Accelero bio founderswork_031717
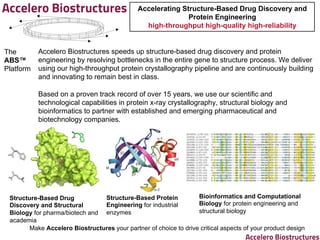
- 1. Accelerating Structure-Based Drug Discovery and Protein Engineering high-throughput high-quality high-reliability Accelero Biostructures speeds up structure-based drug discovery and protein engineering by resolving bottlenecks in the entire gene to structure process. We deliver using our high-throughput protein crystallography pipeline and are continuously building and innovating to remain best in class. Based on a proven track record of over 15 years, we use our scientific and technological capabilities in protein x-ray crystallography, structural biology and bioinformatics to partner with established and emerging pharmaceutical and biotechnology companies. The ABS™ Platform Structure-Based Drug Discovery and Structural Biology for pharma/biotech and academia Structure-Based Protein Engineering for industrial enzymes Bioinformatics and Computational Biology for protein engineering and structural biology Make Accelero Biostructures your partner of choice to drive critical aspects of your product design
- 2. • Application of DEN refinement and automated model building to a difficult case of molecular-replacement phasing: the structure of a putative succinyl-diaminopimelate desuccinylase from Corynebacterium glutamicum. Brunger AT, Das D, Deacon AM, Grant J, Terwilliger TC, Read RJ, Adams PD, Levitt M, Schröder GF. Acta Cryst D Biol Crystallogr. 2012 PMID: 22505259 • Improved molecular replacement by density- and energy-guided protein structure optimization. DiMaio F, Terwilliger TC, Read RJ, Wlodawer A, Oberdorfer G, Wagner U, Valkov E, Alon A, Fass D, Axelrod HL, Das D, Vorobiev SM, Iwaï H, Pokkuluri PR, Baker D. Nature. 2011 PMID: 21532589 • Distributed structure determination at the JCSG. van den Bedem H, Wolf G, Xu Q, Deacon AM. Acta Crystal D Biol Crystallogr. 2011 PMID: 21460455 • The JCSG high-throughput structural biology pipeline. Elsliger MA, Deacon AM, Godzik A, Lesley SA, Wooley J, Wüthrich K, Wilson IA. Acta Crystal Sect F Struct Biol Cryst Commun. 2010 S1744309110038212. PMID: 20944202 • Modeling discrete heterogeneity in X-ray diffraction data by fitting multi-conformers. van den Bedem H, Dhanik A, Latombe JC, Deacon AM. Acta Crystal D Biol Crystallogr. 2009 PMID: 19770508 Selected Methods Development work
- 3. • Three-dimensional structural view of the central metabolic network of Thermotoga maritima. Zhang Y, Thiele I, Weekes D, Li Z, Jaroszewski L, Ginalski K, Deacon AM, Wooley J, Lesley SA, Wilson IA, Palsson B, Osterman A, Godzik A. Science. 2009 PMID: 19762644 • Exploration of uncharted regions of the protein universe. Jaroszewski L, Li Z, Krishna SS, Bakolitsa C, Wooley J, Deacon AM, Wilson IA, Godzik A. PLoS Biol. 2009 PMID: 19787035 • Genome pool strategy for structural coverage of protein families. Jaroszewski L, Slabinski L, Wooley J, Deacon AM, Lesley SA, Wilson IA, Godzik A. Structure. 2008 PMID: 19000818 • An X-ray microsource based system for crystal screening and beamline development during synchrotron shutdown periods. Miller MD, Deacon AM. Nucl Instrum Methods Phys Res A. 2007 PMID: 19005562 • Structure-based inference of molecular functions of proteins of unknown function from Berkeley Structural Genomics Center. Shin DH, Hou J, Chandonia JM, Das D, Choi IG, Kim R, Kim SH. J Struct Funct Genomics. 2007 PMID: 17764033 Selected Methods Development work
