Suraj_Jaladanki_PAX_Presence_Malaclemys_Terrapin_Poster
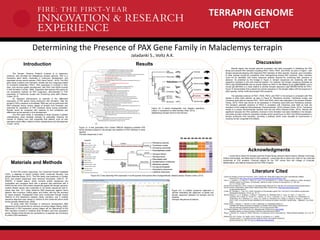
- 1. TERRAPIN GENOME PROJECT Introduction The Terrapin Genome Project’s purpose is to sequence, construct, and annotate the Malaclemys terrapin genome. PAX is a homeobox gene family, implicated in embryonic development, and expressed across animal species (“PAX Gene Family”, 2015). The PAX gene family contains nine members, divided into four subgroups based on functional similarities (“PAX”). PAX expression is involved in limb, brain, and nervous system development, with PAX1 and PAX9 involved in limb formation (LeClair, 1999). Organisms that express PAX genes do not necessarily express all PAX subgroups, demonstrated in PAX expression of Trachemys scripta and Chrysemys picta belili (Paixao- Cortes, 2013). An important phenomenon to examine is the differential expression of PAX genes during embryonic limb formation. After the terrapin’s PAX’s presence is annotated, RNA-seq can be performed with RNA extracted during various stages of terrapin limb formation and examined for expression of PAX members found during annotation. Results would be compared with relatives to find similarities and differences across species in limb development. The PAX gene family is compelling to study because a greater understanding could translate clinically by potentially reducing the number of children born with congenital limb defects, such as limb reductions which affect children’s future independence and development (“Facts”, 2015). Determining the Presence of PAX Gene Family in Malaclemys terrapin Centers for Disease Control and Prevention. (2014, October 28). Facts about Upper and Lower Limb Reduction Defects. Retrieved from http://www.cdc.gov/ncbddd/birthdefects/ul-limbreductiondefects .html. Field, D. J., Gauthier, J. A., King, B. L., Pisani, D., Lyson, T. R. and Peterson, K. J. (2014), Toward consilience in reptile phylogeny: miRNAs support an archosaur, not lepidosaur, affinity for turtles. Evolution & Development, 16: 189–196. doi: 10.1111/ede.12081 Genetics Home Reference. (2015, October 5). PAX gene family. Retrieved from http://ghr.nlm.nih.gov/geneFamily/pax#members. Human Genome Organization. Gene Family: Paired boxes (PAX). Retrieved from http://www.genenames.org/cgi- bin/genefamilies/set/675. LeClair, EE., Bonfiglio, L., Tuan, R.S. (1999). Expression of the pairedbox-genes Pax-1 and Pax-9 in limb skeleton development. Development Dynamics, 214 (2), pp. 101-15. Marchler-Bauer, A., Lu, S., Anderson J., Chitsaz, F., Derbyshire, M., DeWeese-Scott, C., Fong, J.H., Geer, L.Y., Geer, R.C., Gonzales, N.R., Gwadz, M., Hurwitz, D.I., Jackson, J.D., Ke, Z., Lanczycki, C.J., Zu, F., Marchler, G.H., Mullokandov, M., Omelchenko, M.V., Robertson, C.L., Song, J.S., Thanki, N., Yamashita, R.A., Zhang, D., Zhang, N., Zheng, C., Bryant, S. (2011). CDD: a Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Research, 3, pp. D225- D229. Paixao-Cortes, V. Saizano, F., Bortolini, M. (2013, September 2). Evolutionary History of Chordate PAX Genes: Dynamics of Change in a Complex Gene Family. Retrieved from http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0073560. The Reptile Database. (2015, November 23). Deirochelyinae. Retrieved from http://reptile- database.reptarium.cz/search?search=%09Deirochelyinae&submit=Search. Wang, Z., Gerstein, M., Snyder, M. (2009). RNA-Seq: a revolutionary tool for transcriptomics. Nature Reviews Genetics, 10 (1), pp. 57- 63. Waterhouse, A.M., Procter, J.B., Martin, D.M.A, Clamp, M. and Barton, G. J. (2009) "Jalview Version 2 - a multiple sequence alignment editor and analysis workbench" Bioinformatics25 (9) 1189-1191 doi: 10.1093/bioinformatics/btp033 Materials and Methods To find PAX protein sequences, the Conserved Domain Database (CDD), a database to search proteins within conserved domains, was utilized (Marchler-Bauer, 2011). The PAX family was examined in October 2015, and protein sequences were retrieved (Accession: cd00131, ID: 238076). A tblastn search, which converts protein sequences into nucleotides