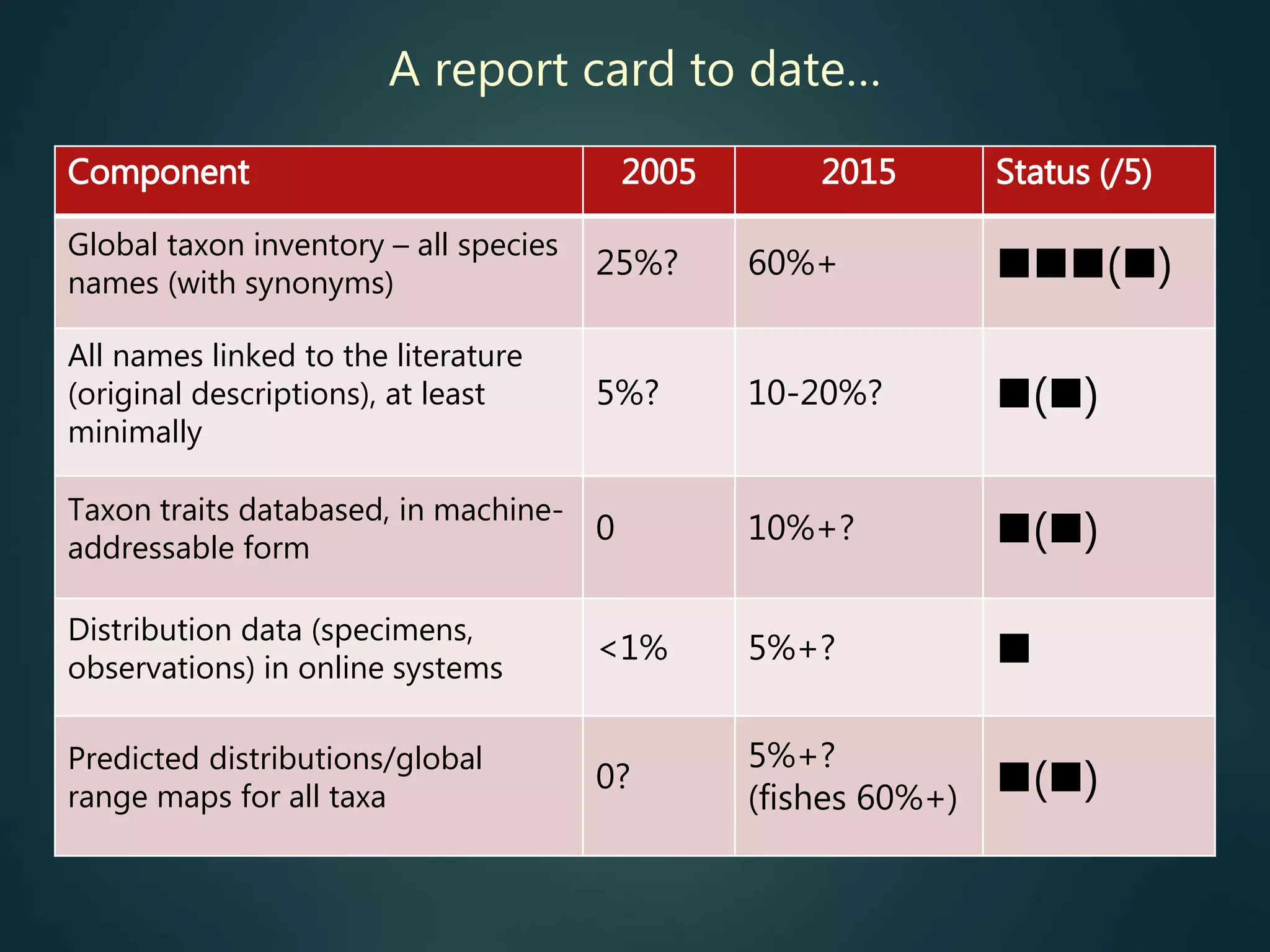
The document discusses the progress made over the past 10 years towards achieving a vision of having comprehensive biodiversity information accessible online. It examines several key components of this vision including developing a global taxon inventory, linking names to primary literature, assembling georeferenced species records, and creating machine-readable taxon trait data. For each component, it provides details on the scope of the task, current resources addressing it, and how complete the work is so far, noting many challenges that still remain.