- 4. • Structural genomics of minimal organisms and protein fold space. Kim SH, Shin DH, Liu J, Oganesyan V, Chen S, Xu QS, Kim JS, Das D, Schulze-Gahmen U, Holbrook SR, Holbrook EL, Martinez BA, Oganesyan N, DeGiovanni A, Lou Y, Henriquez M, Huang C, Jancarik J, Pufan R, Choi IG, Chandonia JM, Hou J, Gold B, Yokota H, Brenner SE, Adams PD, Kim R. J Struct Funct Genomics. 2005 PMID: 16211501 • Real-space protein-model completion: an inverse-kinematics approach. van den Bedem H, Lotan I, Latombe JC, Deacon AM. Acta Crystallogr D Biol Crystallogr. 2005 PMID: 15608370 • Shotgun crystallization strategy for structural genomics II: crystallization conditions that produce high resolution structures for T. maritima proteins. Page R, Deacon AM, Lesley SA, Stevens RC. J Struct Funct Genomics. 2005 PMID: 16211521 • A scaleable and integrated crystallization pipeline applied to mining the Thermotoga maritima proteome. DiDonato M, Deacon AM, Klock HE, McMullan D, Lesley SA. J Struct Funct Genomics. 2004 PMID: 15263852 Selected Methods Development work
- 5. • An automated system to mount cryo-cooled protein crystals on a synchrotron beam line, using compact sample cassettes and a small-scale robot. Cohen AE, Ellis PJ, Miller MD, Deacon AM, Phizackerley RP. J Appl Crystallogr. 2002 PMID: 24899734 • Blu-Ice and the Distributed Control System: software for data acquisition and instrument control at macromolecular crystallography beamlines. McPhillips TM, McPhillips SE, Chiu HJ, Cohen AE, Deacon AM, Ellis PJ, Garman E, Gonzalez A, Sauter NK, Phizackerley RP, Soltis SM, Kuhn P. J Synchrotron Radiat. 2002 PMID: 12409628 • Structural genomics of the Thermotoga maritima proteome implemented in a high- throughput structure determination pipeline. Lesley SA, Kuhn P, Godzik A, Deacon AM, Mathews I, Kreusch A, Spraggon G, Klock HE, McMullan D, Shin T, Vincent J, Robb A, Brinen LS, Miller MD, McPhillips TM, Miller MA, Scheibe D, Canaves JM, Guda C, Jaroszewski L, Selby TL, Elsliger MA, Wooley J, Taylor SS, Hodgson KO, Wilson IA, Schultz PG, Stevens RC. Proc Natl Acad Sci U S A. 2002 PMID: 12193646 • 3-Carboxy-cis,cis-muconate lactonizing enzyme from Neurospora crassa: MAD phasing with 80 selenomethionines. Merckel MC, Kajander T, Deacon AM, Thompson A, Grossmann JG, Kalkkinen N, Goldman A. Acta Cryst D Biol Crystallogr. 2002 PMID: 11976482 Selected Methods Development work
- 6. Selected Applica3ons work • A Distinct Type of Pilus from the Human Microbiome. Xu Q, Shoji M, Shibata S, Naito M, Sato K, Elsliger MA, Grant JC, Axelrod HL, Chiu HJ, Farr CL, Jaroszewski L, Knuth MW, Deacon AM, Godzik A, Lesley SA, Curtis MA, Nakayama K, Wilson IA. Cell. 2016 PMID: 27062925 • UHM-ULM interactions in the RBM39-U2AF65 splicing-factor complex. Stepanyuk GA, Serrano P, Peralta E, Farr CL, Axelrod HL, Geralt M, Das D, Chiu HJ, Jaroszewski L, Deacon AM, Lesley SA, Elsliger MA, Godzik A, Wilson IA, Wüthrich K, Salomon DR, Williamson JR. Acta Cryst D Struct Biol. 2016 PMID: 27050129 • PNT1 Is a C11 Cysteine Peptidase Essential for Replication of the Trypanosome Kinetoplast. Grewal JS, McLuskey K, Das D, Myburgh E, Wilkes J, Brown E, Lemgruber L, Gould MK, Burchmore RJ, Coombs GH, Schnaufer A, Mottram JC. J Biol Chem. 2016 PMID: 26940875 • Crystal Structure and Activity Studies of the C11 Cysteine Peptidase from Parabacteroides merdae in the Human Gut Microbiome. McLuskey K, Grewal JS, Das D, Godzik A, Lesley SA, Deacon AM, Coombs GH, Elsliger MA, Wilson IA, Mottram JC. J Biol Chem. 2016 PMID: 26940874 • Structure of Liver Receptor Homolog-1 (NR5A2) with PIP3 hormone bound in the ligand binding pocket. Sablin EP, Blind RD, Uthayaruban R, Chiu HJ, Deacon AM, Das D, Ingraham HA, Fletterick RJ. J Struct Biol. 2015 PMID: 26416531
