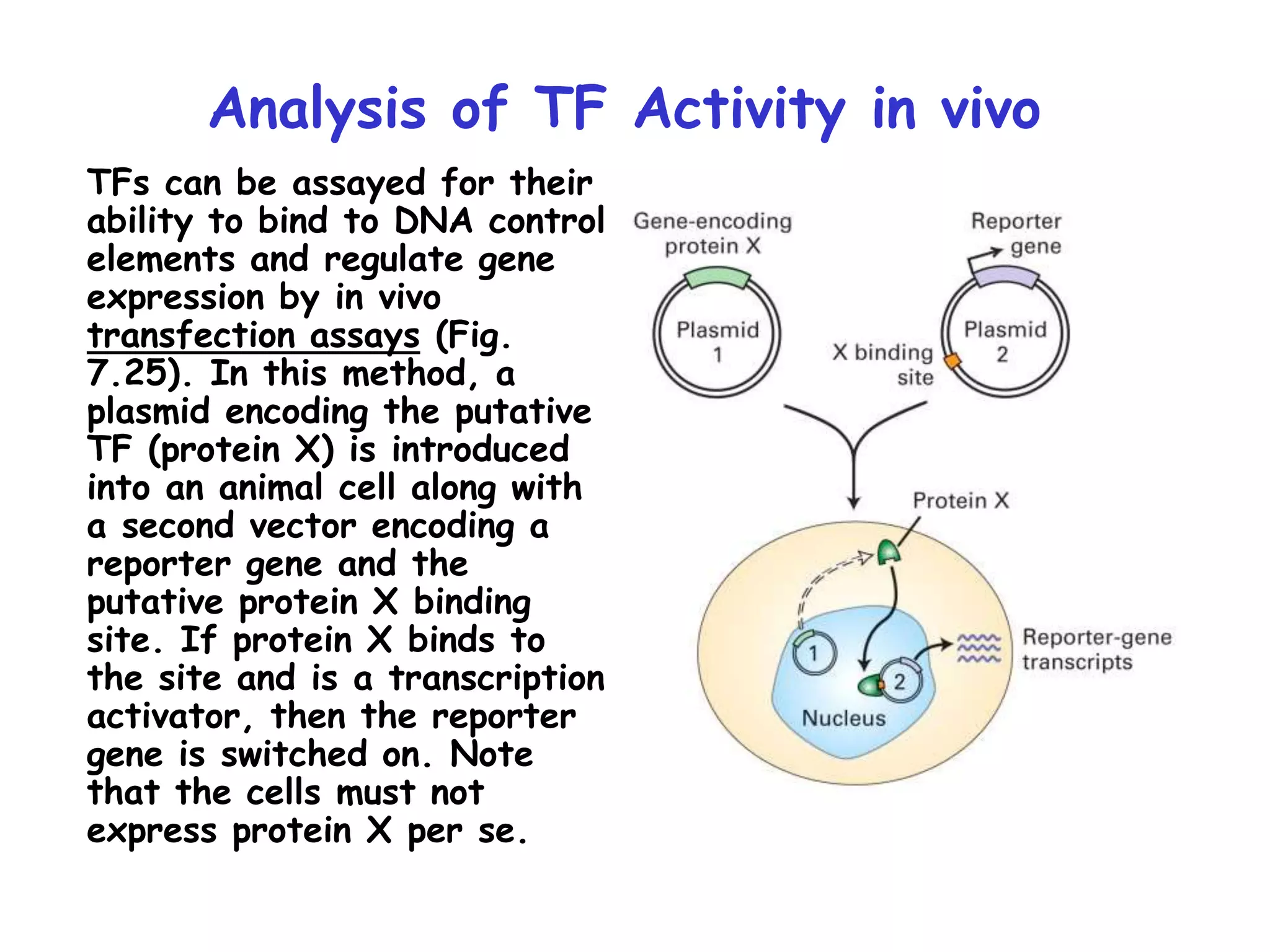
1) The document discusses the mechanisms of transcriptional control in eukaryotes. It describes the core promoter elements, general transcription factors, and how they assemble to form the pre-initiation complex at RNA polymerase II promoters.
