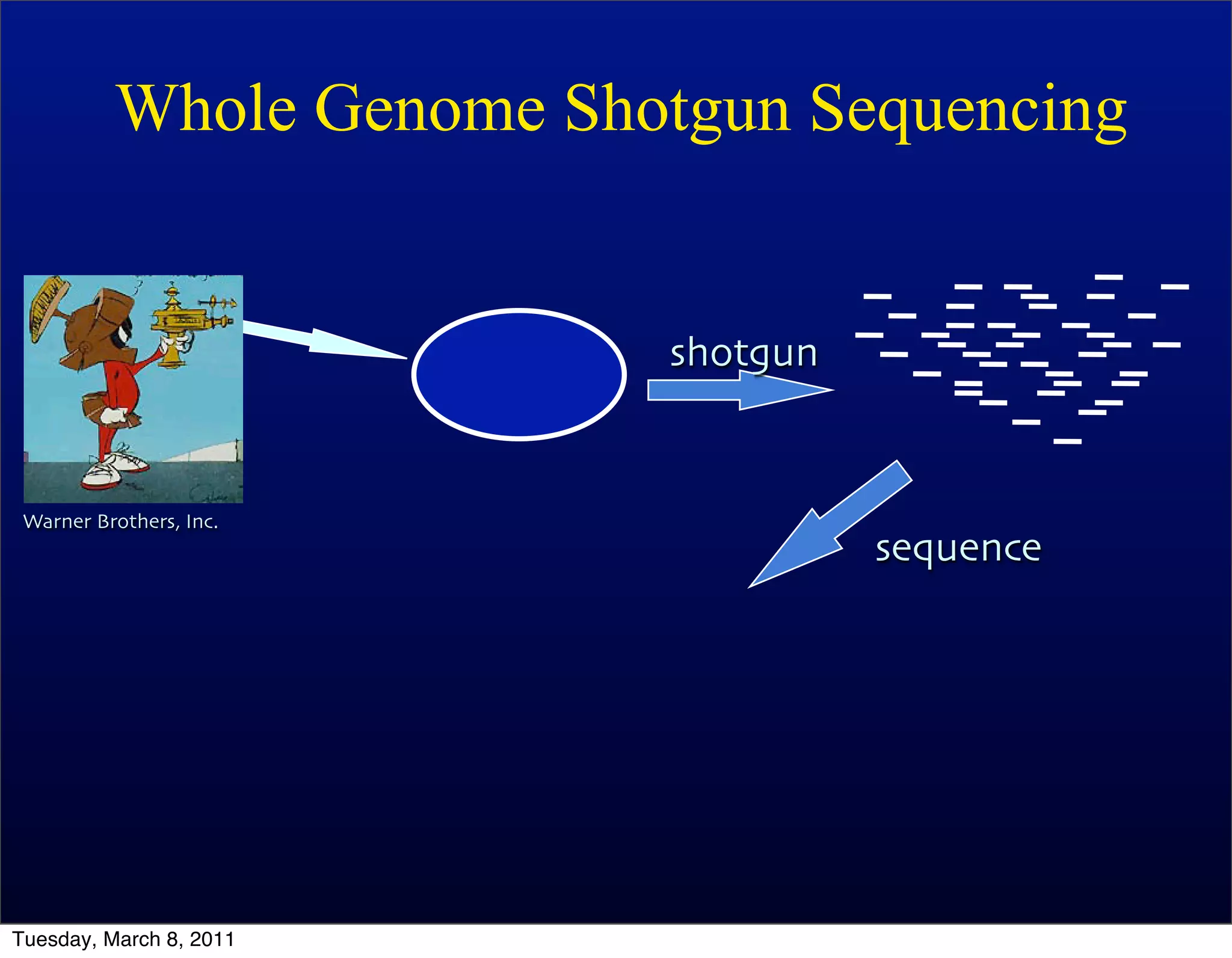
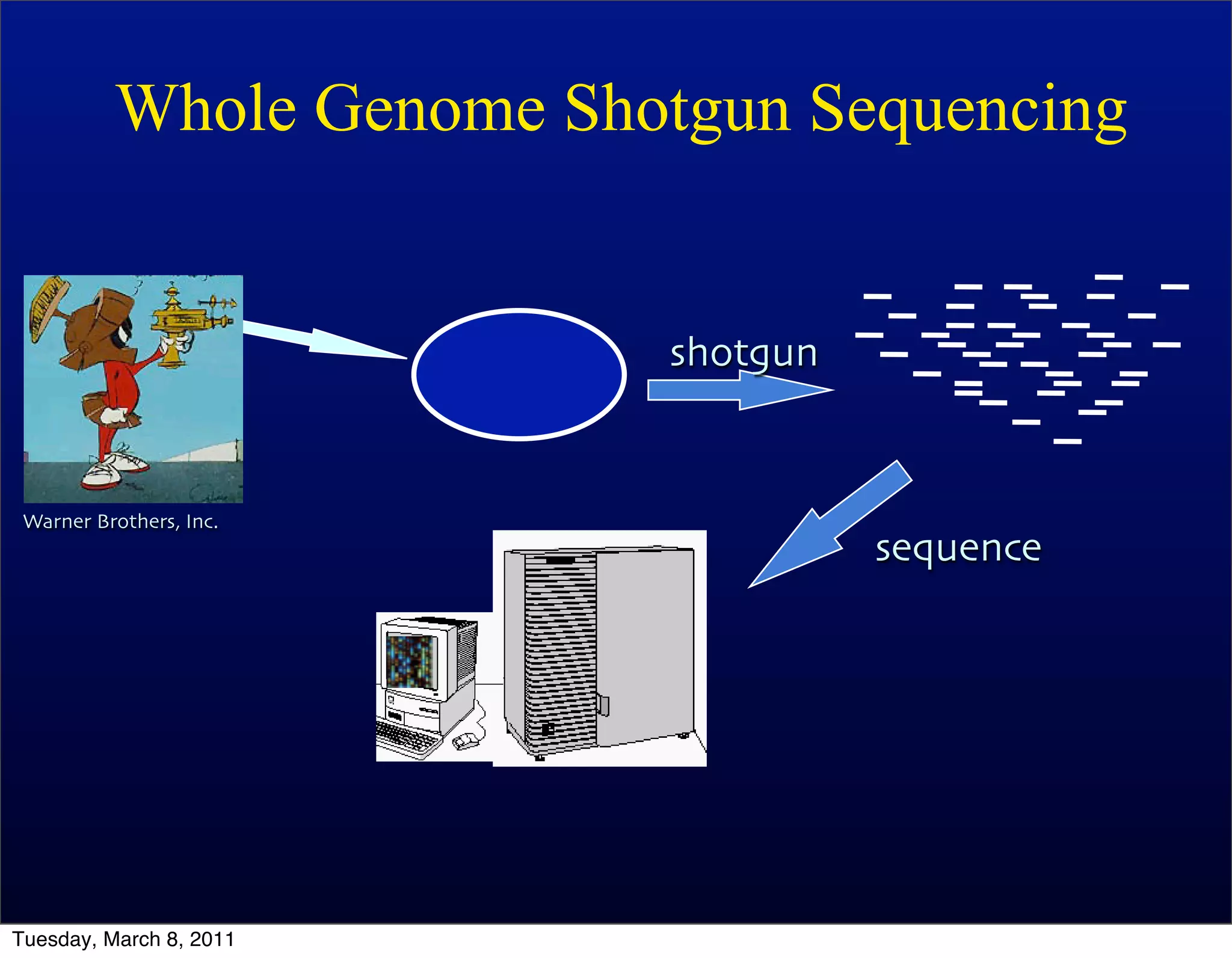
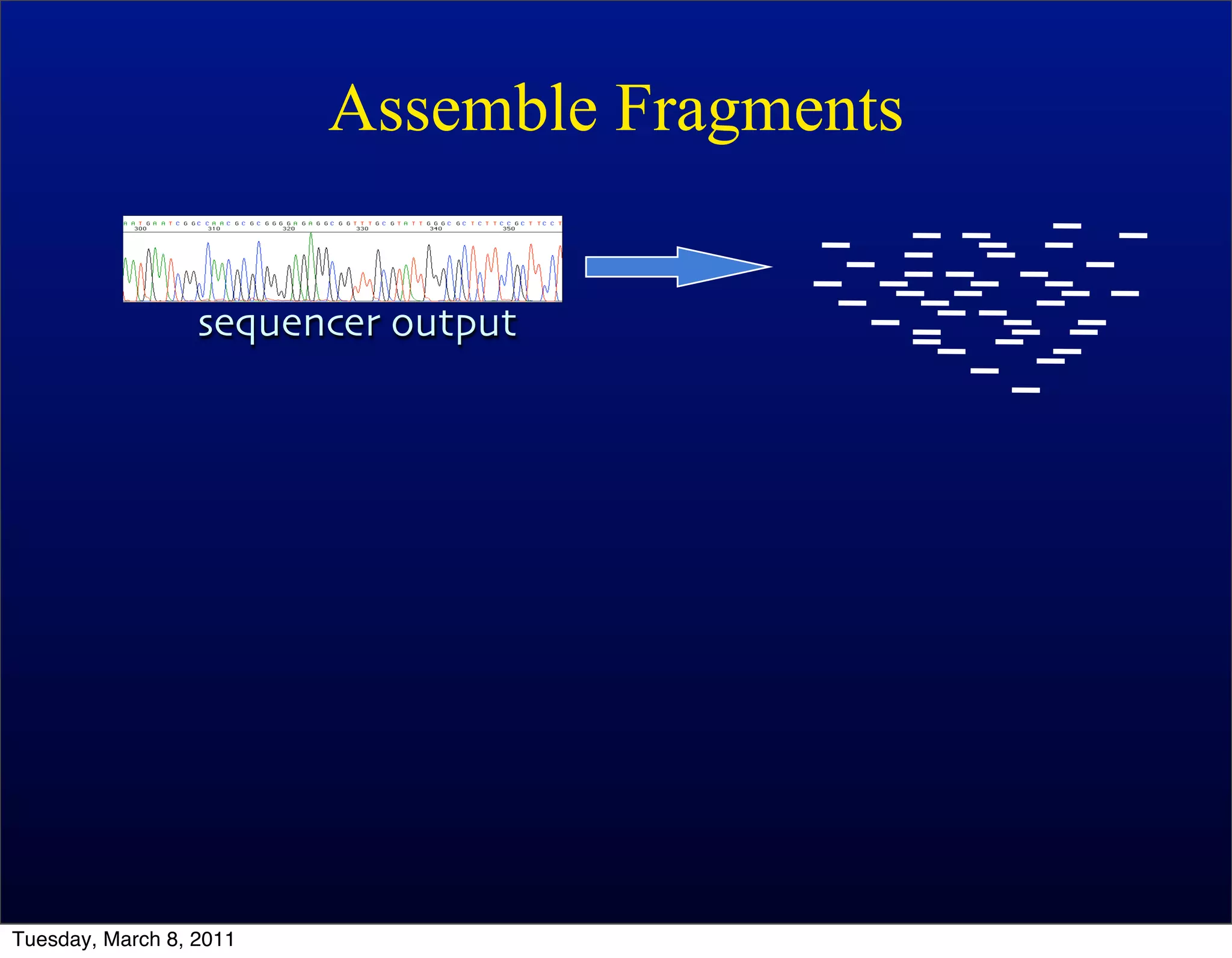
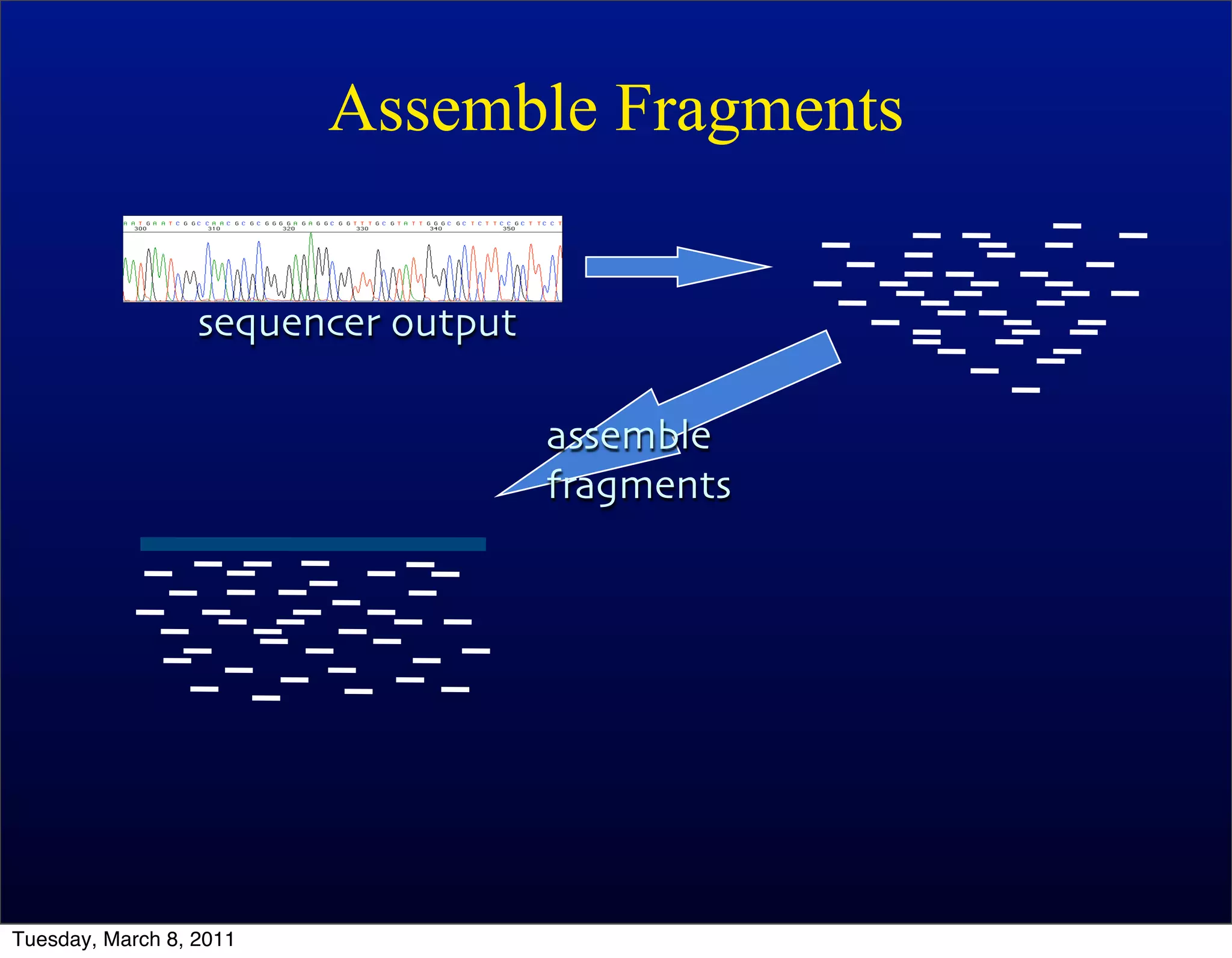