![The vision: “Biodiversity information on every
desktop” [ / device]…
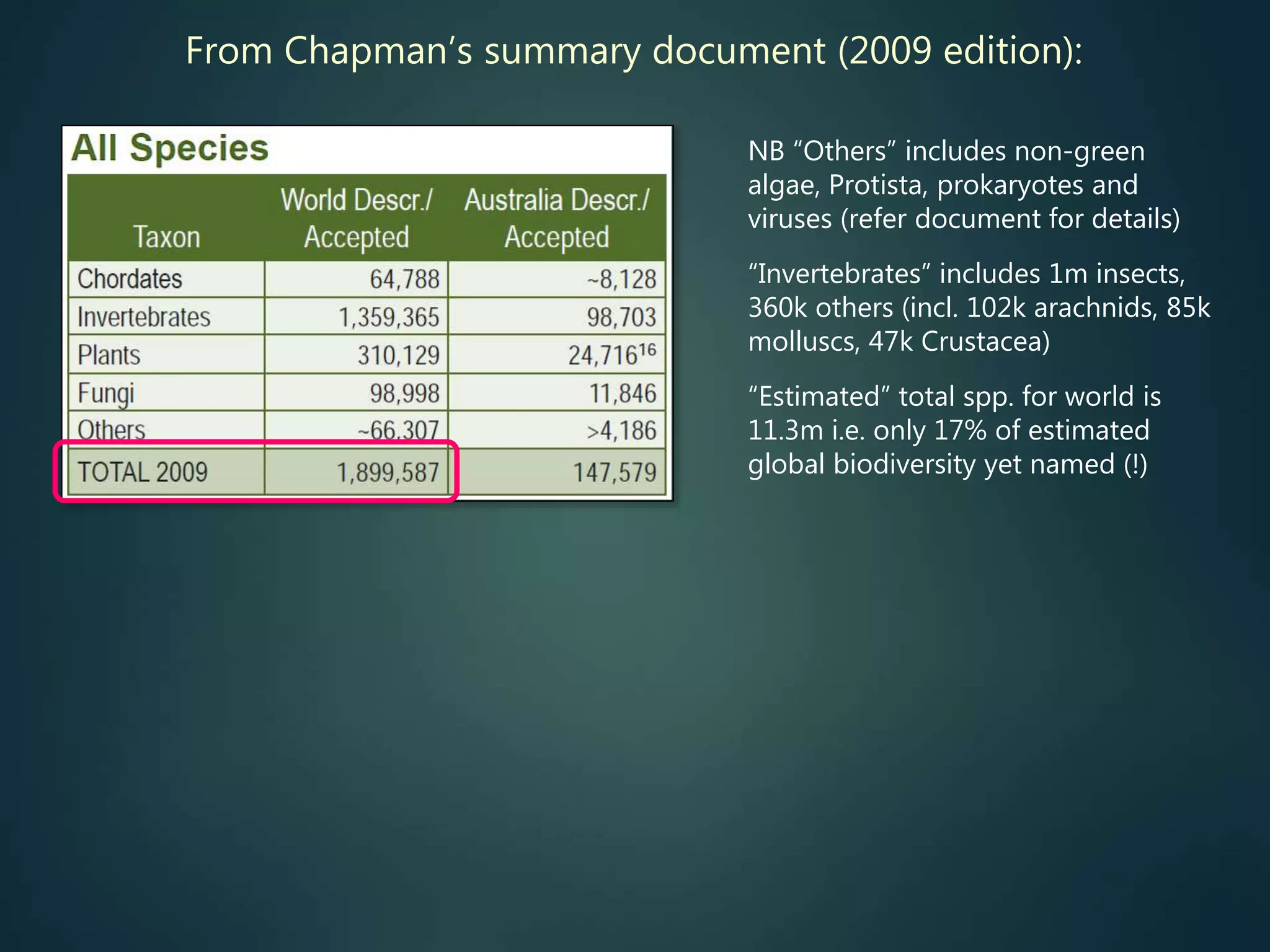
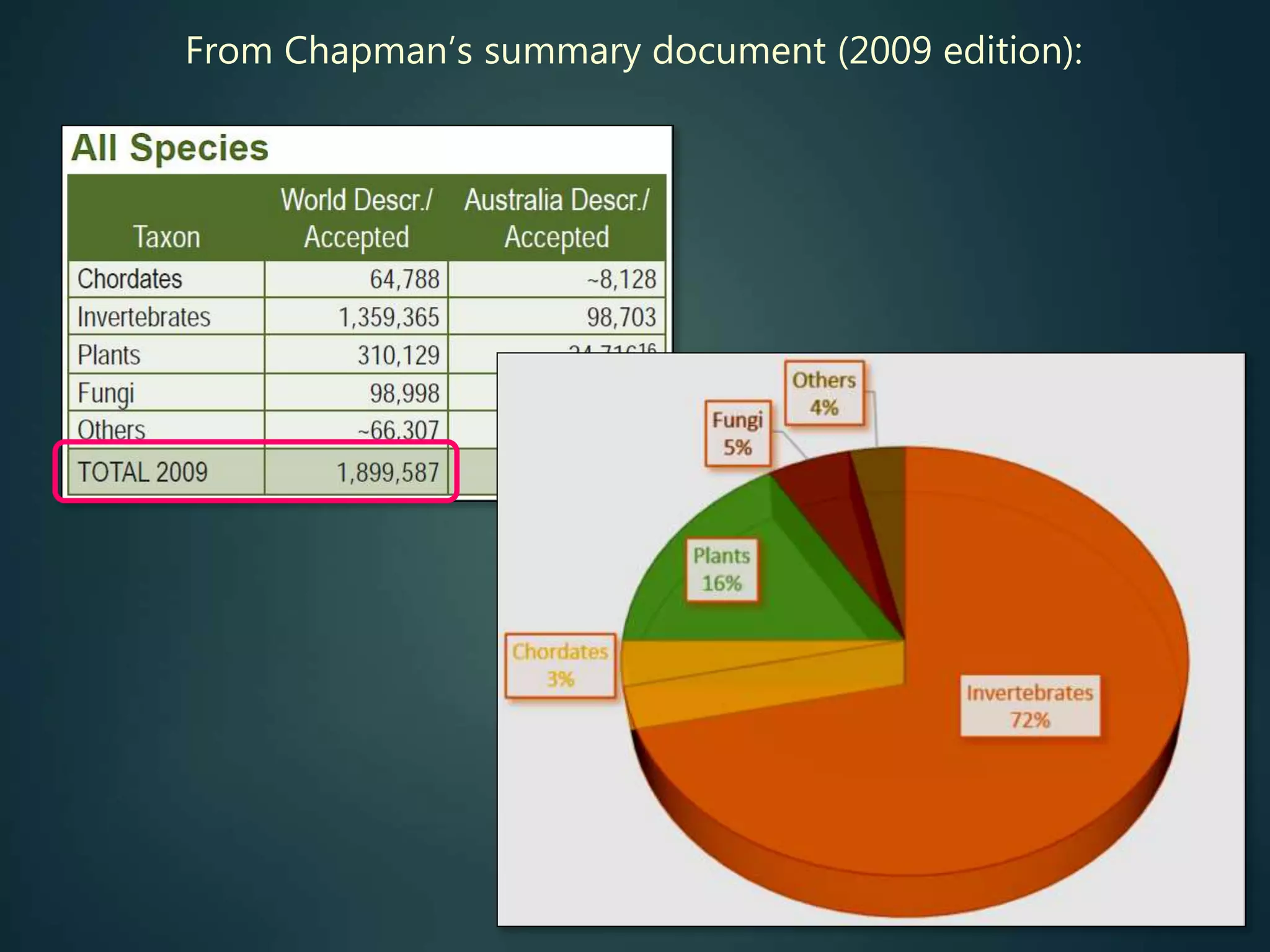