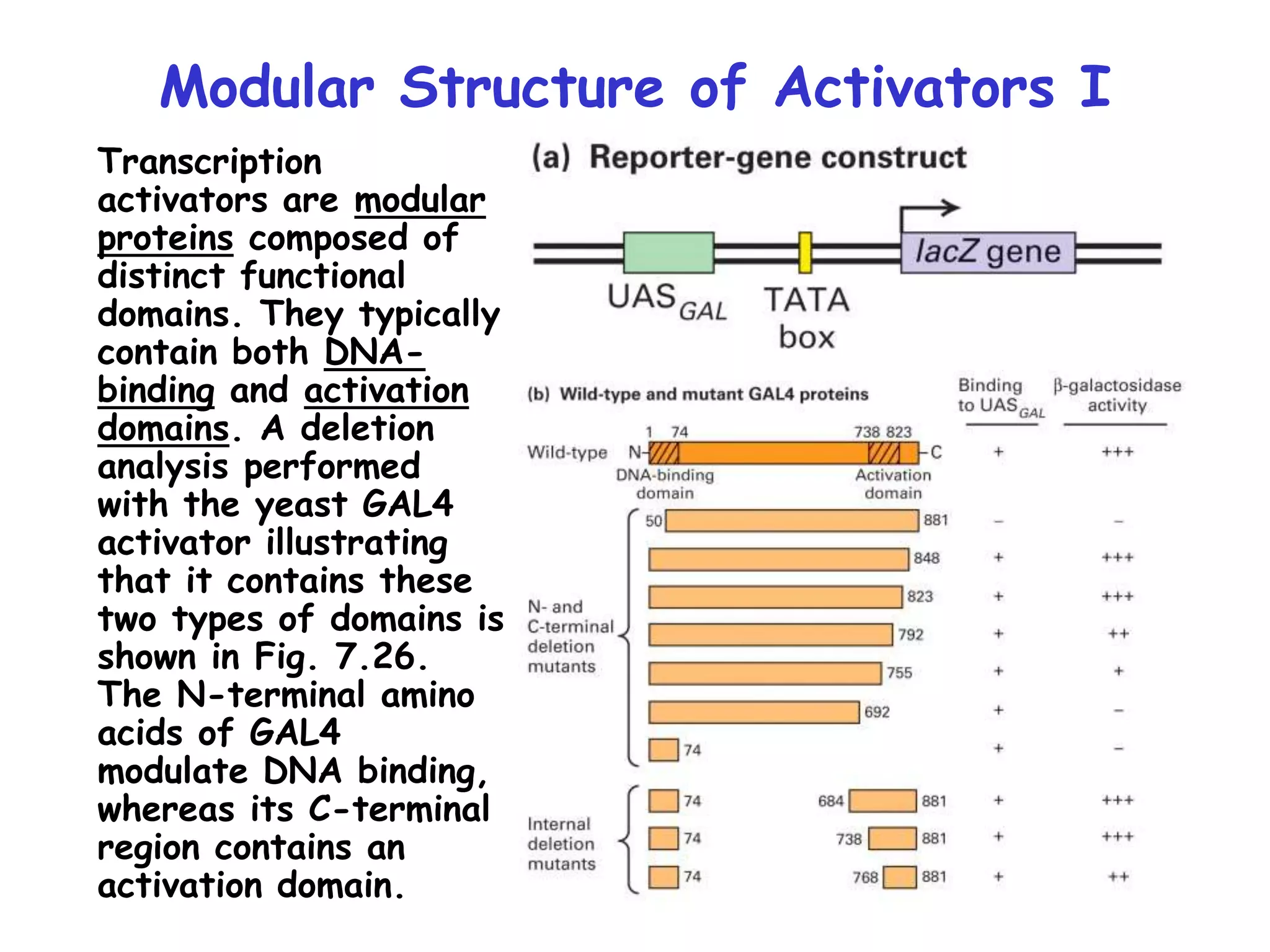
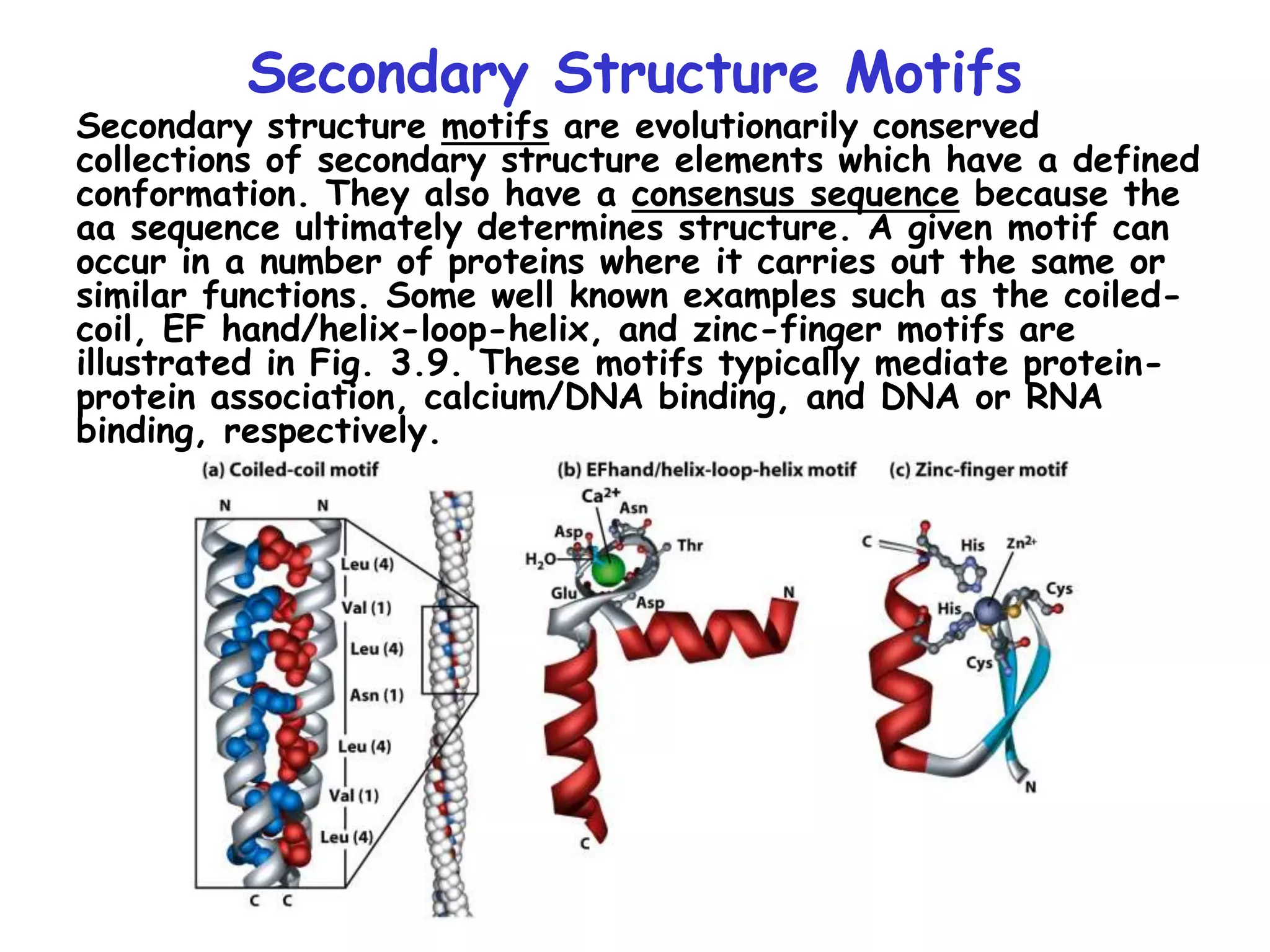