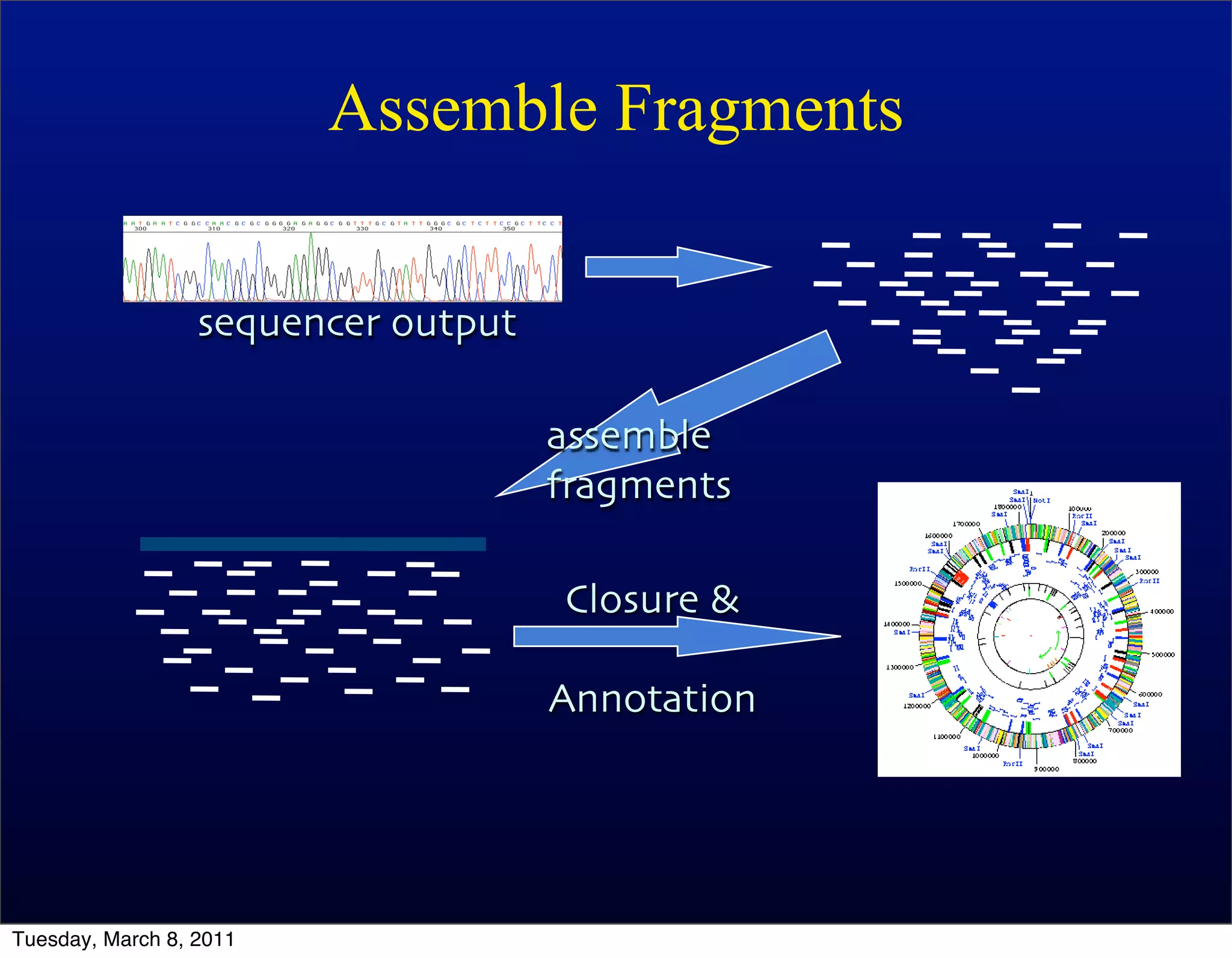
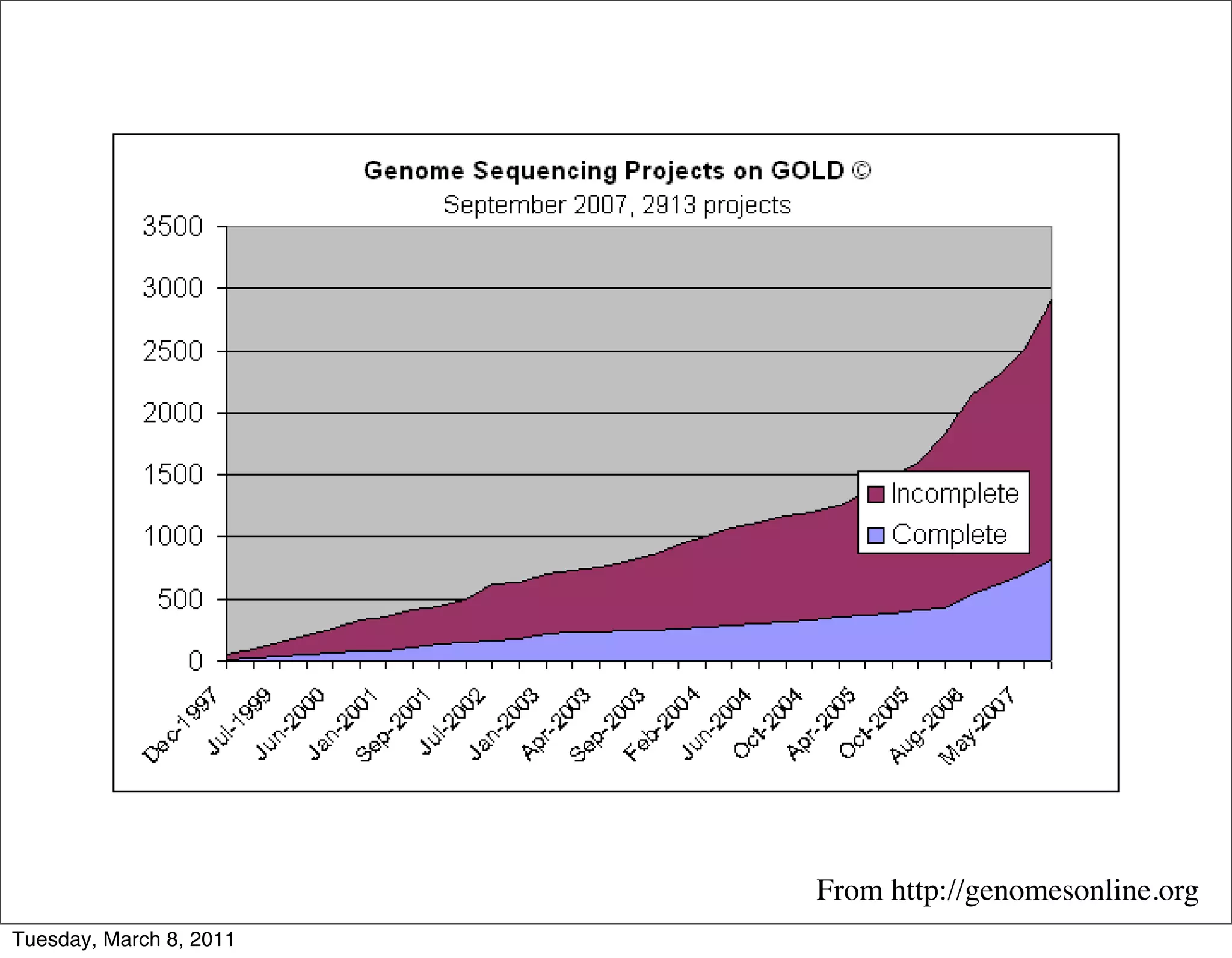
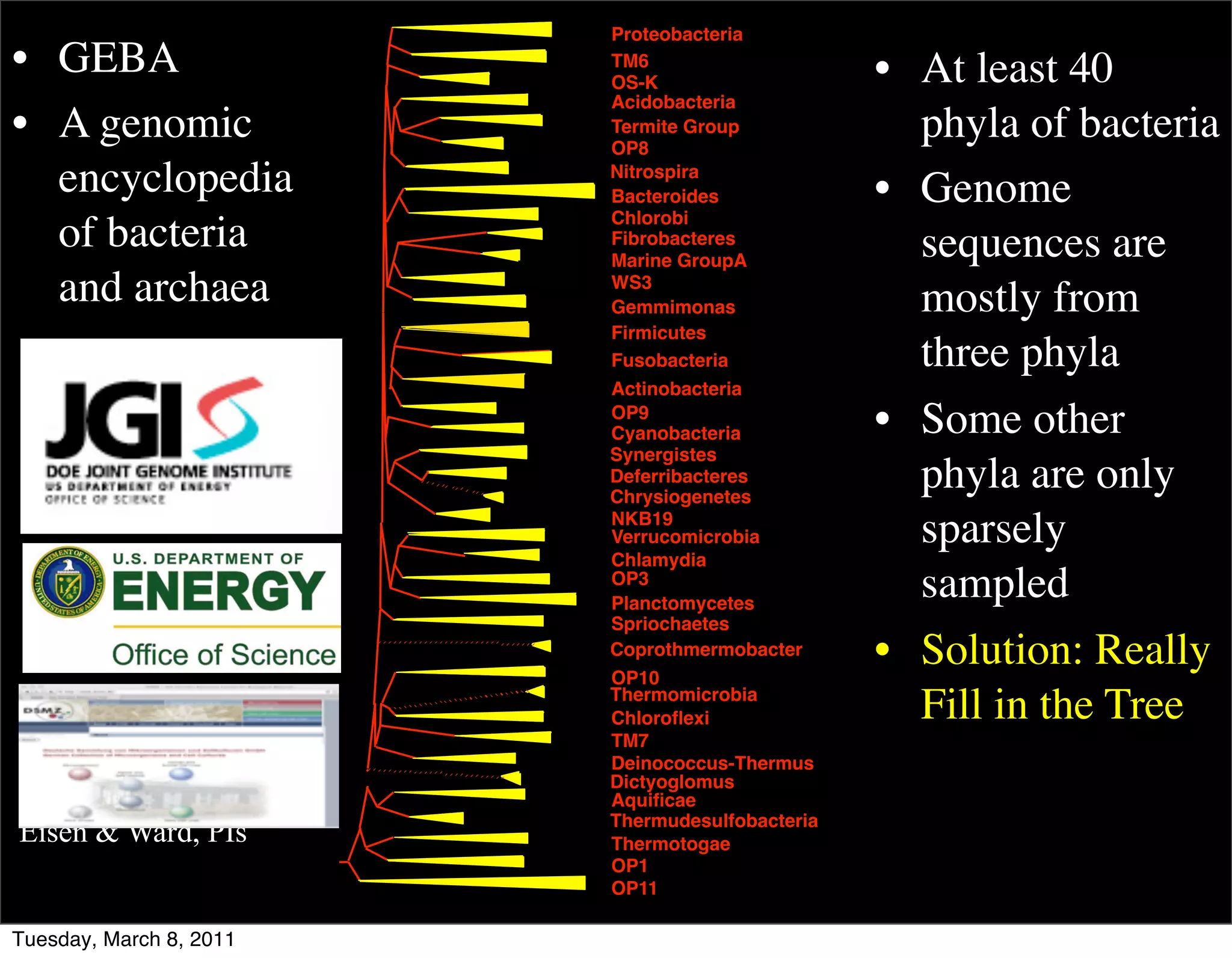
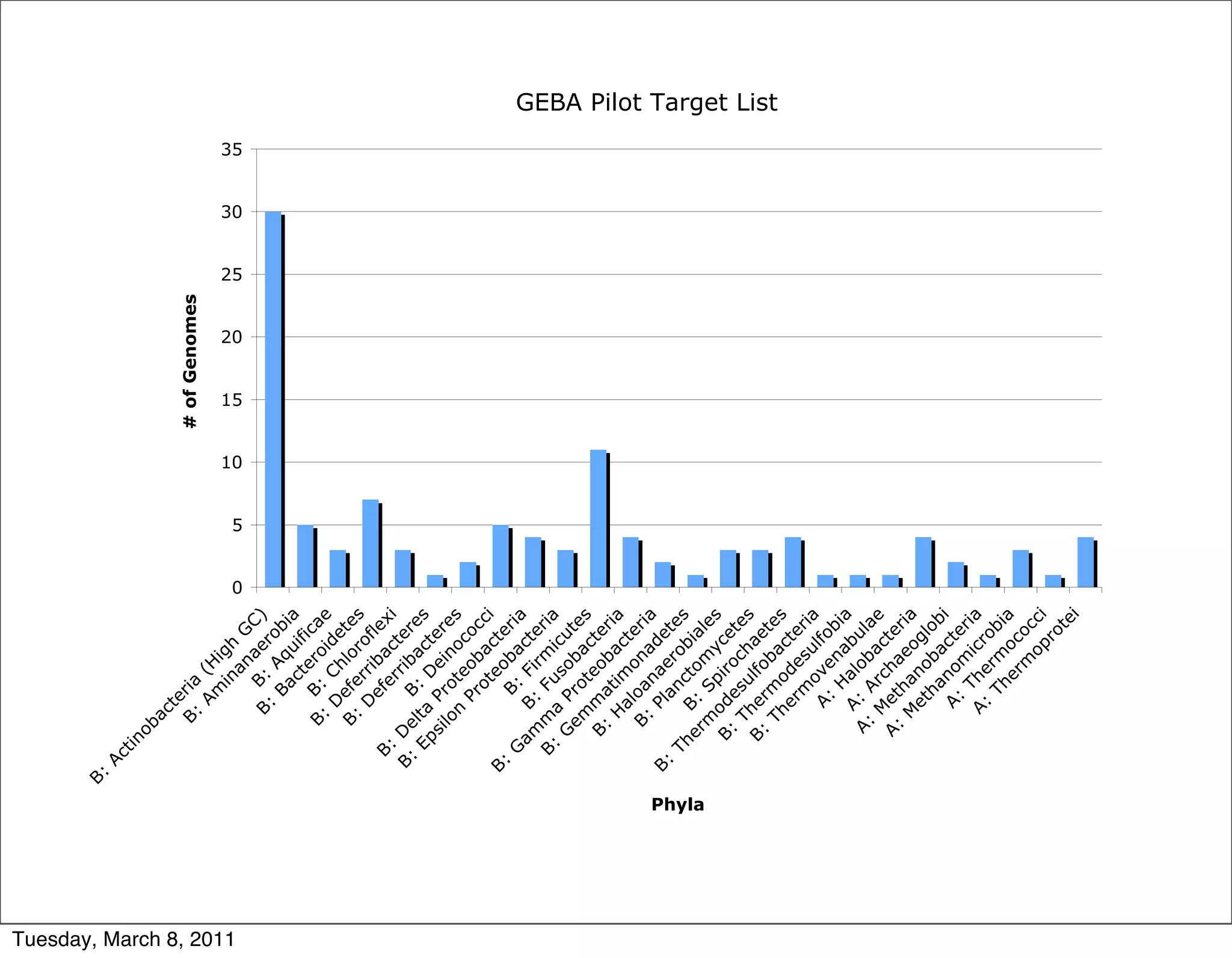
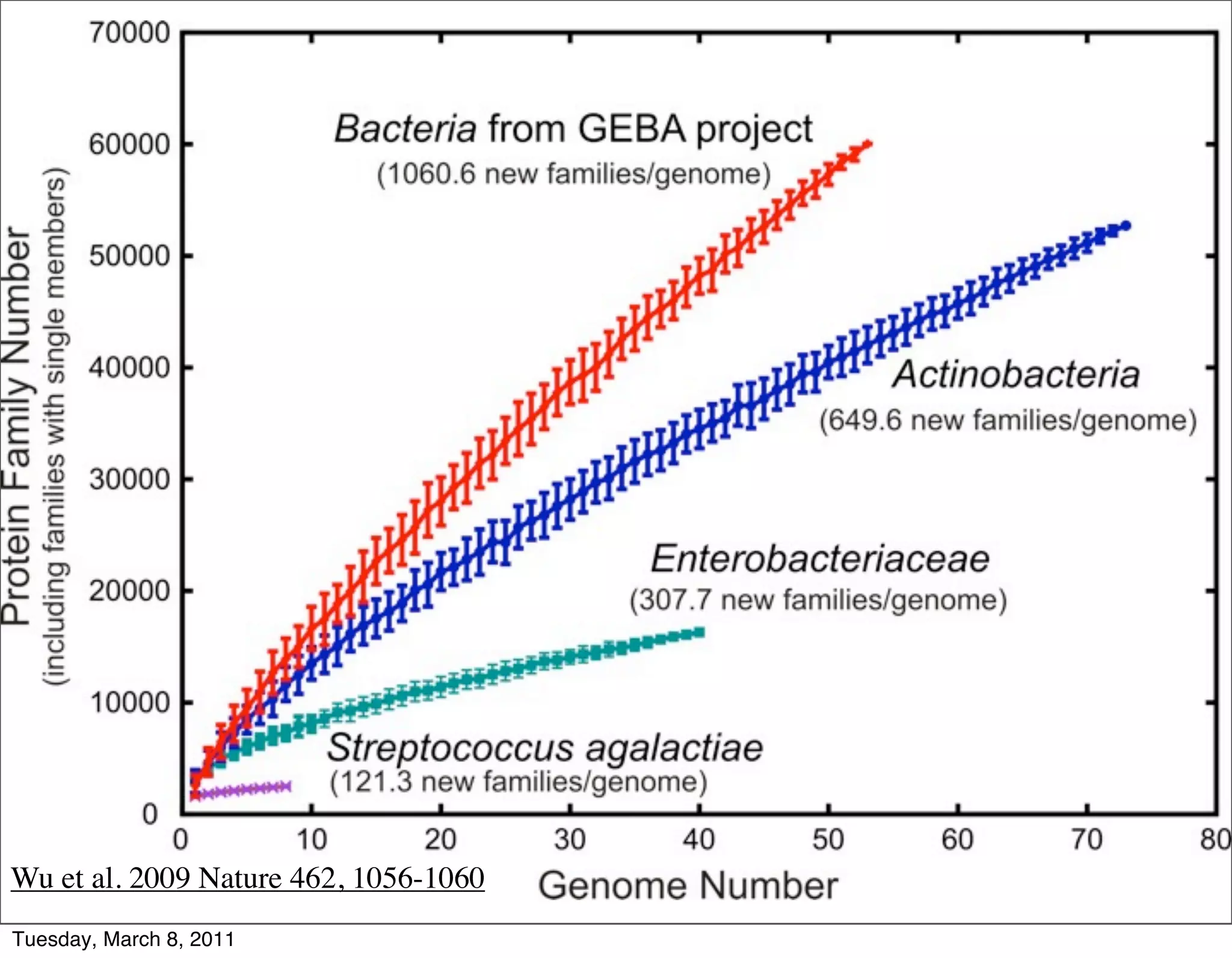