and compares them with a genome was performed with a FASTA format of the CDD protein sequences against the terrapin genome. Another tblastn search was conducted on all protein sequences with E- values less than 10-20 using reference RNA sequences against Homo sapiens, Mus musculus, Gallus gallus, and turtles, with top hits recorded for future analysis. Phylogenetic trees were constructed through Clustal OMEGA to find distinctions between family members, and a multiple sequence alignment was viewed in JalView to find conserved amino acids across species (Waterhouse, 2009). To study turtle limb formation in embryonic development, RNA sequencing will be performed on embryos at various stages (Wang, 2009). Differences in PAX expression among stages will be determined to find PAX members involved in vertebral body formation and their expression period. Western blots will also be conducted on a separate set of embryos to confirm PAX expression. Figure (I): A tree generated from Clustal OMEGA displaying potential PAX family members present in the terrapin and relations to PAX members in other species (Terrapin sequences in red) Results Jaladanki S., Voltz A.K. Literature Cited Discussion Results signify that terrapin genome annotation was fairly successful in identifying the PAX family and several PAX members including PAX1, PAX3, PAX7, and PAX9, as seen in Figure I, with terrapin sequences aligning with respective PAX members of other species. However, poor annotation of other species should be considered when distinguishing among PAX members. Other members could not be identified, with further sequence analysis required for confirming their presence or absence. As predicted by the lineage in Figure II, terrapin sequences are clustering with their respective members in bird and mammal species. For instance, the terrapin sequence predicted as PAX3 (utg7180000004508) has its closest relative as PAX3 in the Mallard Duck (gi880796890), and a mouse (gi12643442) is a close relative to another terrapin sequence (utg7180000010439) for PAX1. Figure IV demonstrates that a paired box domain sequence in the terrapin aligns with the sequence in other species, further confirming terrapin PAX presence. The potential evidence of PAX1, PAX9, PAX3, and PAX7 in the terrapin is consistent with PAX studies in other turtle species, as seen in Figure III. Pelodiscus sinensis, Trachemys scripta, and Chrsemys picta belili all express PAX1, PAX7 and PAX9, which was replicated in the terrapin (Paixao- Cortes, 2013). PAX3 was shown to be expressed in Chrsemys picta belili and Pelodiscus sinensis. The terrapin’s potential presence of PAX3 is consistent with Chrsemys picta belili as both are members of the subfamily Deirochelyinae (“Deirochelyinae”, 2015) (Paixao-Cortes, 2013). Trachemys scripta is a unique Deirochelyniae member that has lost PAX3 (Paixao-Cortes, 2013). Results are significant because the PAX subfamily involved in limb formation has been identified in the terrapin. Outcomes provide a basis for expression studies which would focus on PAX1 and PAX9 expression in terrapin embryonic limb formation, providing a pathway which could translate to improvement in reducing human congenital limb defects. Figure (III): A tree depicting PAX expression in turtle species and several other phylogenetically related species (Paixao-Cortes, 2013) Figure (II): A partial phylogenetic tree diagram specifying turtles’ in comparison to other families (Field, 2014) (Malaclemys terrapin found in the red box) Acknowledgments I would like to thank the Terrapin genome Project faculty advisors Steven Mount, Michael Cummings, Sridhar Hannenhalli, and Mihai Pop for their guidance. I would also like to thank Amy Voltz for her continued mentorship of TGP students. Financial support for the TGP comes from the College of Computer, Mathematical, and Natural Sciences and the FIRE Program. Figure (IV): A multiple sequence alignment in JalView illustrating the alignment of paired box domain sequence between the terrapin and its relatives (Terrapin Sequences at bottom)