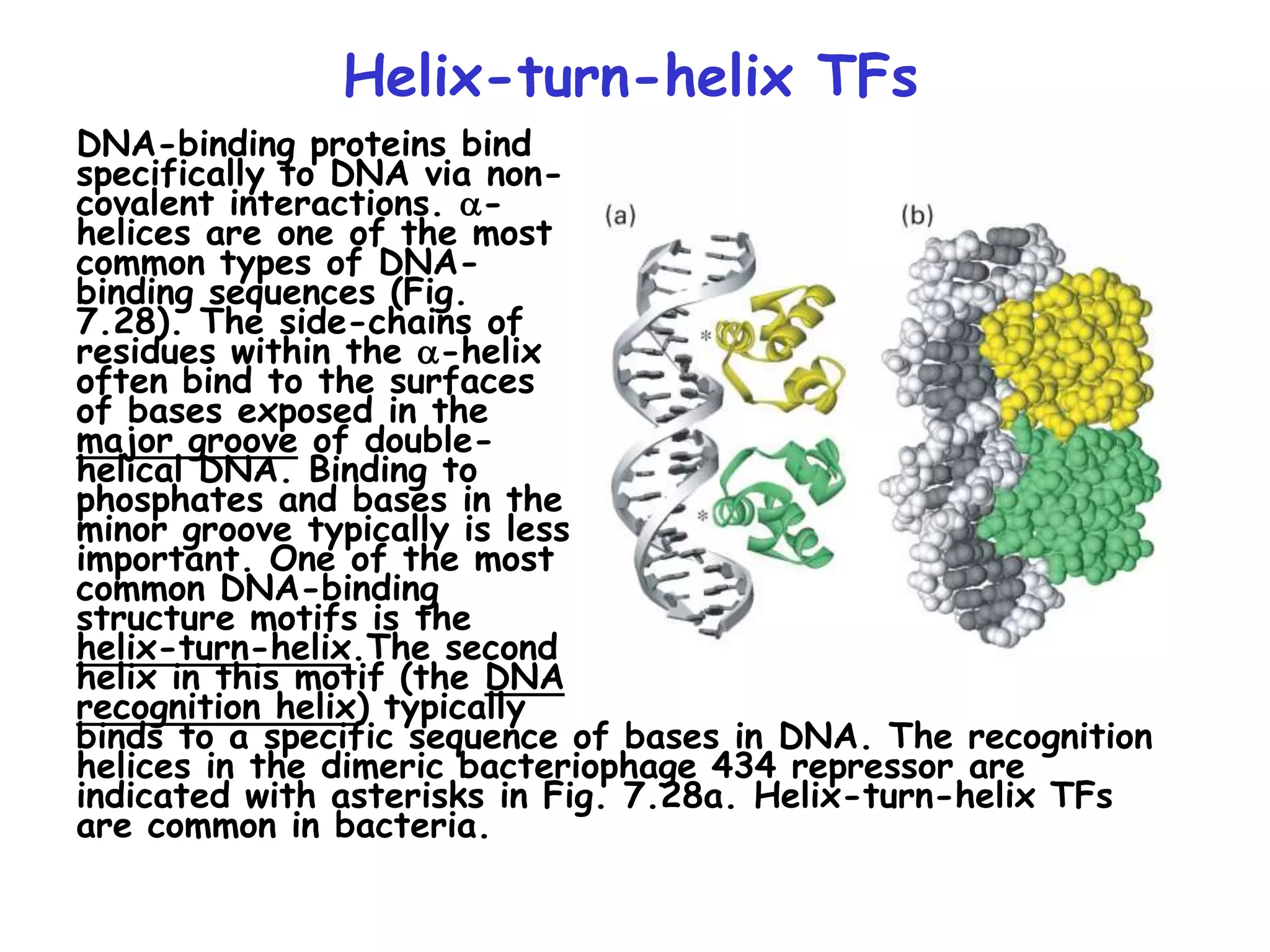
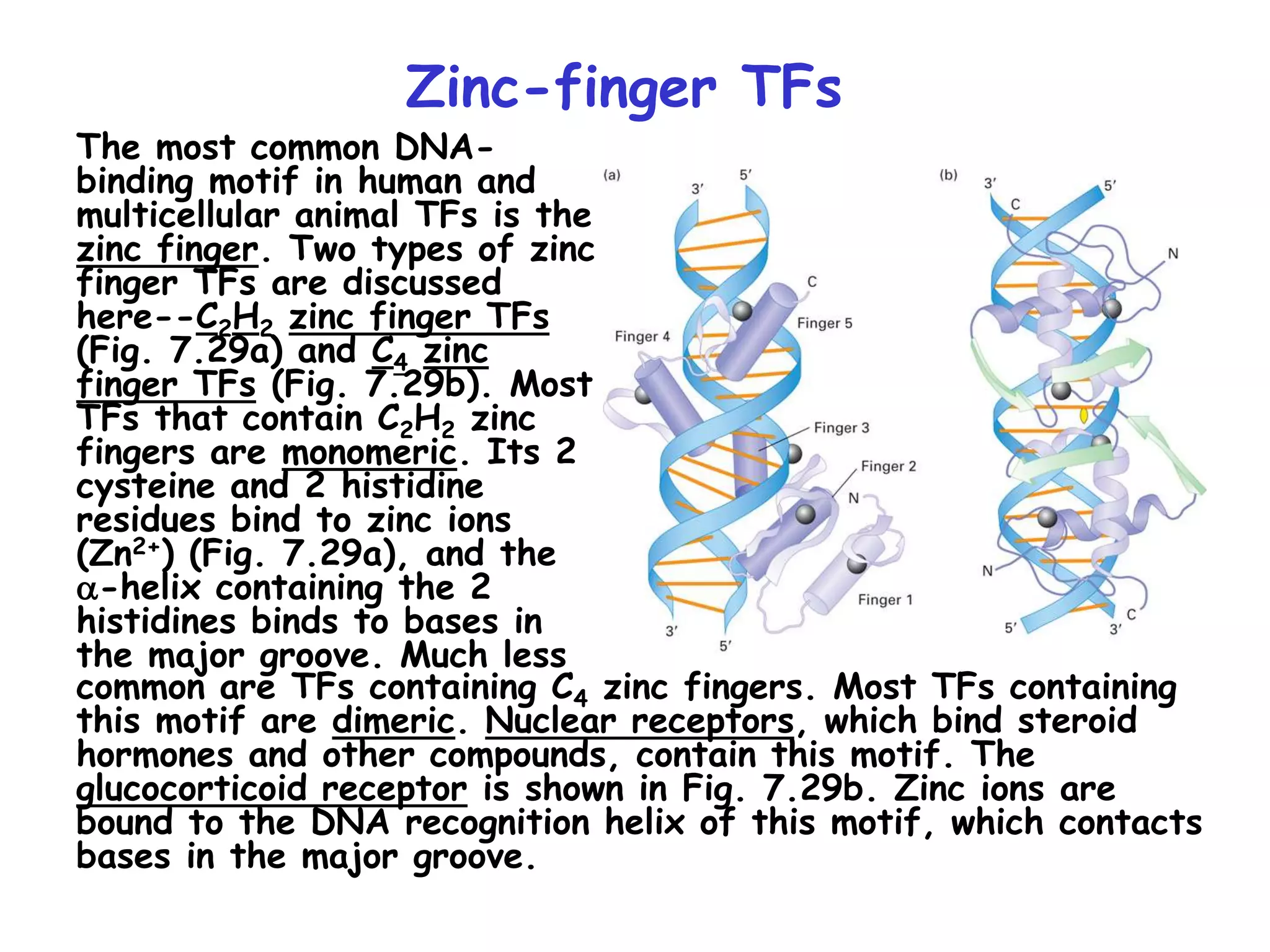