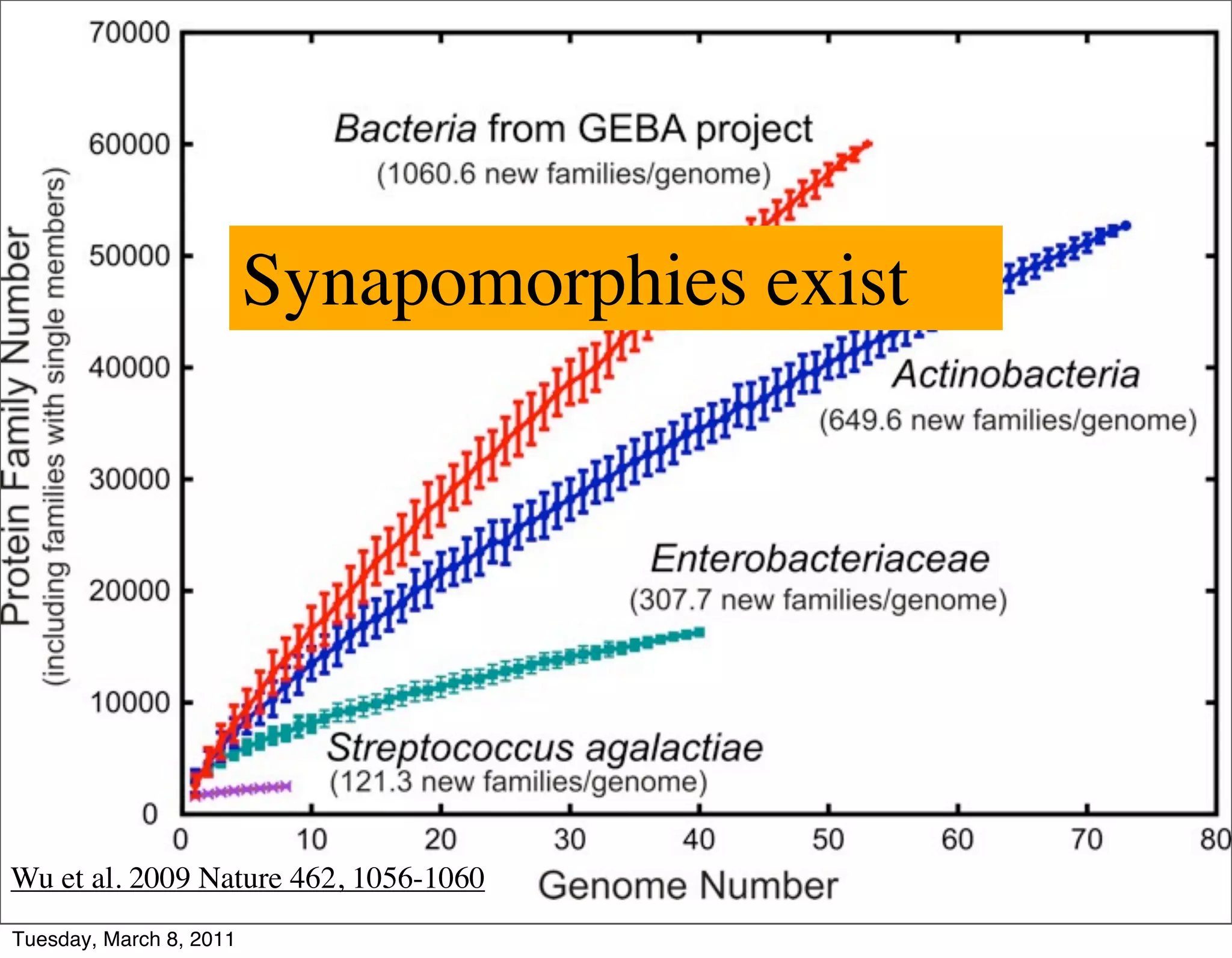
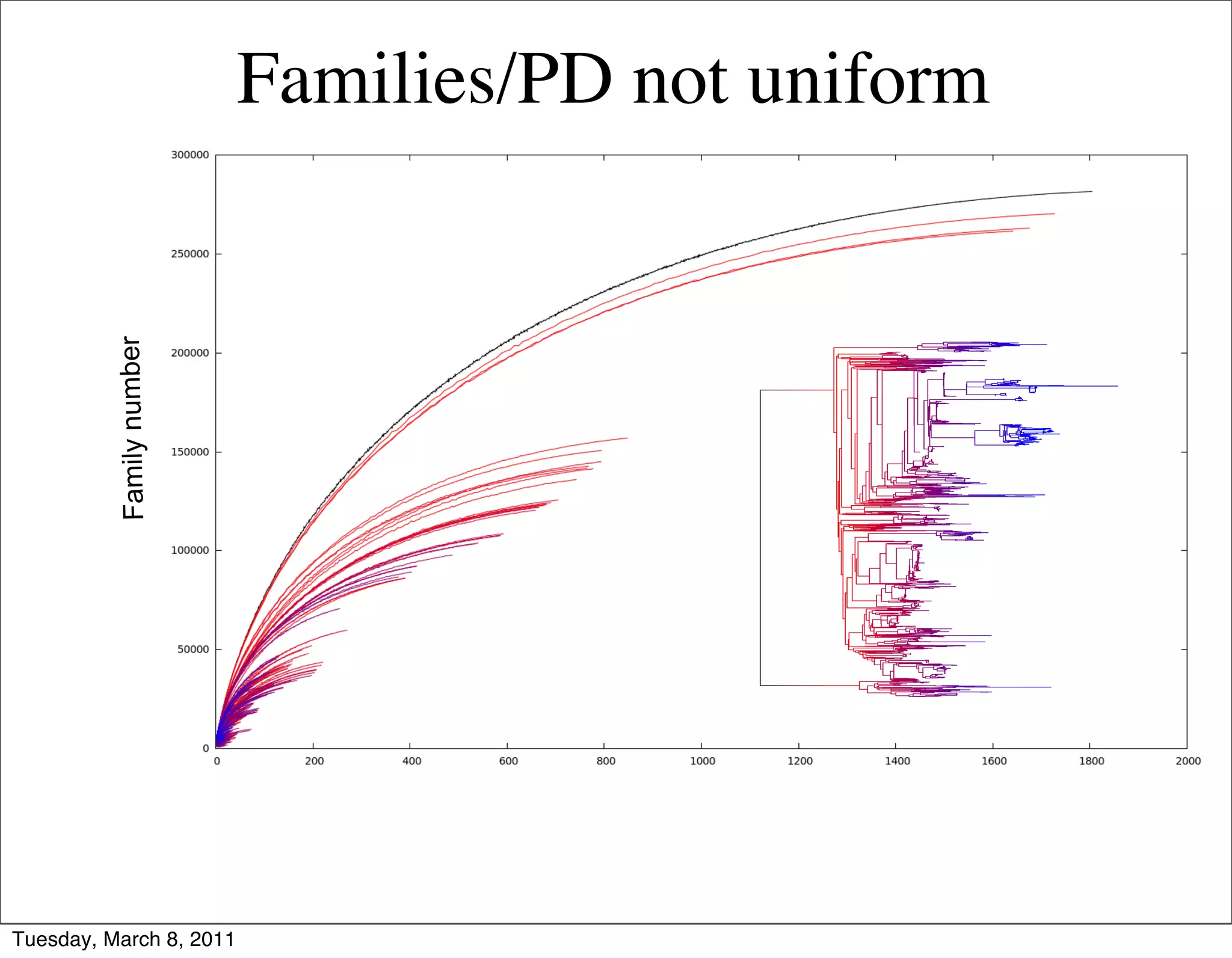
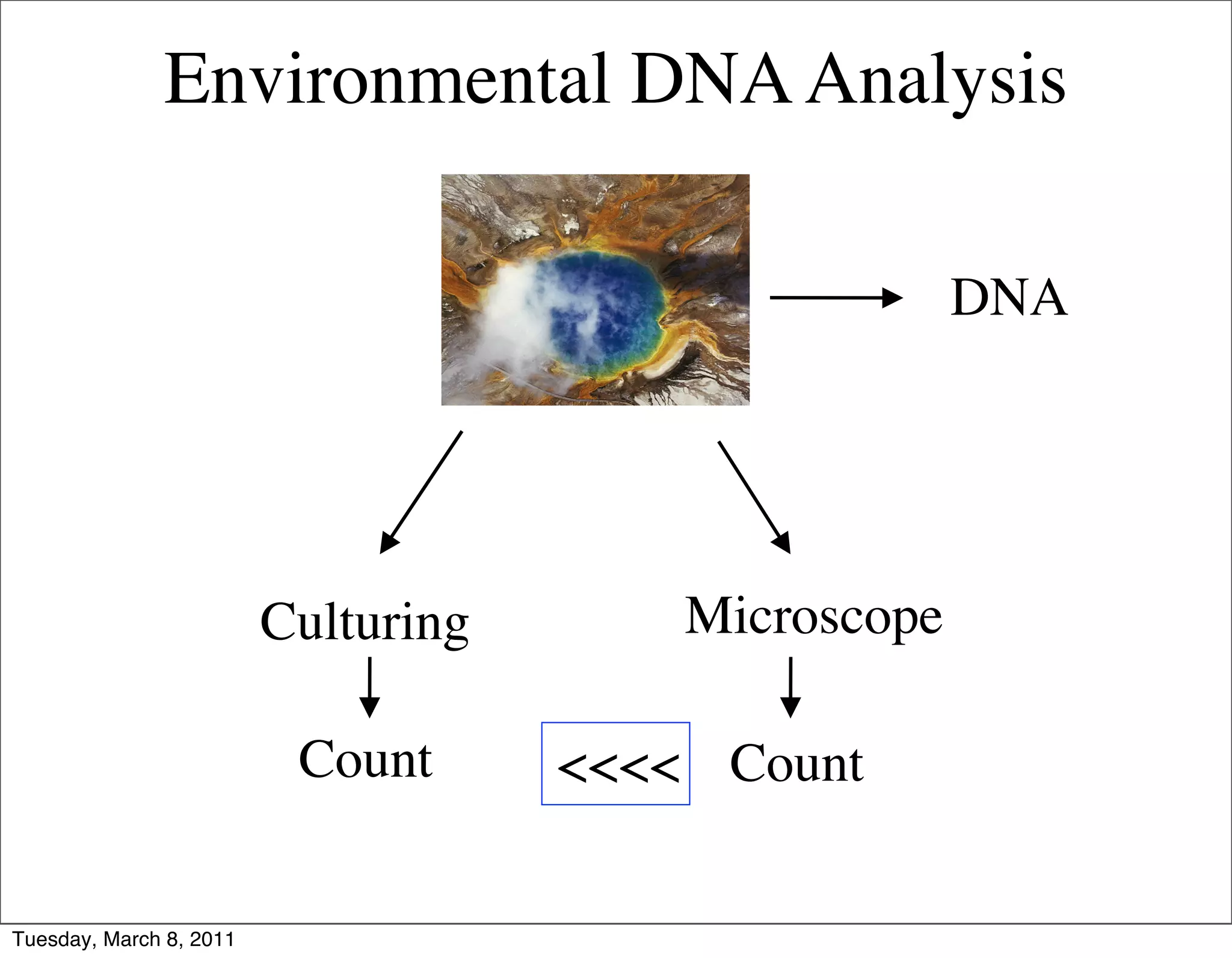
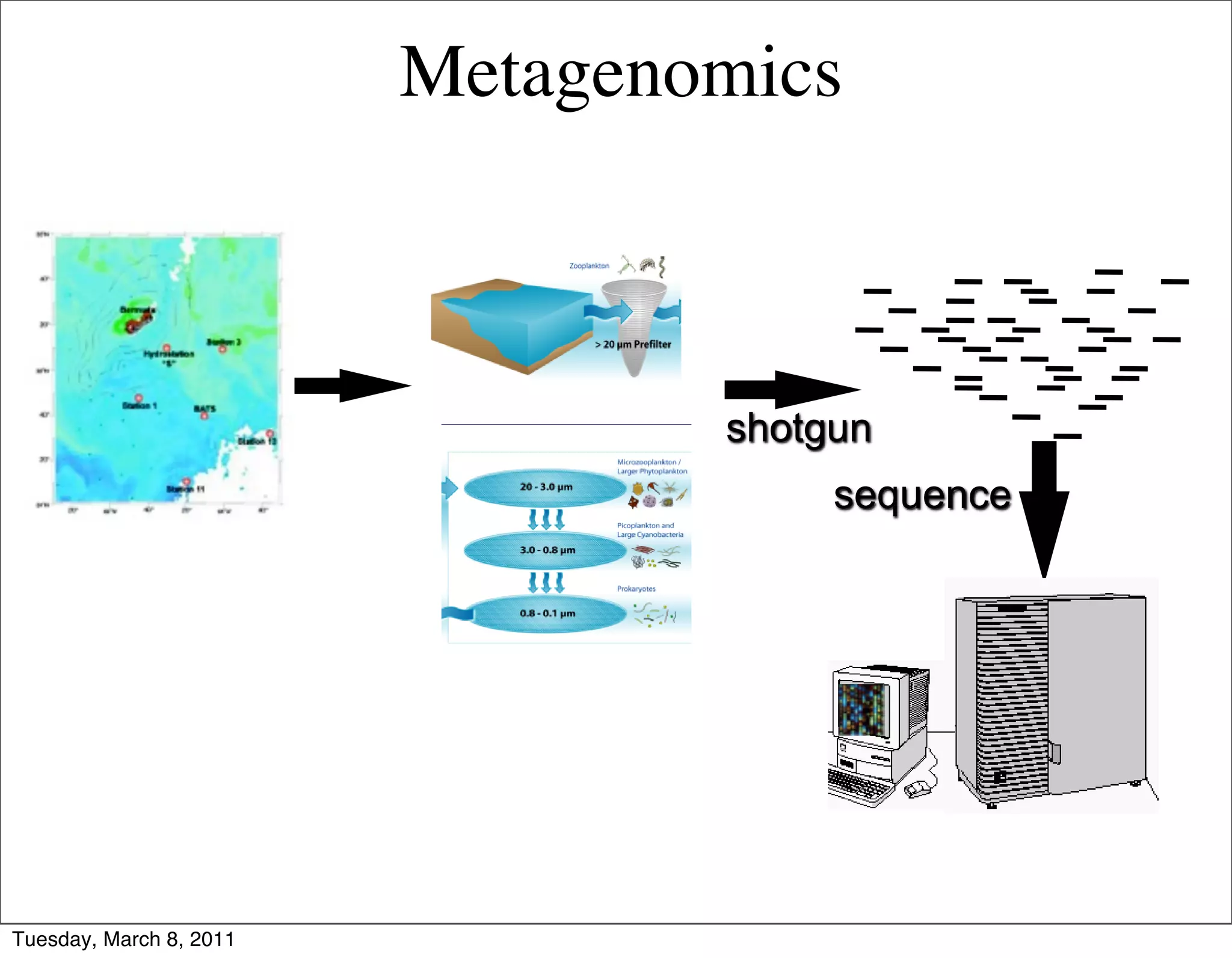