A global taxon inventory
up-to-date species lists,
synonymies, etc. (for all groups)
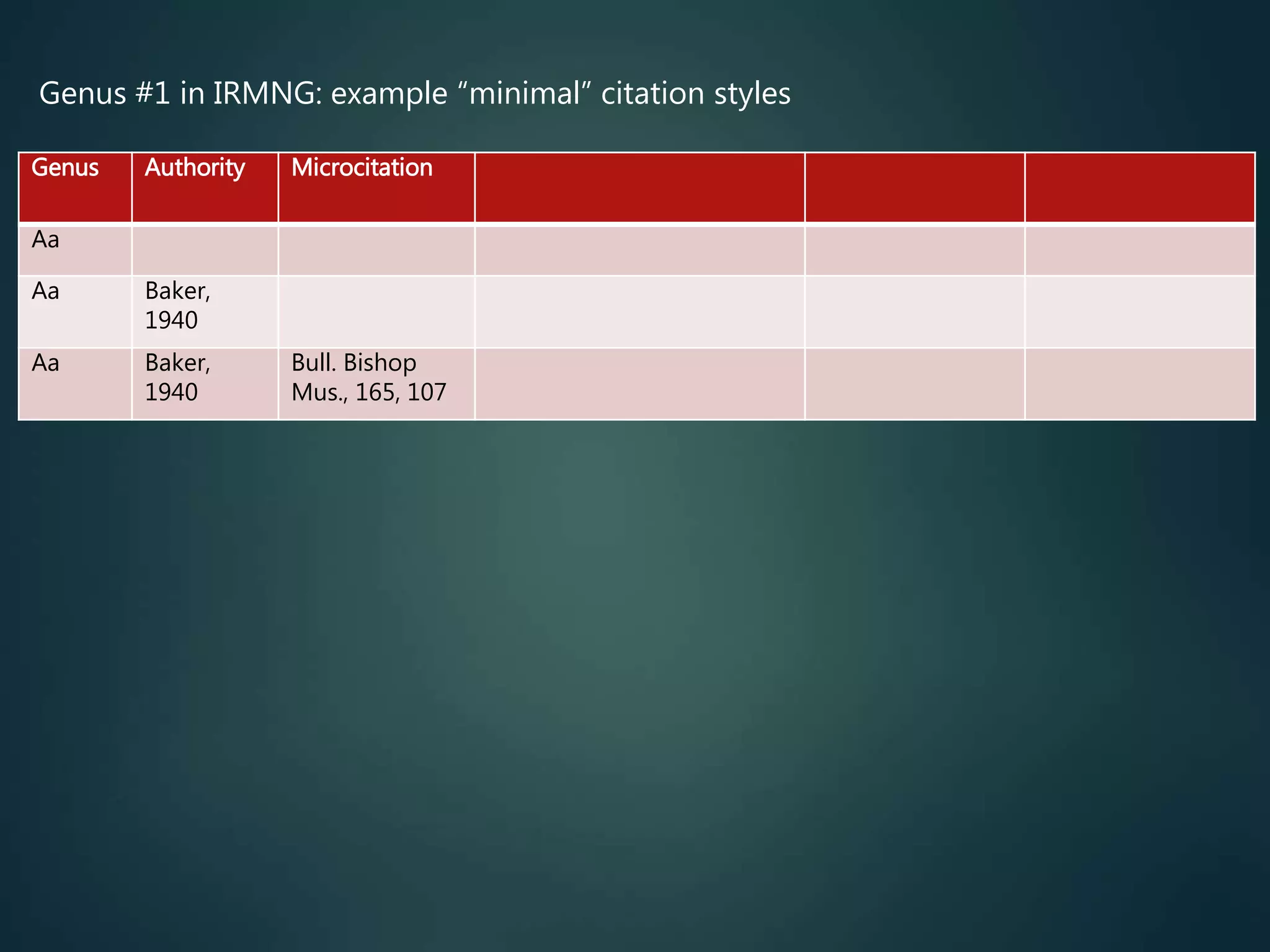
