How to Sequence the lmmune Repertoire
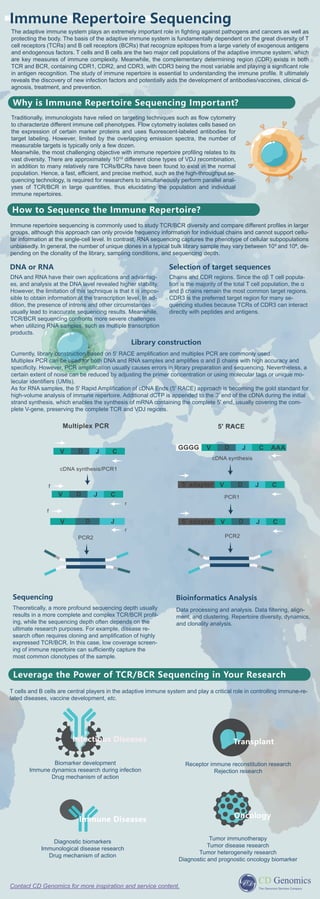
- 1. The adaptive immune system plays an extremely important role in fighting against pathogens and cancers as well as protecting the body. The basis of the adaptive immune system is fundamentally dependent on the great diversity of T cell receptors (TCRs) and B cell receptors (BCRs) that recognize epitopes from a large variety of exogenous antigens and endogenous factors. T cells and B cells are the two major cell populations of the adaptive immune system, which are key measures of immune complexity. Meanwhile, the complementary determining region (CDR) exists in both TCR and BCR, containing CDR1, CDR2, and CDR3, with CDR3 being the most variable and playing a significant role in antigen recognition. The study of immune repertoire is essential to understanding the immune profile. It ultimately reveals the discovery of new infection factors and potentially aids the development of antibodies/vaccines, clinical di- agnosis, treatment, and prevention. Contact CD Genomics for more inspiration and service content. Immune Repertoire Sequencing Why is Immune Repertoire Sequencing Important? Immune repertoire sequencing is commonly used to study TCR/BCR diversity and compare different profiles in larger groups, although this approach can only provide frequency information for individual chains and cannot support cellu- lar information at the single-cell level. In contrast, RNA sequencing captures the phenotype of cellular subpopulations unbiasedly. In general, the number of unique clones in a typical bulk library sample may vary between 10³ and 10⁶, de- pending on the clonality of the library, sampling conditions, and sequencing depth. How to Sequence the Immune Repertoire? Traditionally, immunologists have relied on targeting techniques such as flow cytometry to characterize different immune cell phenotypes. Flow cytometry isolates cells based on the expression of certain marker proteins and uses fluorescent-labeled antibodies for target labeling. However, limited by the overlapping emission spectra, the number of measurable targets is typically only a few dozen. Meanwhile, the most challenging objective with immune repertoire profiling relates to its vast diversity. There are approximately 10¹³ different clone types of VDJ recombination, in addition to many relatively rare TCRs/BCRs have been found to exist in the normal population. Hence, a fast, efficient, and precise method, such as the high-throughput se- quencing technology, is required for researchers to simultaneously perform parallel anal- yses of TCR/BCR in large quantities, thus elucidating the population and individual immune repertoires. DNA or RNA Selection of target sequences Chains and CDR regions. Since the αβ T cell popula- tion is the majority of the total T cell population, the α and β chains remain the most common target regions. CDR3 is the preferred target region for many se- quencing studies because TCRs of CDR3 can interact directly with peptides and antigens. Library construction Sequencing Theoretically, a more profound sequencing depth usually results in a more complete and complex TCR/BCR profil- ing, while the sequencing depth often depends on the ultimate research purposes. For example, disease re- search often requires cloning and amplification of highly expressed TCR/BCR. In this case, low coverage screen- ing of immune repertoire can sufficiently capture the most common clonotypes of the sample. Bioinformatics Analysis Data processing and analysis. Data filtering, align- ment, and clustering. Repertoire diversity, dynamics, and clonality analysis. Leverage the Power of TCR/BCR Sequencing in Your Research T cells and B cells are central players in the adaptive immune system and play a critical role in controlling immune-re- lated diseases, vaccine development, etc. DNA and RNA have their own applications and advantag- es, and analysis at the DNA level revealed higher stability. However, the limitation of this technique is that it is impos- sible to obtain information at the transcription level. In ad- dition, the presence of introns and other circumstances usually lead to inaccurate sequencing results. Meanwhile, TCR/BCR sequencing confronts more severe challenges when utilizing RNA samples, such as multiple transcription products. Currently, library construction based on 5' RACE amplification and multiplex PCR are commonly used. Multiplex PCR can be used for both DNA and RNA samples and amplifies α and β chains with high accuracy and specificity. However, PCR amplification usually causes errors in library preparation and sequencing. Nevertheless, a certain extent of noise can be reduced by adjusting the primer concentration or using molecular tags or unique mo- lecular identifiers (UMIs). As for RNA samples, the 5' Rapid Amplification of cDNA Ends (5' RACE) approach is becoming the gold standard for high-volume analysis of immune repertoire. Additional dCTP is appended to the 3' end of the cDNA during the initial strand synthesis, which enables the synthesis of mRNA containing the complete 5' end, usually covering the com- plete V-gene, preserving the complete TCR and VDJ regions. Infectious Diseases Biomarker development Immune dynamics research during infection Drug mechanism of action Transplant Receptor immune reconstitution research Rejection research Oncology Tumor immunotherapy Tumor disease research Tumor heterogeneity research Diagnostic and prognostic oncology biomarker Diagnostic biomarkers Immunological disease research Drug mechanism of action Immune Diseases V D C J V D J V D C J cDNA synthesis/PCR1 PCR2 f r f r Multiplex PCR PCR1 PCR2 V D C J 5’ adapter V D C J GGGG AAA cDNA synthesis V D C J 5’ adapter 5' RACE