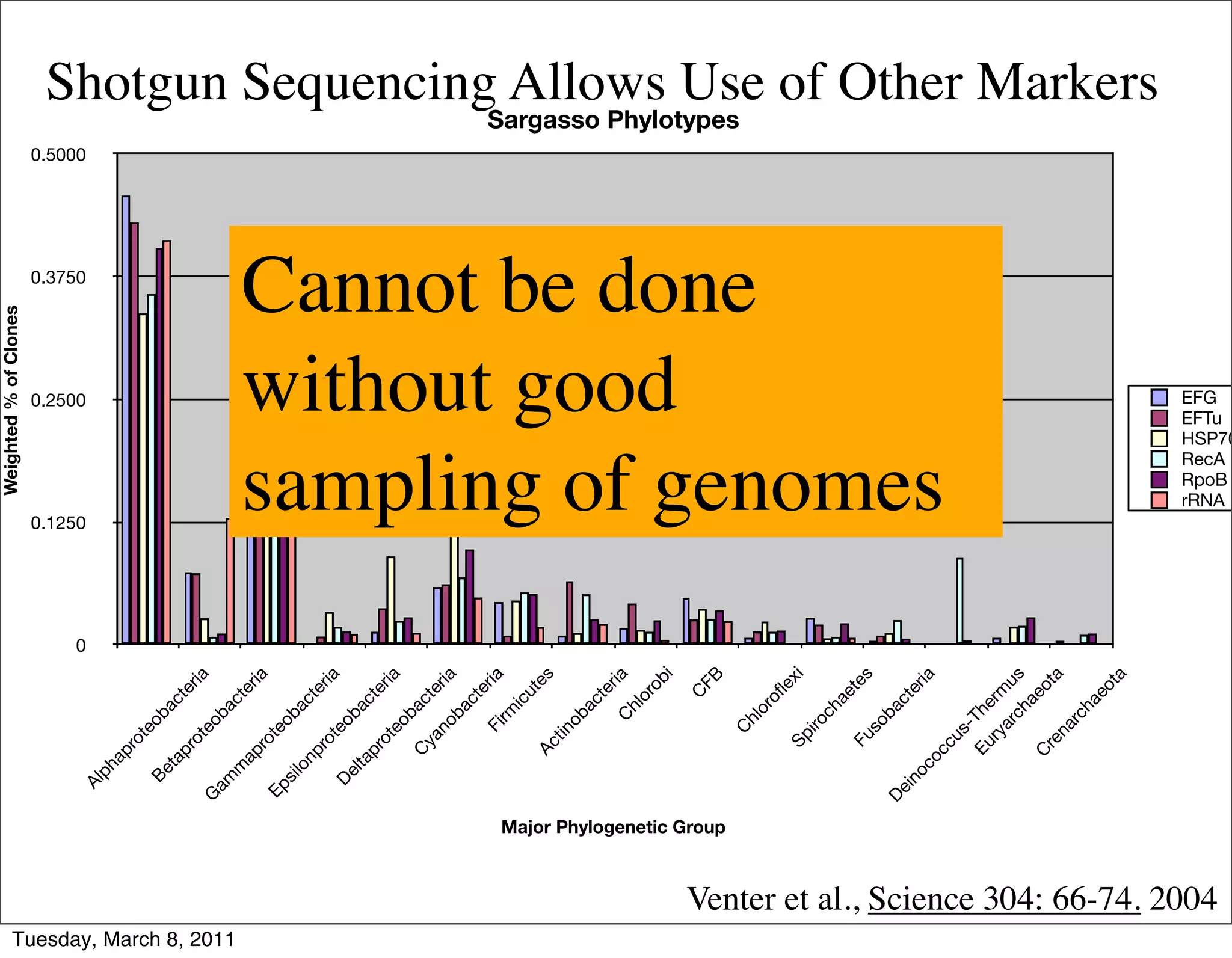
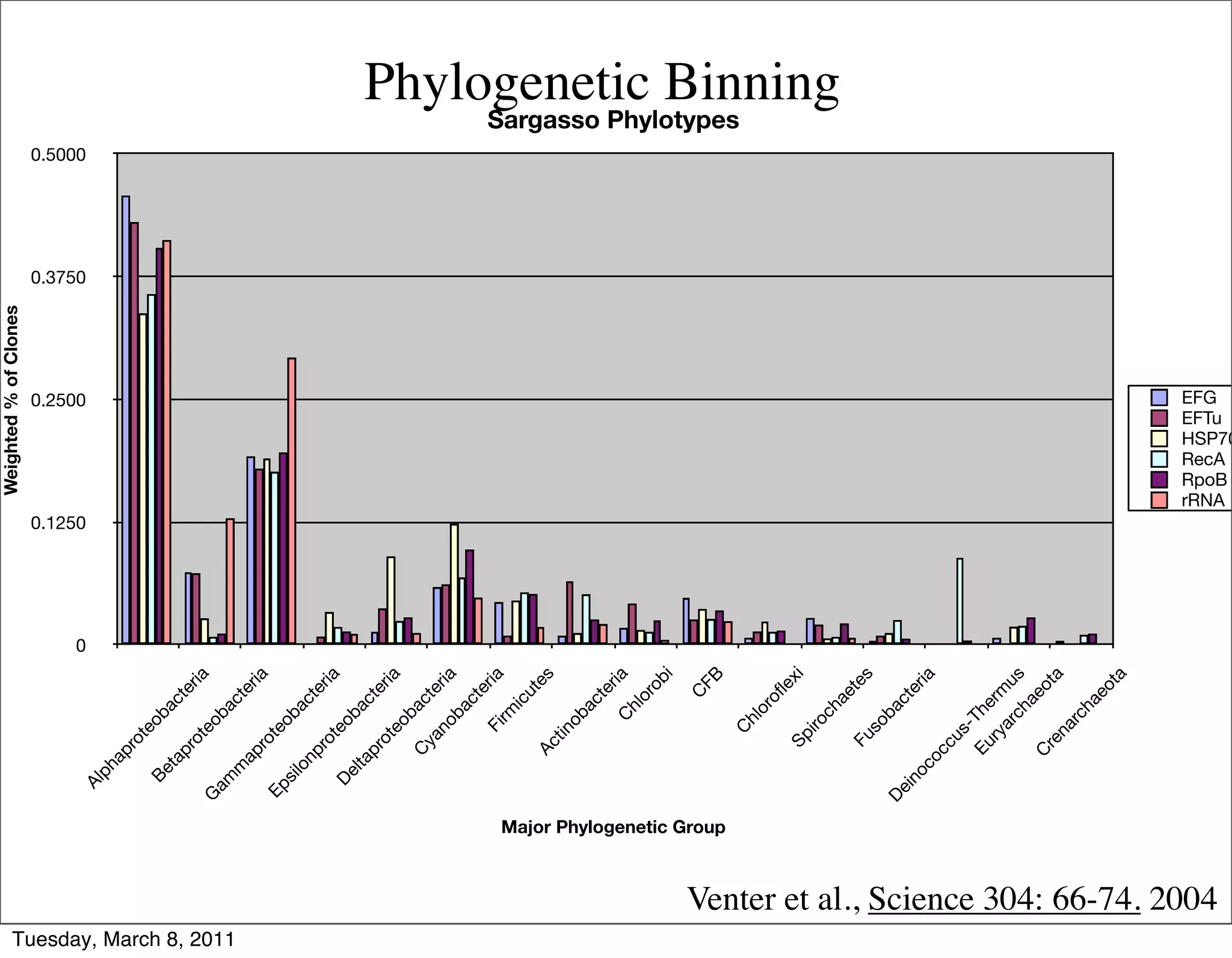
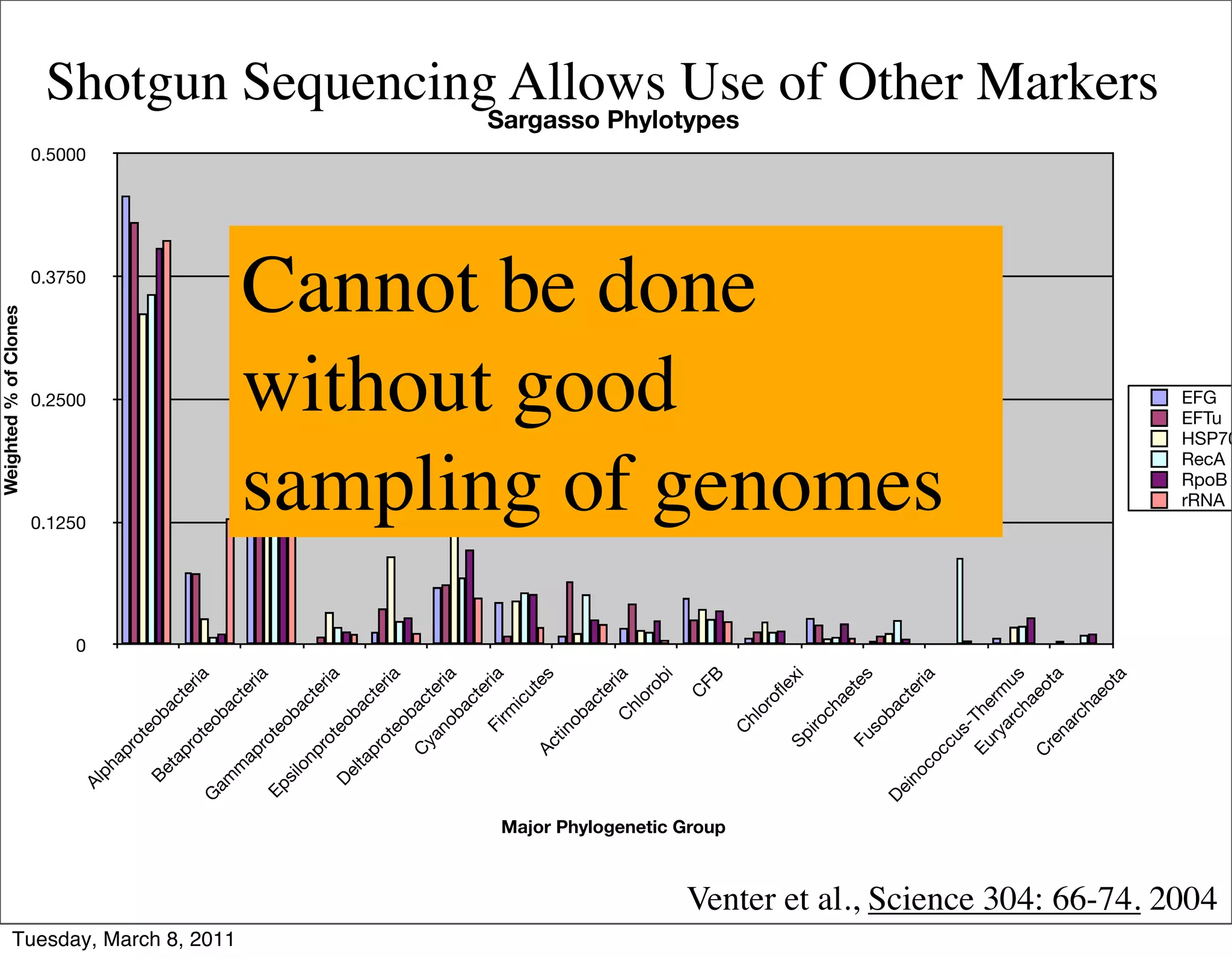
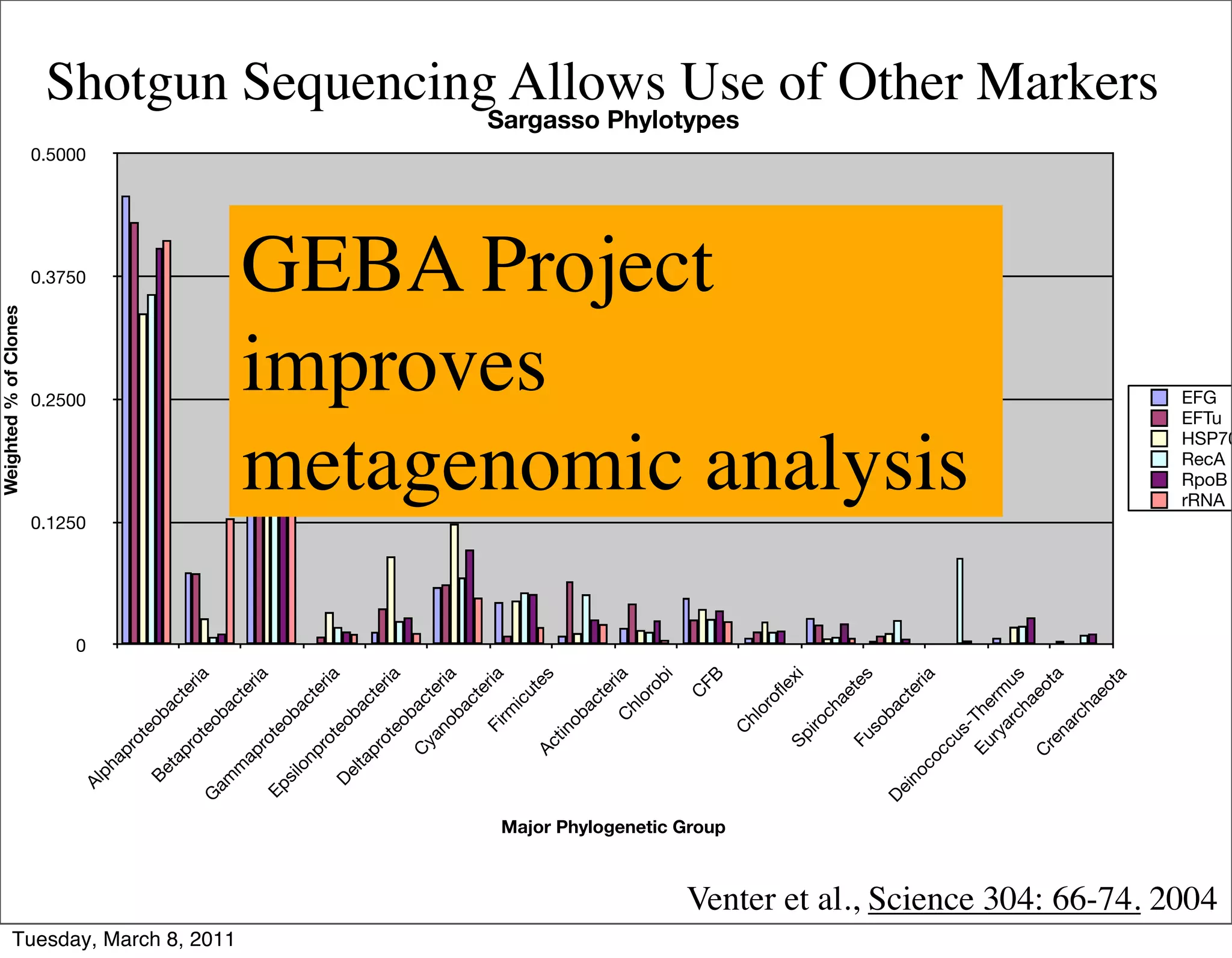