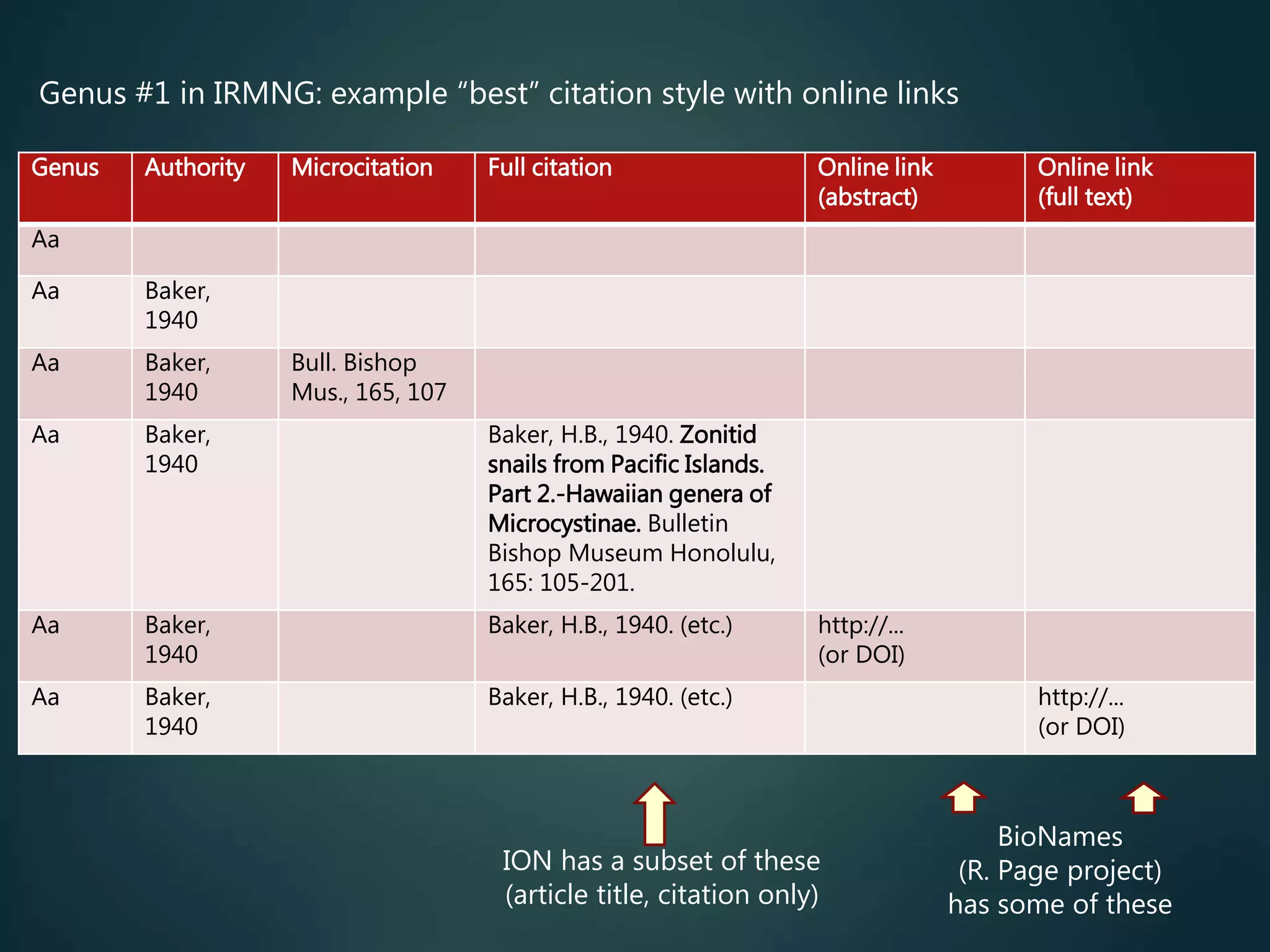
Citations, links to primary
literature
direct access to the primary
taxonomic literature (for all
