phylogenetic tree.pptx
•Download as PPTX, PDF•
0 likes•70 views
phylogenetic tree
Report
Share
Report
Share
Recommended
More Related Content
What's hot
What's hot (7)
An information retrieval training tool targeting the PBL students at the Univ...

An information retrieval training tool targeting the PBL students at the Univ...
Looking at chemistry - protein - papers connectivity in ELIXIR

Looking at chemistry - protein - papers connectivity in ELIXIR
A network framework to explore phylogenetic structure in genomic data

A network framework to explore phylogenetic structure in genomic data
TreeScaper: Software to Visualize and Extract Phylogenetic Signals from Sets ...

TreeScaper: Software to Visualize and Extract Phylogenetic Signals from Sets ...
Linkages to EHRs and Related Standards. What can we learn from the Parallel U...

Linkages to EHRs and Related Standards. What can we learn from the Parallel U...
Similar to phylogenetic tree.pptx
Similar to phylogenetic tree.pptx (20)
MIB200A at UCDavis Module: Microbial Phylogeny; Class 2

MIB200A at UCDavis Module: Microbial Phylogeny; Class 2
Basic concepts in systamatics,taxonomy and phylogenetic tree

Basic concepts in systamatics,taxonomy and phylogenetic tree
Recently uploaded
TỔNG ÔN TẬP THI VÀO LỚP 10 MÔN TIẾNG ANH NĂM HỌC 2023 - 2024 CÓ ĐÁP ÁN (NGỮ Â...

TỔNG ÔN TẬP THI VÀO LỚP 10 MÔN TIẾNG ANH NĂM HỌC 2023 - 2024 CÓ ĐÁP ÁN (NGỮ Â...Nguyen Thanh Tu Collection
Mehran University Newsletter Vol-X, Issue-I, 2024

Mehran University Newsletter Vol-X, Issue-I, 2024Mehran University of Engineering & Technology, Jamshoro
Recently uploaded (20)
Unit 3 Emotional Intelligence and Spiritual Intelligence.pdf

Unit 3 Emotional Intelligence and Spiritual Intelligence.pdf
ICT Role in 21st Century Education & its Challenges.pptx

ICT Role in 21st Century Education & its Challenges.pptx
NO1 Top Black Magic Specialist In Lahore Black magic In Pakistan Kala Ilam Ex...

NO1 Top Black Magic Specialist In Lahore Black magic In Pakistan Kala Ilam Ex...
Jual Obat Aborsi Hongkong ( Asli No.1 ) 085657271886 Obat Penggugur Kandungan...

Jual Obat Aborsi Hongkong ( Asli No.1 ) 085657271886 Obat Penggugur Kandungan...
TỔNG ÔN TẬP THI VÀO LỚP 10 MÔN TIẾNG ANH NĂM HỌC 2023 - 2024 CÓ ĐÁP ÁN (NGỮ Â...

TỔNG ÔN TẬP THI VÀO LỚP 10 MÔN TIẾNG ANH NĂM HỌC 2023 - 2024 CÓ ĐÁP ÁN (NGỮ Â...
HMCS Max Bernays Pre-Deployment Brief (May 2024).pptx

HMCS Max Bernays Pre-Deployment Brief (May 2024).pptx
Exploring_the_Narrative_Style_of_Amitav_Ghoshs_Gun_Island.pptx

Exploring_the_Narrative_Style_of_Amitav_Ghoshs_Gun_Island.pptx
General Principles of Intellectual Property: Concepts of Intellectual Proper...

General Principles of Intellectual Property: Concepts of Intellectual Proper...
Fostering Friendships - Enhancing Social Bonds in the Classroom

Fostering Friendships - Enhancing Social Bonds in the Classroom
On National Teacher Day, meet the 2024-25 Kenan Fellows

On National Teacher Day, meet the 2024-25 Kenan Fellows
Sensory_Experience_and_Emotional_Resonance_in_Gabriel_Okaras_The_Piano_and_Th...

Sensory_Experience_and_Emotional_Resonance_in_Gabriel_Okaras_The_Piano_and_Th...
Food safety_Challenges food safety laboratories_.pdf

Food safety_Challenges food safety laboratories_.pdf
phylogenetic tree.pptx
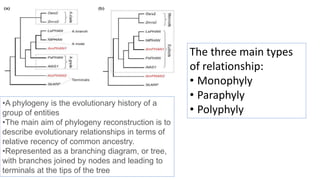
- 1. •A phylogeny is the evolutionary history of a group of entities •The main aim of phylogeny reconstruction is to describe evolutionary relationships in terms of relative recency of common ancestry. •Represented as a branching diagram, or tree, with branches joined by nodes and leading to terminals at the tips of the tree The three main types of relationship: • Monophyly • Paraphyly • Polyphyly
- 2. Overview of phylogenetic analysis Choosing a study group • Survey the literature pertaining to the group of interest. Sampling the group of interest • 4 sources of data for reconstructing the phylogeny of a gene family: published sequences of characterized, gene databases, EST project databases, and unpublished data from colleagues. Sequence retrieval • Sequences retrieved from BLAST searches varies depending on the size of the gene family, and what is chosen for inclusion in further analyses will vary accordingly. • When sequences are retrieved from BLAST searches they are allocated an e- score, which is an indication of the degree of similarity between the initial sequence used for searches and the sequence retrieved. Amino acid or nucleotide data analysis • Data as amino lebih banyak kemungkinan keadaan karakter dibandingkan dengan nukleotida (20:4). • Alignment as amino lbh mudah Alignment • Phylogenetic methodology relies on the assumption that the characters used to generate trees are homologous.
- 4. Interpreting a phylogenetic tree • Look at the strict consensus parsimony tree. A totally unresolved tree (the branches all come out from the same node) may indicate a rapid radiation, insufficient phylogenetic information in the original alignment or incongruence between tree topologies. • Compare the strict consensus parsimony and ML trees to see if they are congruent • Look at the relationships between the terminals of interest and see how they answer the biological question that was originally posed in terms of orthology and paralogy, and in terms of monophyly, paraphyly and polyphyly.