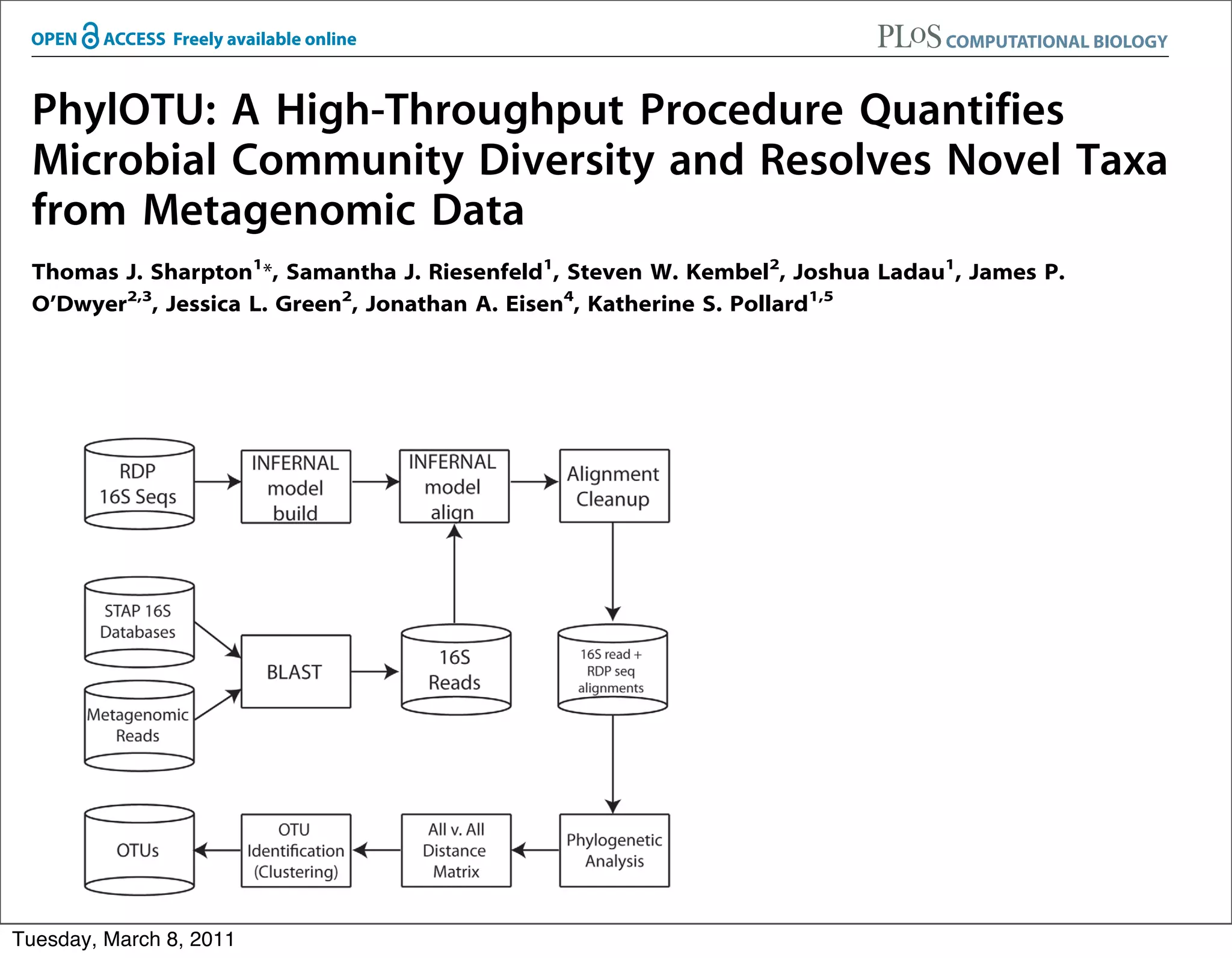
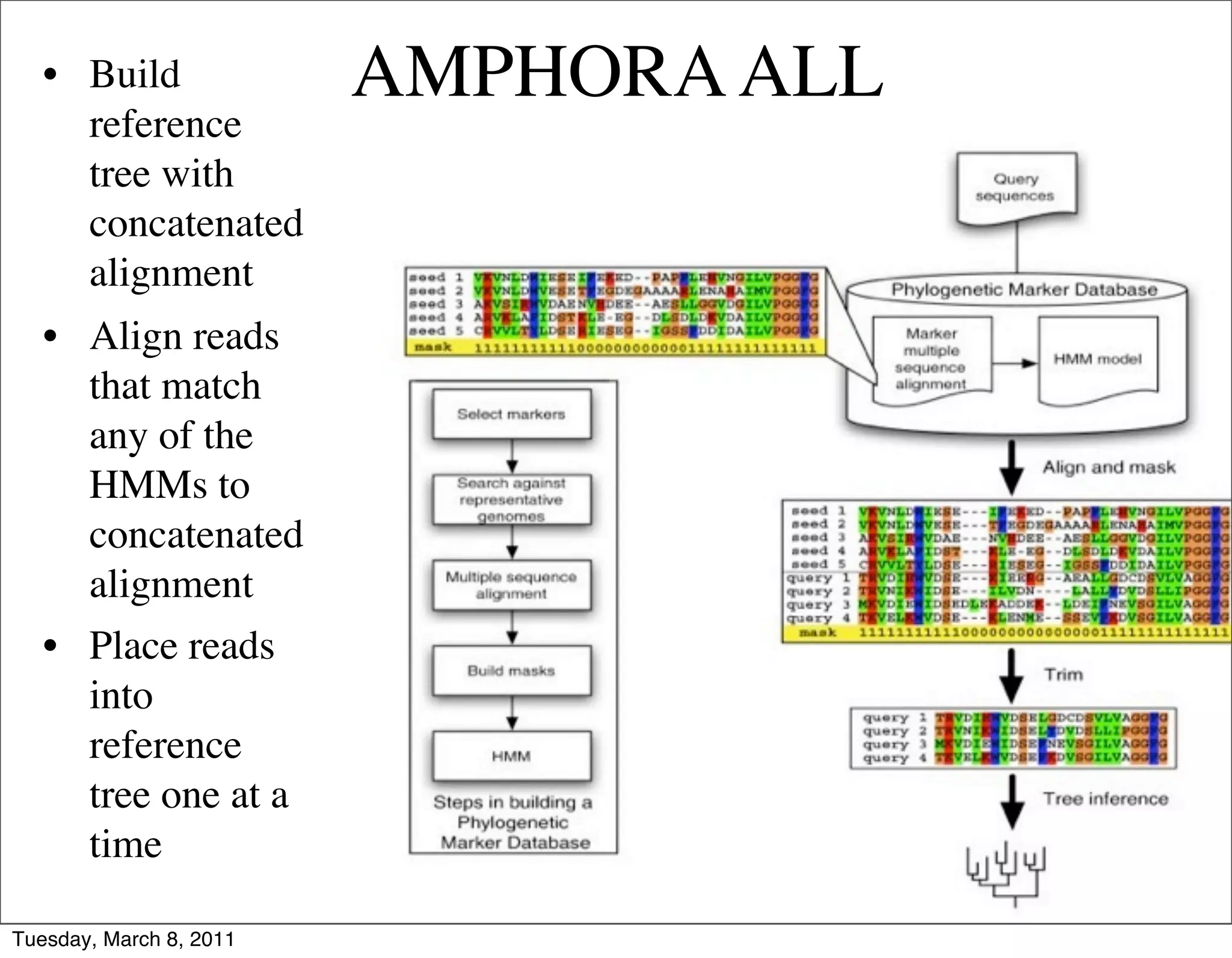
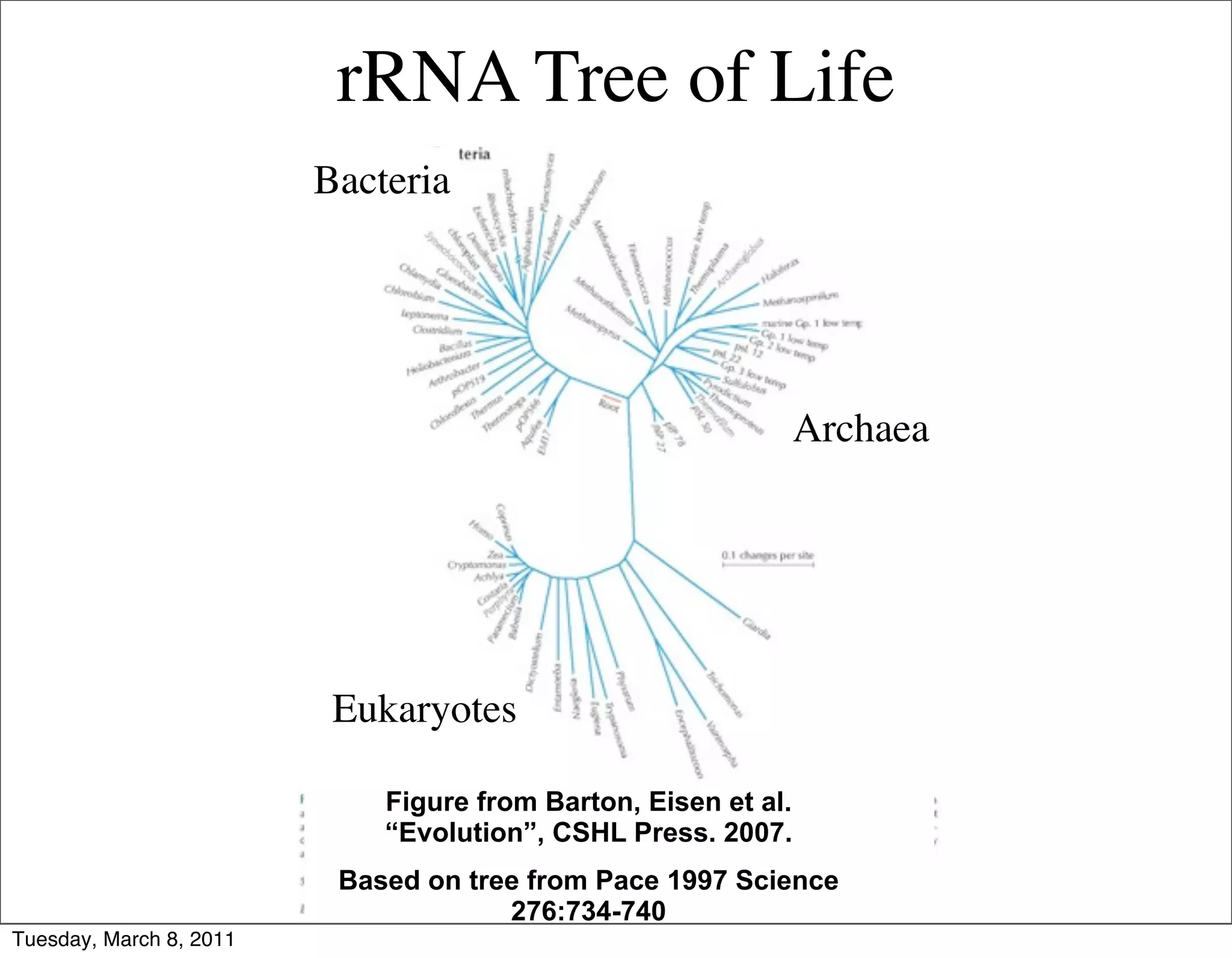
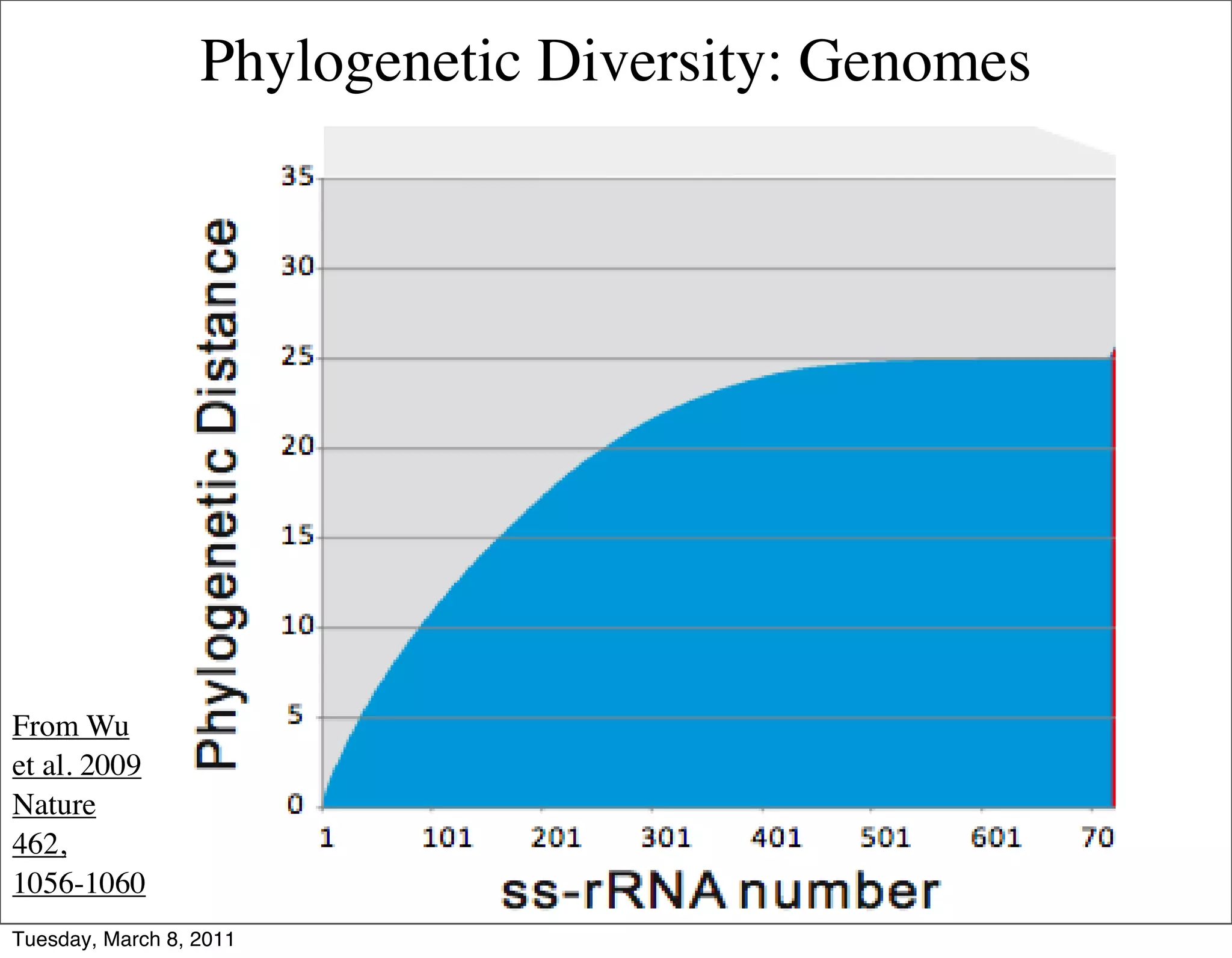