described taxa), including full text
(preferably…)
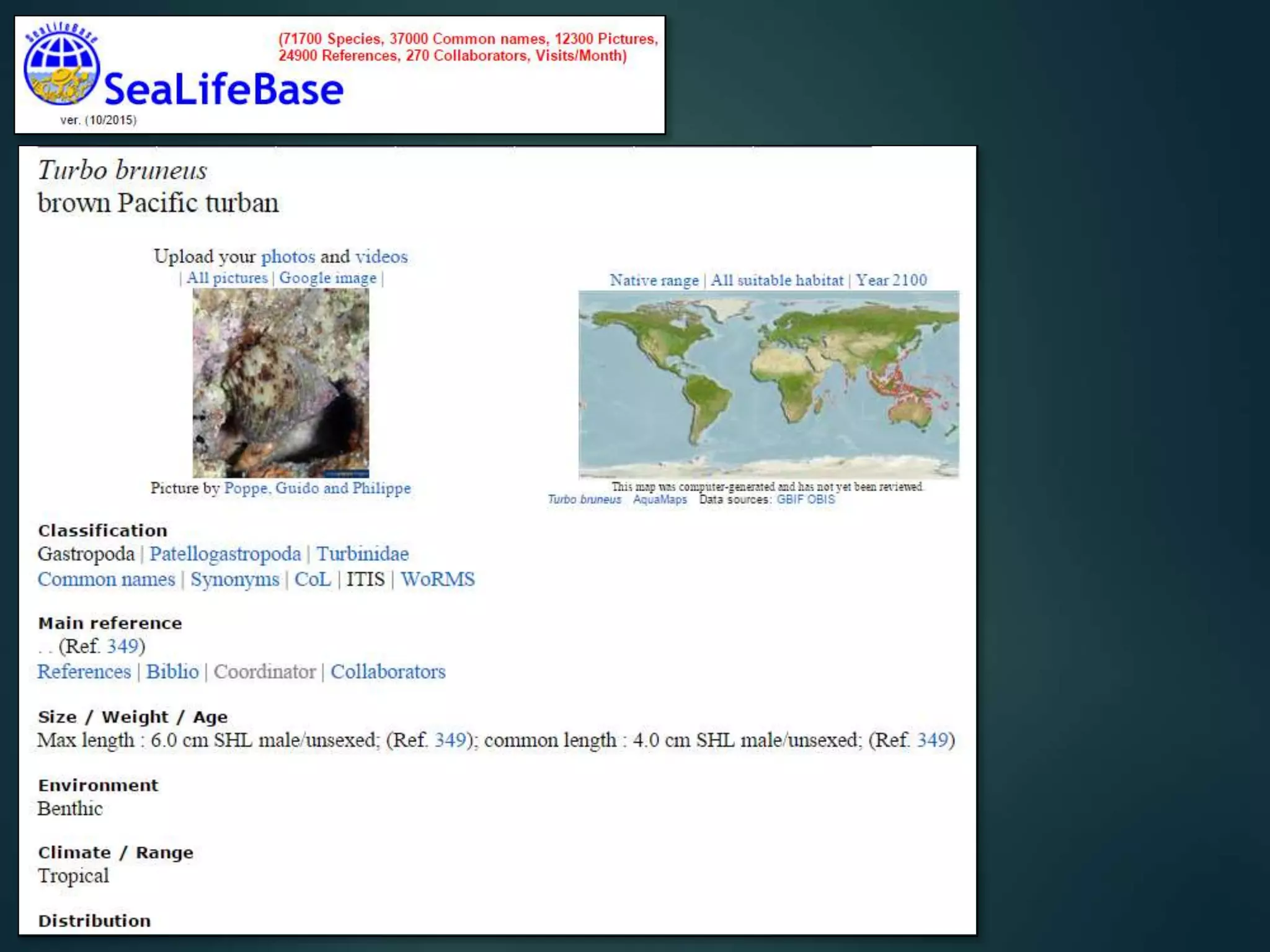
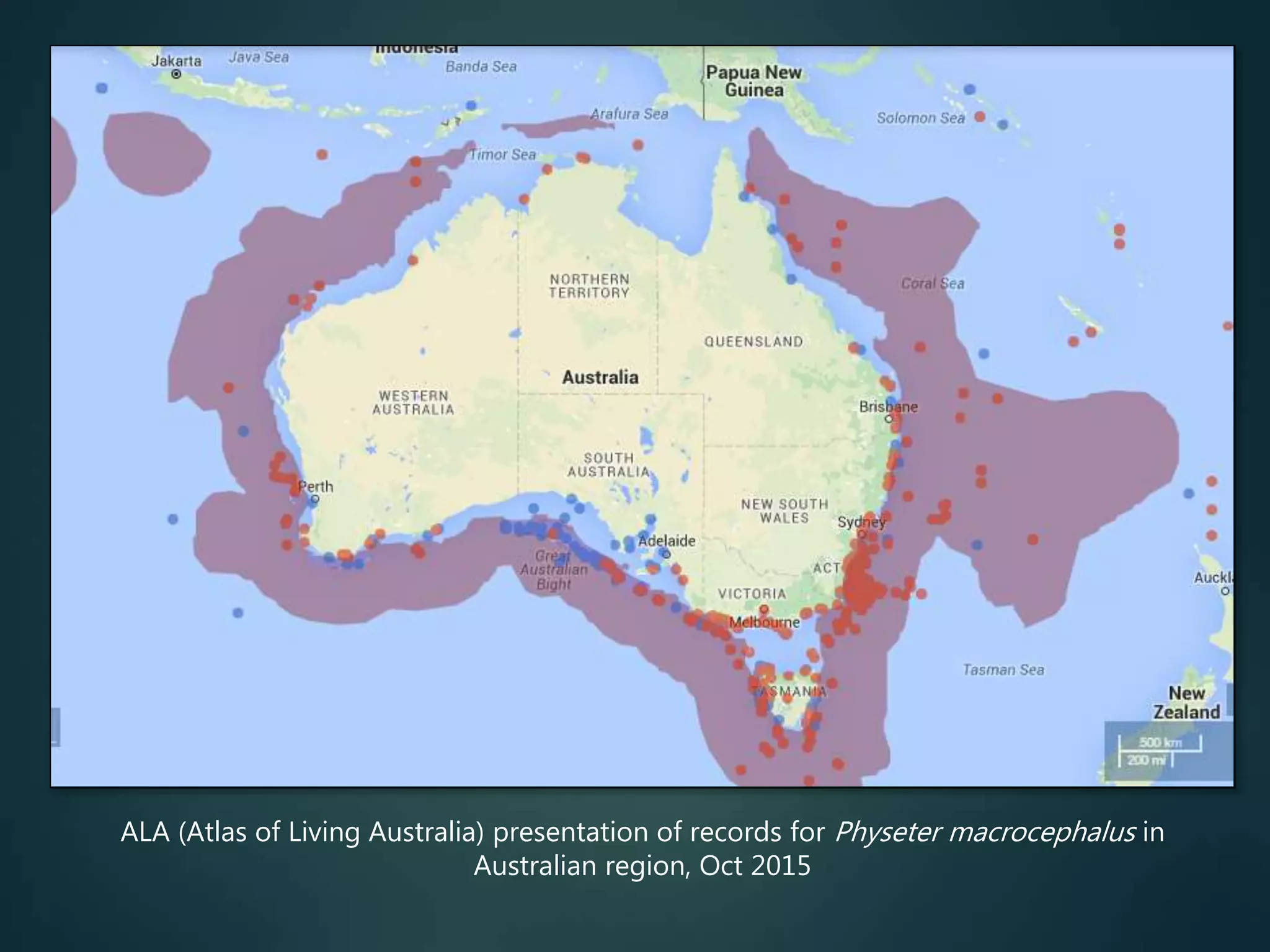