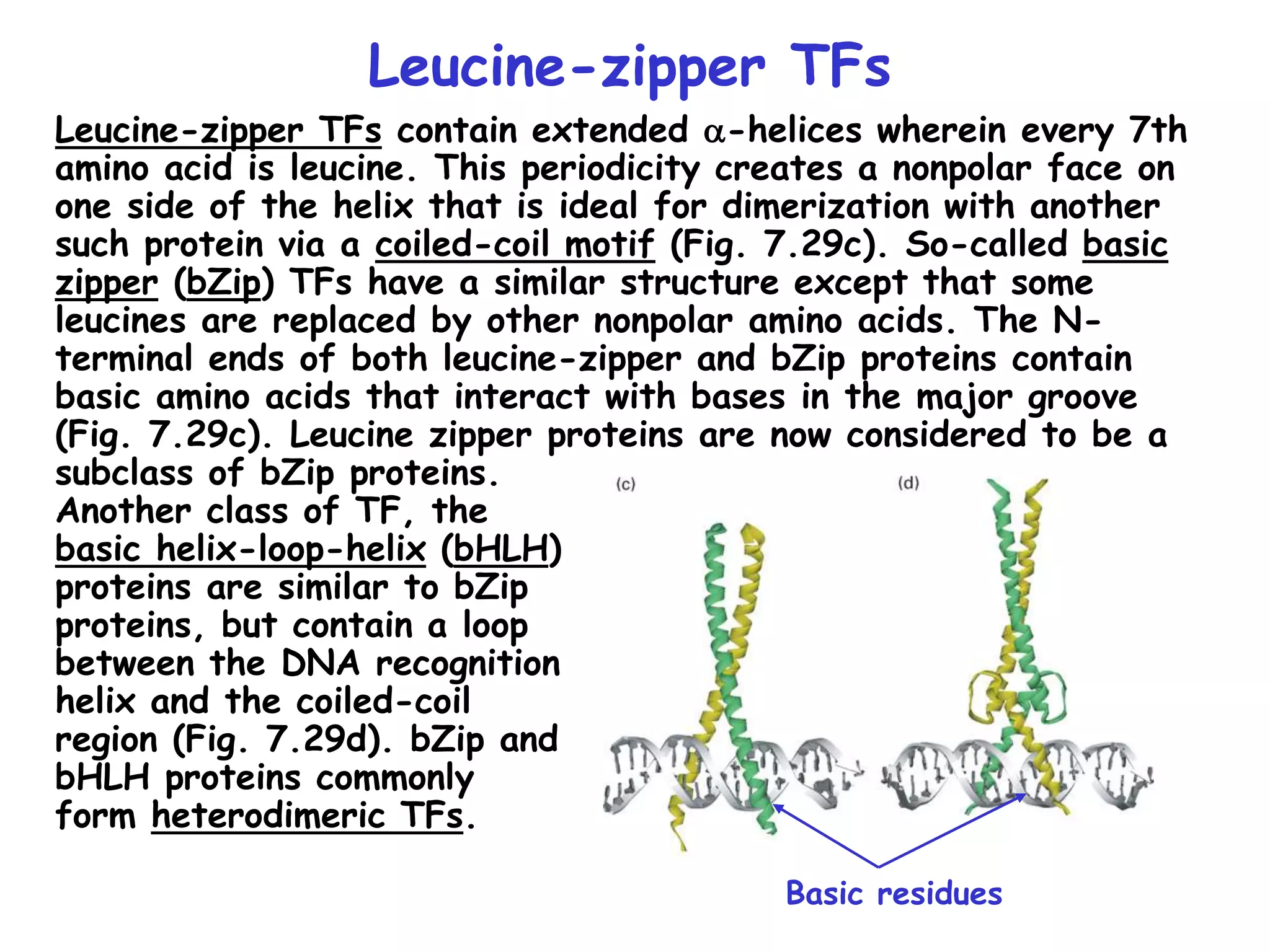
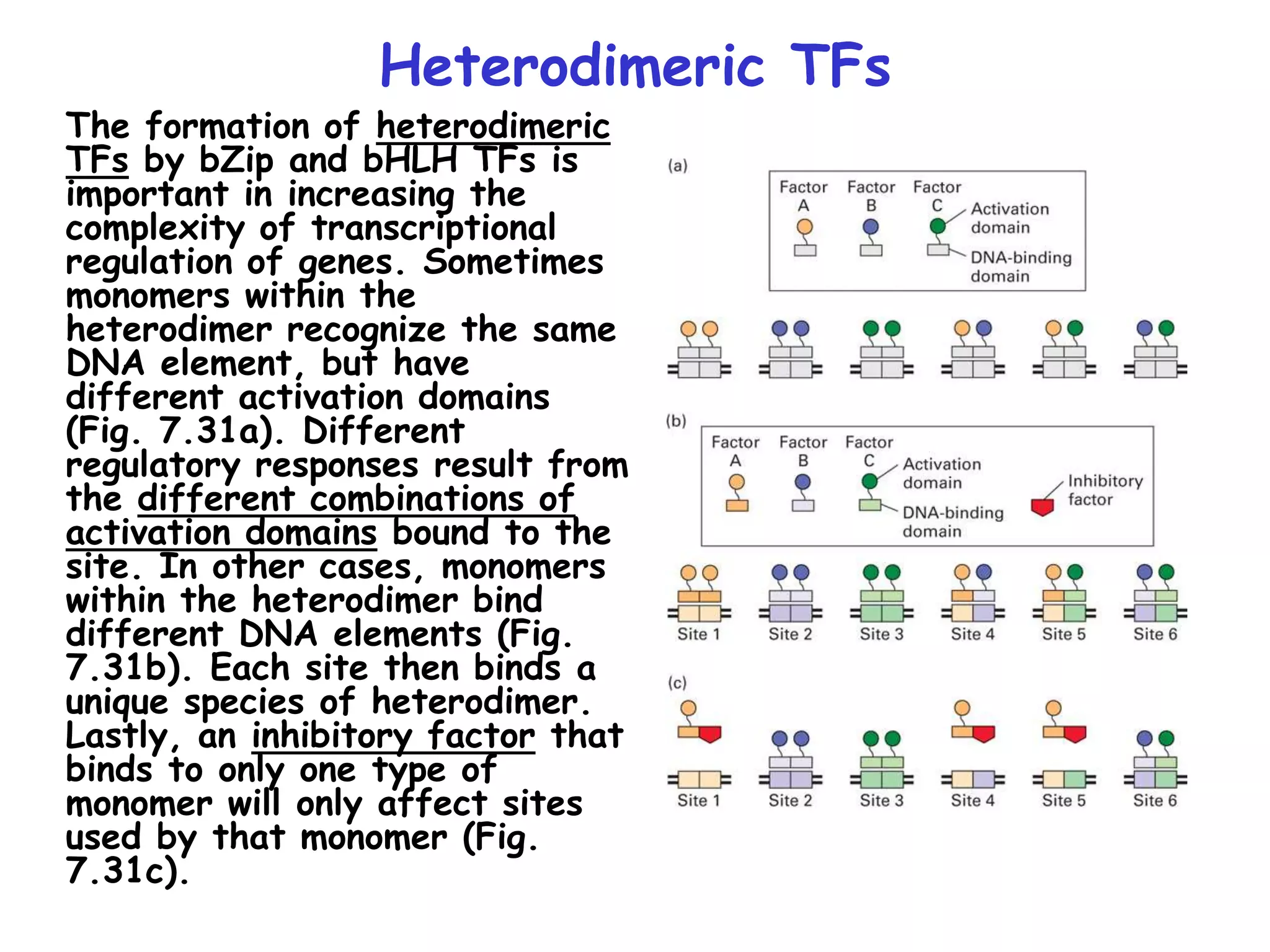
2) It also discusses the various classes of transcription factors that regulate gene expression, including their DNA-binding domains like zinc fingers, helix-turn-helices, and leucine zippers.