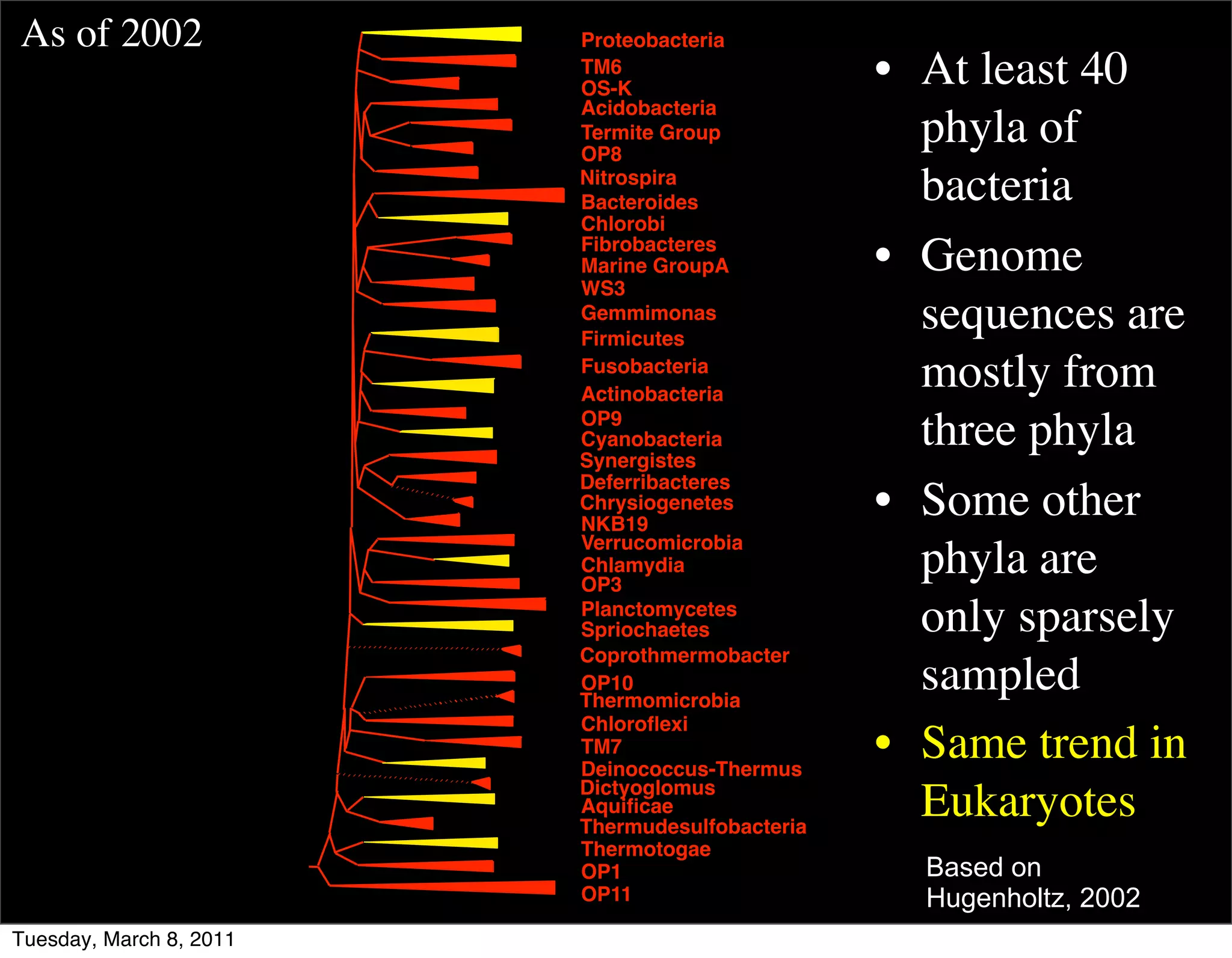
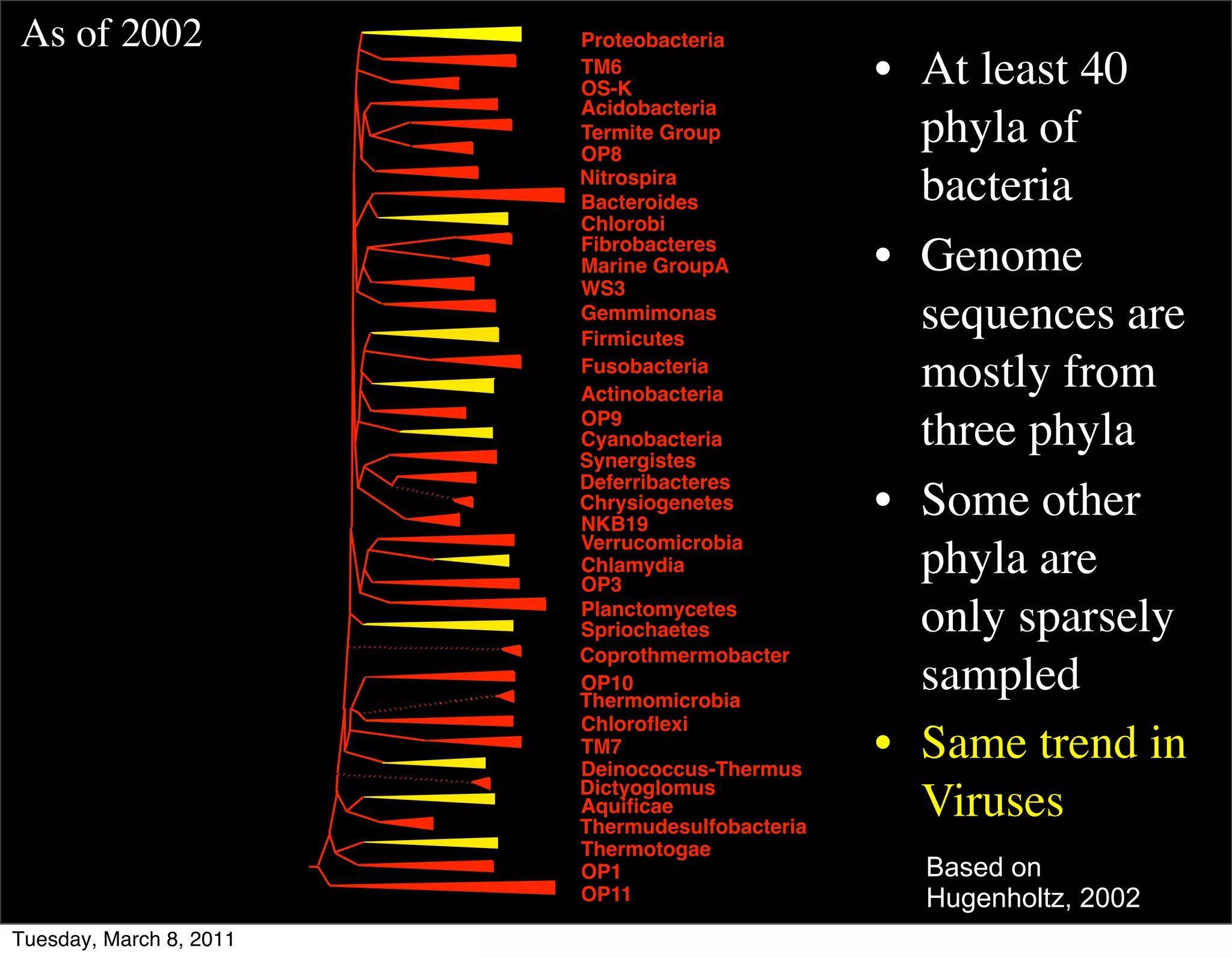
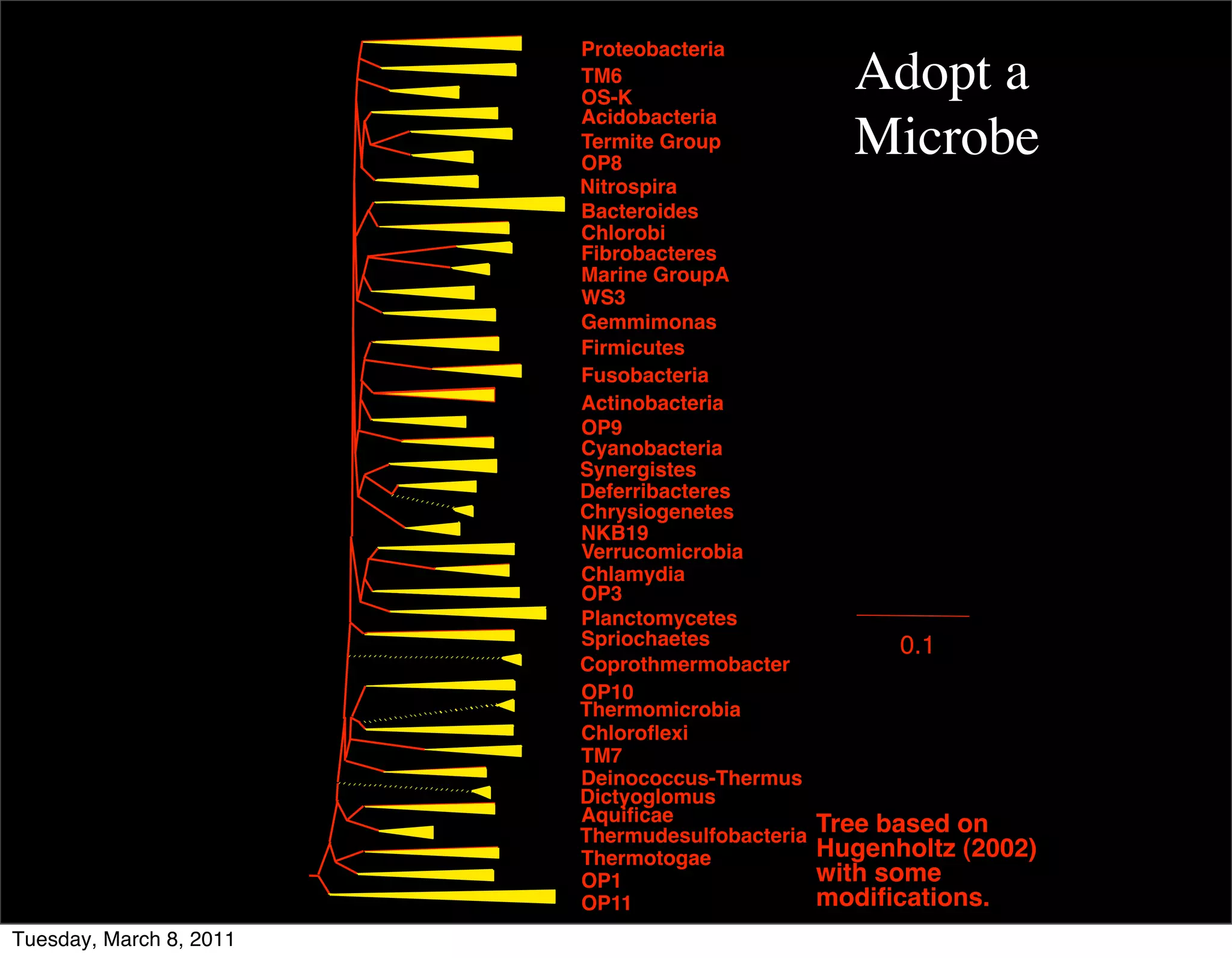
The document discusses whole genome shotgun sequencing. It explains that shotgun sequencing involves breaking the genome into random fragments, sequencing the fragments, and then assembling the sequences back into the original genome. It notes that this approach was pioneered by The Institute for Genomic Research and has revolutionized genome sequencing by allowing rapid determination of entire microbial genomes.