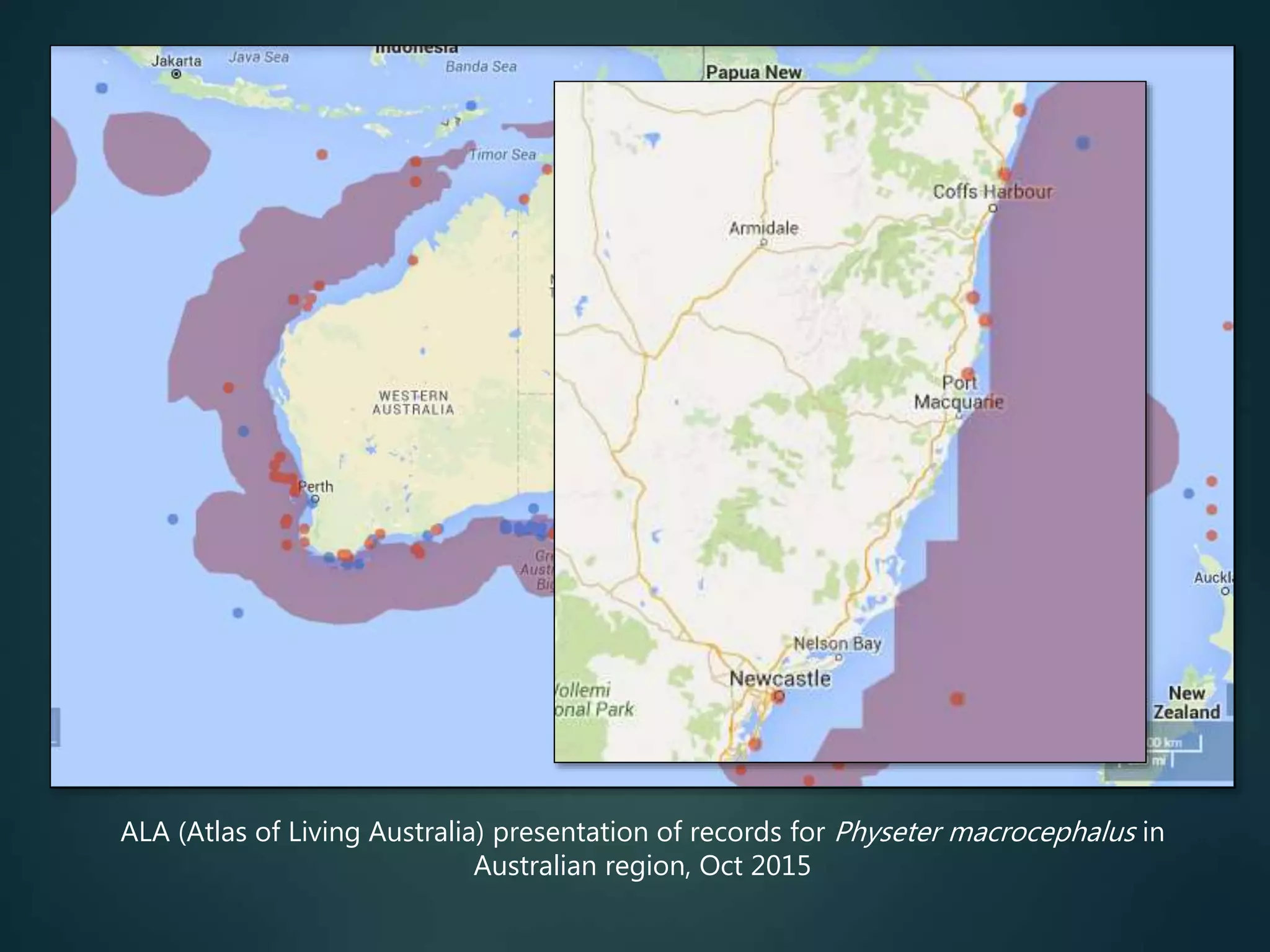
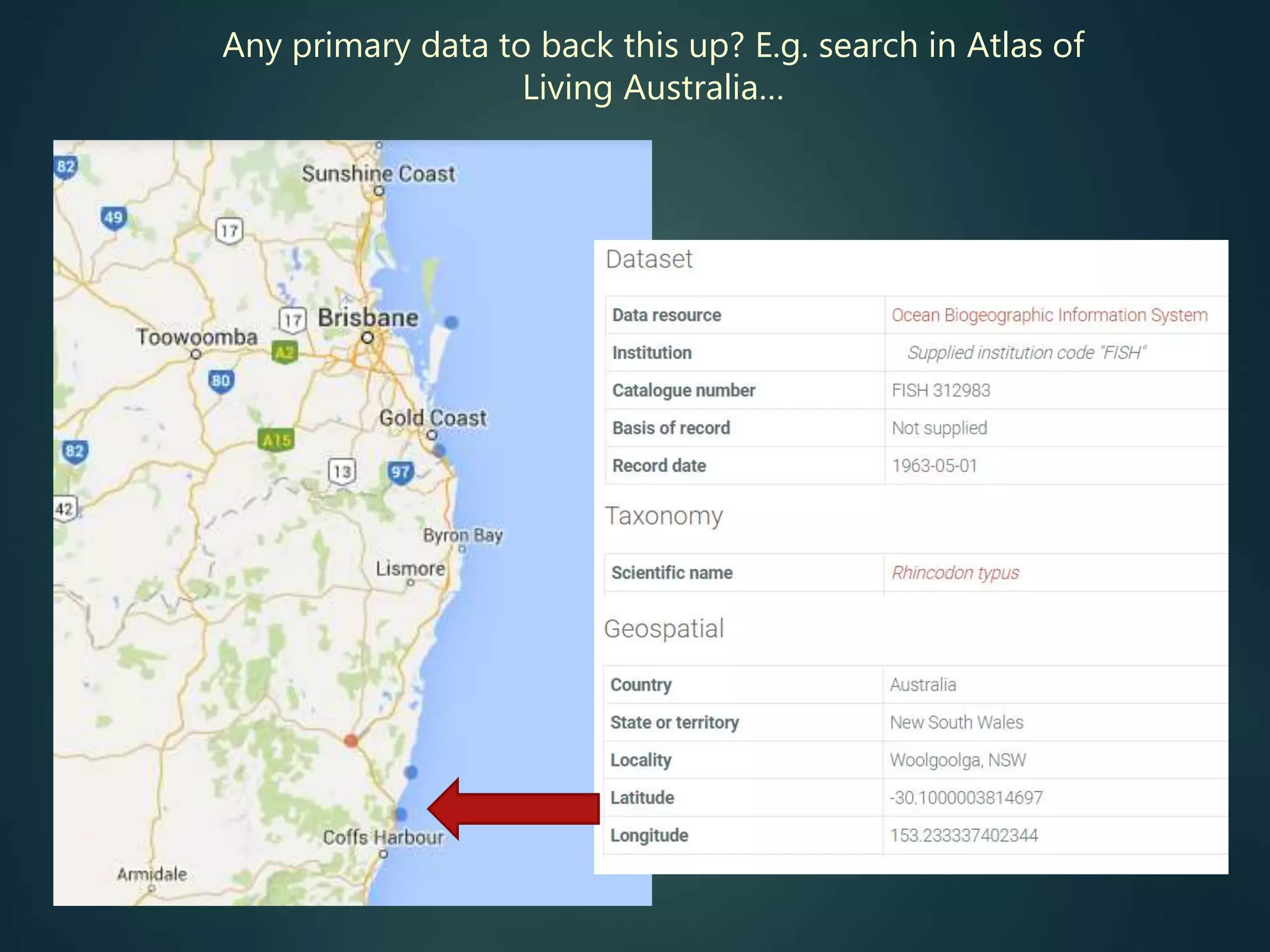
“All” georeferenced records
accessible, for all species
no need for individuals to
