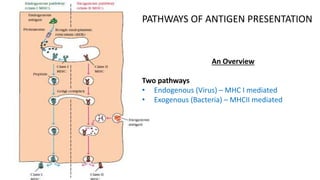
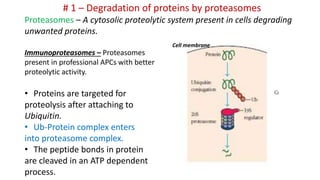
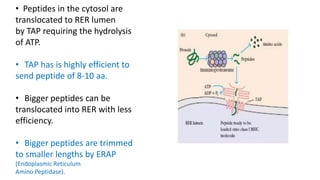
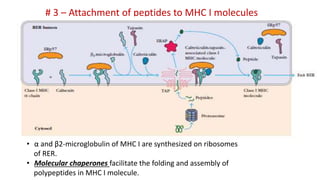
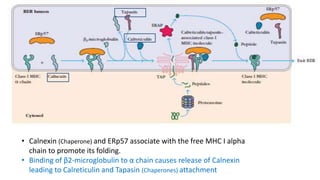
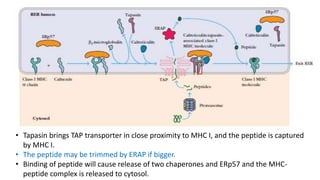
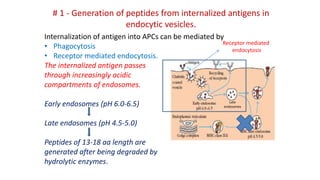
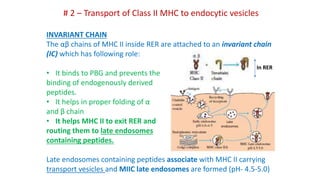
The document provides an overview of the historical development of protective immunity. It discusses key findings and contributors such as Edward Jenner who demonstrated that cowpox fluid could protect against smallpox in 1798. Louis Pasteur introduced the concept of vaccination in 1885 through experiments showing attenuated pathogens caused less severe disease. Emil von Behring showed in 1890 that serum from animals immunized with attenuated diphtheria or tetanus viruses could cure untreated animals infected with these pathogens, providing evidence for passive immunity.