- 7. • Structure-based discovery of NANOG variant with enhanced properties to promote self- renewal and reprogramming of pluripotent stem cells. Hayashi Y, Caboni L, Das D, Yumoto F, Clayton T, Deller MC, Nguyen P, Farr CL, Chiu HJ, Miller MD, Elsliger MA, Deacon AM, Godzik A, Lesley SA, Tomoda K, Conklin BR, Wilson IA, Yamanaka S, Fletterick RJ. Proc Natl Acad Sci U S A. 2015 PMID: 25825768 • The signaling phospholipid PIP3 creates a new interaction surface on the nuclear receptor SF-1. Blind RD, Sablin EP, Kuchenbecker KM, Chiu HJ, Deacon AM, Das D, Fletterick RJ, Ingraham HA. Proc Natl Acad Sci U S A. 2014 PMID: 25288771 • Cell fate regulation governed by a repurposed bacterial histidine kinase. Childers WS, Xu Q, Mann TH, Mathews II, Blair JA, Deacon AM, Shapiro L. PLoS Biol. 2014 PMID: 25349992 • Molecular characterization of novel pyridoxal-5'-phosphate-dependent enzymes from the human microbiome. Fleischman NM, Das D, Kumar A, Xu Q, Chiu HJ, Jaroszewski L, Knuth MW, Klock HE, Miller MD, Elsliger MA, Godzik A, Lesley SA, Deacon AM, Wilson IA, Toney MD. Protein Sci. 2014 PMID: 24888348 • Structure and computational analysis of a novel protein with metallopeptidase-like and circularly permuted winged-helix-turn-helix domains reveals a possible role in modified polysaccharide biosynthesis. Das D, Murzin AG, Rawlings ND, Finn RD, Coggill P, Bateman A, Godzik A, Aravind L. BMC Bioinformatics. 2014 PMID: 24646163 Selected Applica3ons work
- 8. • Crystal structure of a putative quorum sensing-regulated protein (PA3611) from the Pseudomonas-specific DUF4146 family. Das D, Chiu HJ, Farr CL, Grant JC, Jaroszewski L, Knuth MW, Miller MD, Tien HJ, Elsliger MA, Deacon AM, Godzik A, Lesley SA, Wilson IA. Proteins. 2014 PMID: 24174223 • Structure and function of a novel LD-carboxypeptidase a involved in peptidoglycan recycling. Das D, Hervé M, Elsliger MA, Kadam RU, Grant JC, Chiu HJ, Knuth MW, Klock HE, Miller MD, Godzik A, Lesley SA, Deacon AM, Mengin-Lecreulx D, Wilson IA. J Bacteriol. 2013 PMID: 24123814 • Structure and function of the DUF2233 domain in bacteria and in the human mannose 6- phosphate uncovering enzyme. Das D, Lee WS, Grant JC, Chiu HJ, Farr CL, Vance J, Klock HE, Knuth MW, Miller MD, Elsliger MA, Deacon AM, Godzik A, Lesley SA, Kornfeld S, Wilson IA. J Biol Chem. 2013 PMID: 23572527 • Structure and function of the first full-length murein peptide ligase (Mpl) cell wall recycling protein. Das D, Hervé M, Feuerhelm J, Farr CL, Chiu HJ, Elsliger MA, Knuth MW, Klock HE, Miller MD, Godzik A, Lesley SA, Deacon AM, Mengin-Lecreulx D, Wilson IA. PLoS One. 2011 PMID: 21445265 Selected Applica3ons work
- 9. • Crystal structure of the first eubacterial Mre11 nuclease reveals novel features that may discriminate substrates during DNA repair. Das D, Moiani D, Axelrod HL, Miller MD, McMullan D, Jin KK, Abdubek P, Astakhova T, Burra P, Carlton D, Chiu HJ, Clayton T, Deller MC, Duan L, Ernst D, Feuerhelm J, Grant JC, Grzechnik A, Grzechnik SK, Han GW, Jaroszewski L, Klock HE, Knuth MW, Kozbial P, Krishna SS, Kumar A, Marciano D, Morse AT, Nigoghossian E, Okach L, Paulsen J, Reyes R, Rife CL, Sefcovic N, Tien HJ, Trame CB, van den Bedem H, Weekes D, Xu Q, Hodgson KO, Wooley J, Elsliger MA, Deacon AM, Godzik A, Lesley SA, Tainer JA, Wilson IA. J Mol Biol. 2010 PMID: 20122942 • Crystal structure of the multidrug efflux transporter AcrB at 3.1A resolution reveals the N- terminal region with conserved amino acids. Das D, Xu QS, Lee JY, Ankoudinova I, Huang C, Lou Y, DeGiovanni A, Kim R, Kim SH. J Struct Biol. 2007 PMID: 17275331 • The crystal structure of the monomeric reverse transcriptase from Moloney murine leukemia virus. Das D, Georgiadis MM. Structure. 