do the data aggregation
map local / regional / global
records, show details for any
data item
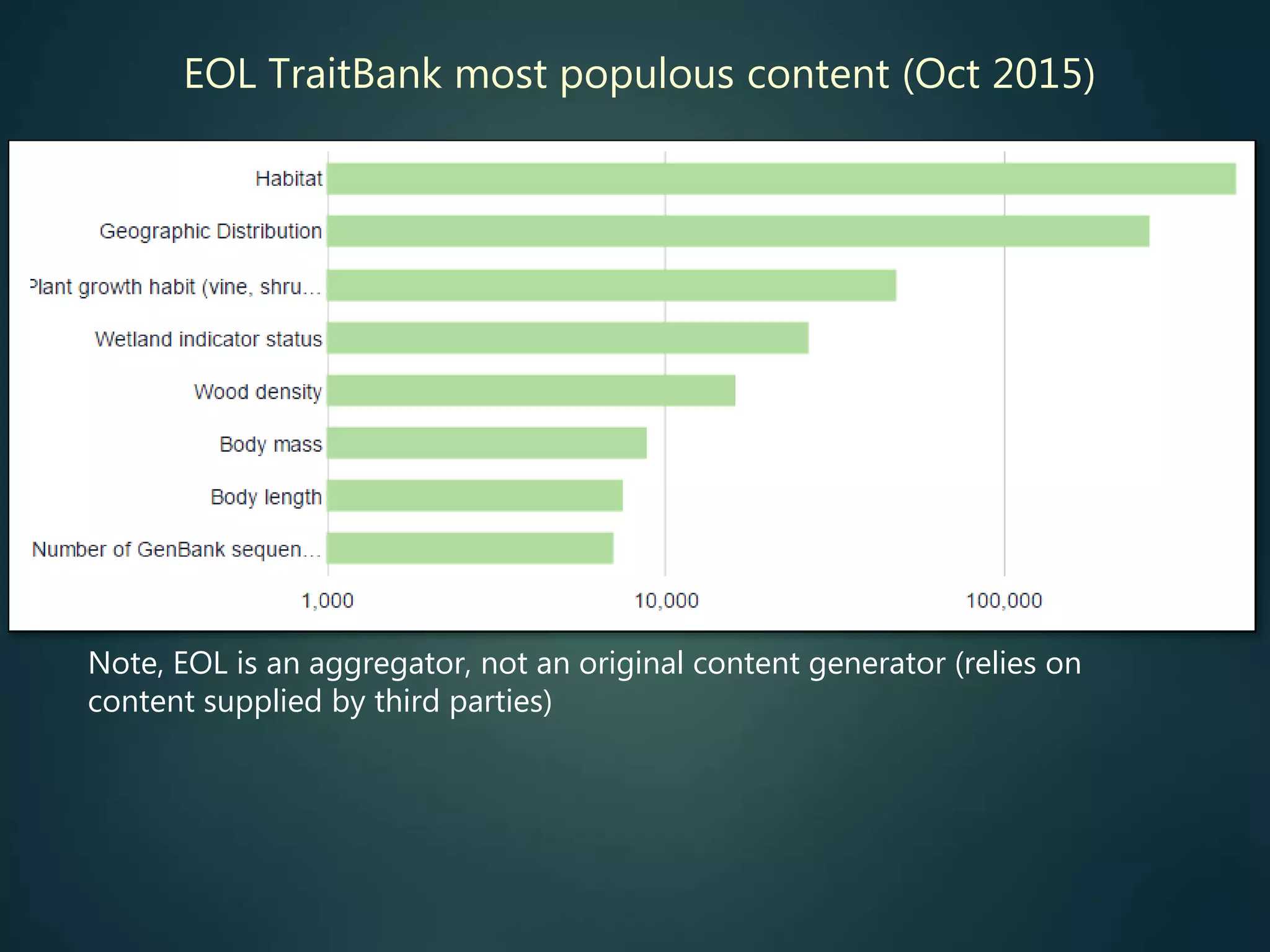