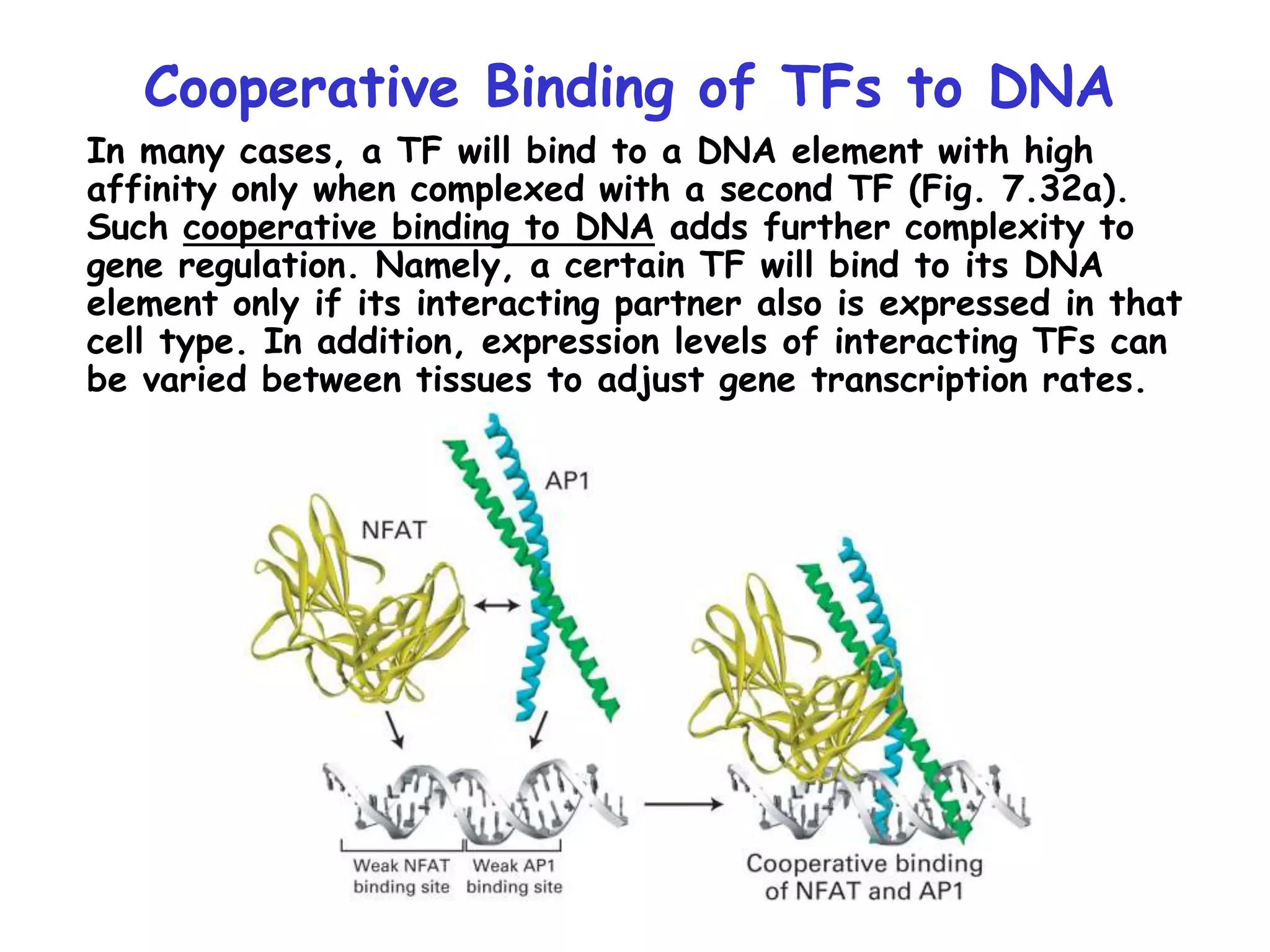
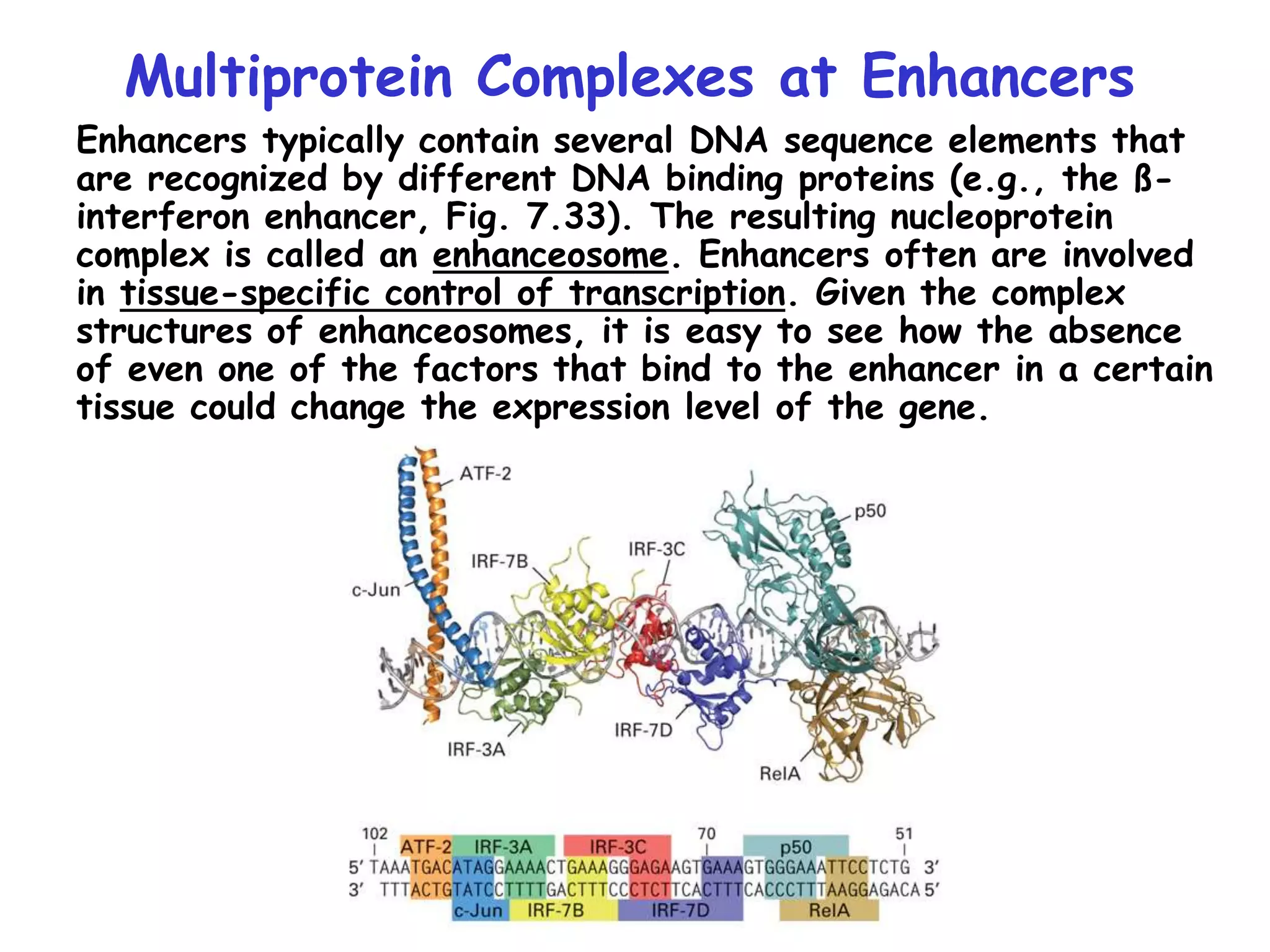
3) Transcription factor activity is regulated by ligands, co-factors, cooperative binding, and their assembly into enhanceosomes at gene enhancer elements.