2004 PMID: 15130474 • Functional analysis of substrate and cofactor complex structures of a thymidylate synthase-complementing protein. Mathews II, Deacon AM, Canaves JM, McMullan D, Lesley SA, Agarwalla S, Kuhn P. Structure. 2003 PMID: 12791256 Selected Applica3ons work
- 10. Debanu Das, PhD, Co-‐founder & CEO Debanu has 15 years of experience in protein x-ray crystallography and structural biology. His expertise includes structure-function analysis of a broad range of proteins involved in cell development, cell wall recycling, DNA repair, human microbiome, cell signaling, viral replication and drug transport. Prior to Accelero Biostructures, he served as Staff Scientist in the Structural Genomics Division at SSRL (Stanford Synchrotron Radiation Lightsource) at SLAC National Accelerator Laboratory, Menlo Park. There he determined crystal structures of several hundred novel proteins as part of the JCSG and established multiple productive partnerships with other renowned research groups for applying the results. He received his postdoctoral training at Lawrence Berkeley National Laboratory and University of California Berkeley on structural genomics and membrane protein crystallography with Sung-Hou Kim, a pioneer of structural genomics and founder of the drug discovery company Plexxikon. As an industry expert, Debanu also consults with startups on structure-based drug discovery and business development. He holds an Integrated MS in Science and Engineering from the Indian Institute of Technology Kanpur and received his Ph.D. from Rutgers, NJ. He has also taken professional development courses in business and law in the Continuing Studies Program at Stanford University. . Ashley Deacon, PhD, Co-‐founder & CSO Ashley has 25 years of experience in synchrotron-based protein crystallography research. His professional experience and passion lies in the development and application of new technology to high impact structural biology research. Prior to founding Accelero Biostructures, Ashley held multiple positions at SSRL at SLAC National Accelerator Laboratory, Menlo Park, including Staff Scientist, Senior Staff Scientist, and Group Leader of the Structural Genomics Division and JCSG Structure Determination Core. Ashley helped develop a synchrotron-integrated platform for high quality structure determination which JCSG applied to a broad array of important biological problems, including the de novo structure determination of hundreds of proteins from the human microbiome, human stem cell and T-cell biology and the prokaryotic cell cycle. This led to the determination of about 1500 novel 3D structures, including several protein-protein and protein-nucleotide complexes and numerous protein-ligand complexes. During postdoctoral training at Cornell University, Ashley worked closely with Nobel Laureate Herbert Hauptman to apply “direct methods” of structure determination to a broad class of biological problems using synchrotron-based multi-wavelength anomalous dispersion and ultra-high resolution diffraction data. Ashley holds a Ph.D. from the University of Manchester, UK, and is an alumni of the Stanford Manager Academy at Stanford University. www.accelerobio.com info@accelerobio.com Team
