Genetics
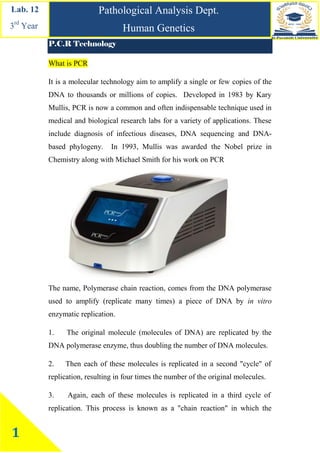
- 1. Pathological Analysis Dept. Human Genetics Lab. 12 3rd Year 1 P.C.R Technology What is PCR It is a molecular technology aim to amplify a single or few copies of the DNA to thousands or millions of copies. Developed in 1983 by Kary Mullis, PCR is now a common and often indispensable technique used in medical and biological research labs for a variety of applications. These include diagnosis of infectious diseases, DNA sequencing and DNA- based phylogeny. In 1993, Mullis was awarded the Nobel prize in Chemistry along with Michael Smith for his work on PCR PCR application The name, Polymerase chain reaction, comes from the DNA polymerase used to amplify (replicate many times) a piece of DNA by in vitro enzymatic replication. 1. The original molecule (molecules of DNA) are replicated by the DNA polymerase enzyme, thus doubling the number of DNA molecules. 2. Then each of these molecules is replicated in a second "cycle" of replication, resulting in four times the number of the original molecules. 3. Again, each of these molecules is replicated in a third cycle of replication. This process is known as a "chain reaction" in which the
- 2. Pathological Analysis Dept. Human Genetics Lab. 12 3rd Year 2 original DNA template is exponentially amplified with PCR it is possible to amplify a single piece of DNA, or a very small number of pieces of DNA, over many cycles, generating millions of copies of original DNA molecule Types of PCR • Conventional (Qualitative)PCR. • Multiplex PCR. • Nested PCR. • RT-PCR and qRT-PCR. • Quantitative PCR. • Hot-start PCR. • Touchdown PCR. • Assembly PCR. • Colony PCR. • Methylation-specific PCR. • LAMP assay. PCR requirement PCR requires several components and reagents. These components include: 1. DNA template that contains the DNA region (target) to the amplified. 2. One or more primers, which are complementary to the DNA regions at the 5 (five prime) and 3(three prime) ends of the DNA region. 3. DNA polymerase such as Taq polymerase or another DNA polymerase with a temperature optimum at around 70 C°.
- 3. Pathological Analysis Dept. Human Genetics Lab. 12 3rd Year 3 4. Deoxynucleoside triphosphates (dNTPs; also very commonly and erroneously called deoxynucleotide triphosphates), the building blocks from which the DNA polymerases synthesizes a new DNA strand. 5. Buffer solution, providing a suitable chemical environment for optimum activity and stability of the DNA polymerase. 6. Divalent cations, magnesium Mg+2 ion is used. 7. Monovalent cation potassium ions. Procedure The PCR usually consists of a series of 20 to 35 repeated temperature changes called cycles; each cycle typically consists of 2-3 discrete temperature steps. Most commonly PCR is carried out with cycles that have three temperature steps. The temperatures used and the length of time they are applied in each cycle depend on a variety of parameters. These include the enzyme used for DNA synthesis, the concentration of
- 4. Pathological Analysis Dept. Human Genetics Lab. 12 3rd Year 4 divalent ions and dNTPs in the reaction, and the melting temperature (Tm) of the primers. • Denaturation step: this step is the first regular cycling event and consists of heating the reaction to 94-98 C° for 20-30 seconds. It causes melting of DNA template and primers by disrupting the hydrogen bonds between complementary bases of the DNA strands, yielding single strands of DNA. • Annealing step: the reaction temperature is lowered to 50 -65 C° for 2040 seconds allowing annealing of the primers to the single - stranded DNA template. Typically the annealing temperature is about 3-5 degrees Celsius below the Tm of the primers used. Stable DNA-DNA hydrogen bonds are only formed when the primer sequence very closely matches the template sequence. The polymerase binds to the primer- template hybrid and begins DNA synthesis. • Extension / elongation step: the temperature at this step depends on the DNA polymerase used; Taq polymerase has its optimum activity temperature at 75-80 C°, and commonly a temperature of 72 C° is used with this enzyme. At this step the DNA polymerase synthesizes a new DNA strand complementary to the DNA template strands by adding dNTPs that are complementary to the template. The extension time depends both on the DNA polymerase used and on the length of the DNA fragment to be amplified. At its optimum temperature, the DNA polymerase will polymerize a thousand bases in one minute.
- 5. Pathological Analysis Dept. Human Genetics Lab. 12 3rd Year 5 • Final elongation: this single step is occasionally performed at a temperature of 70-74 C° for 5-15 minutes after the last PCR cycle to ensure that any remaining single - stranded DNA is fully extended. s Dr.Hayder Nasser