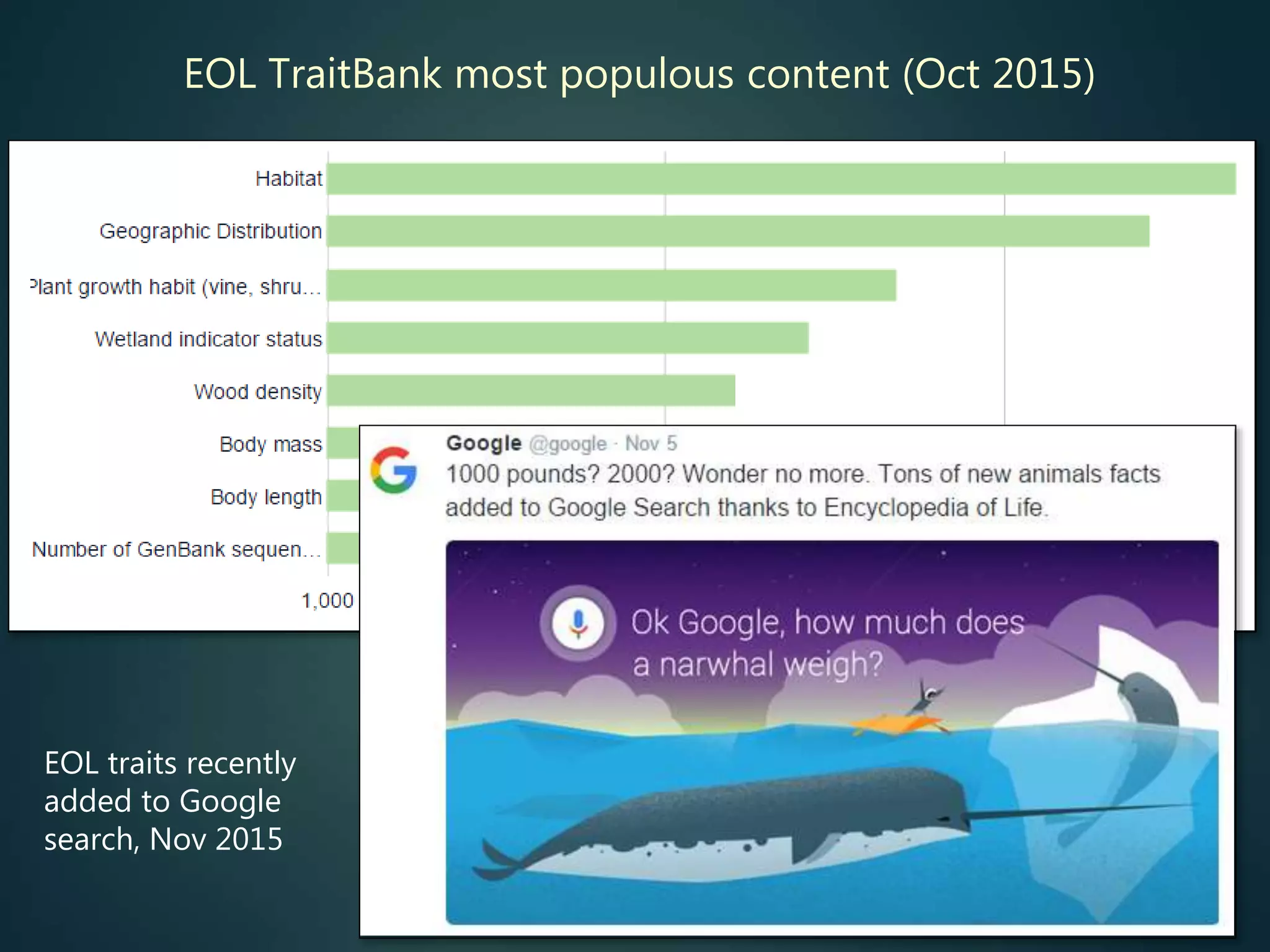
Indexes of taxon traits
e.g. to support sort /
filter / group by…
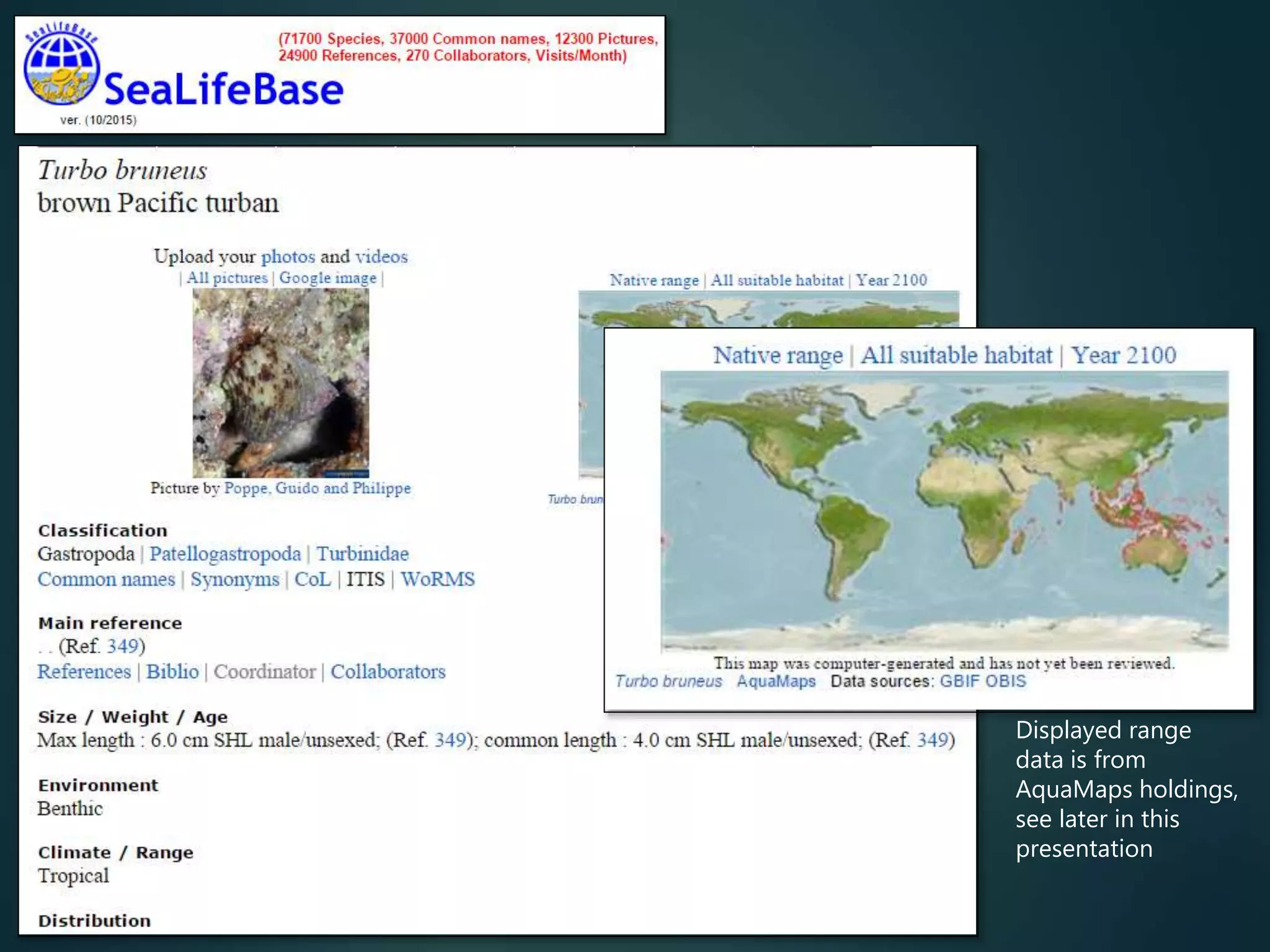
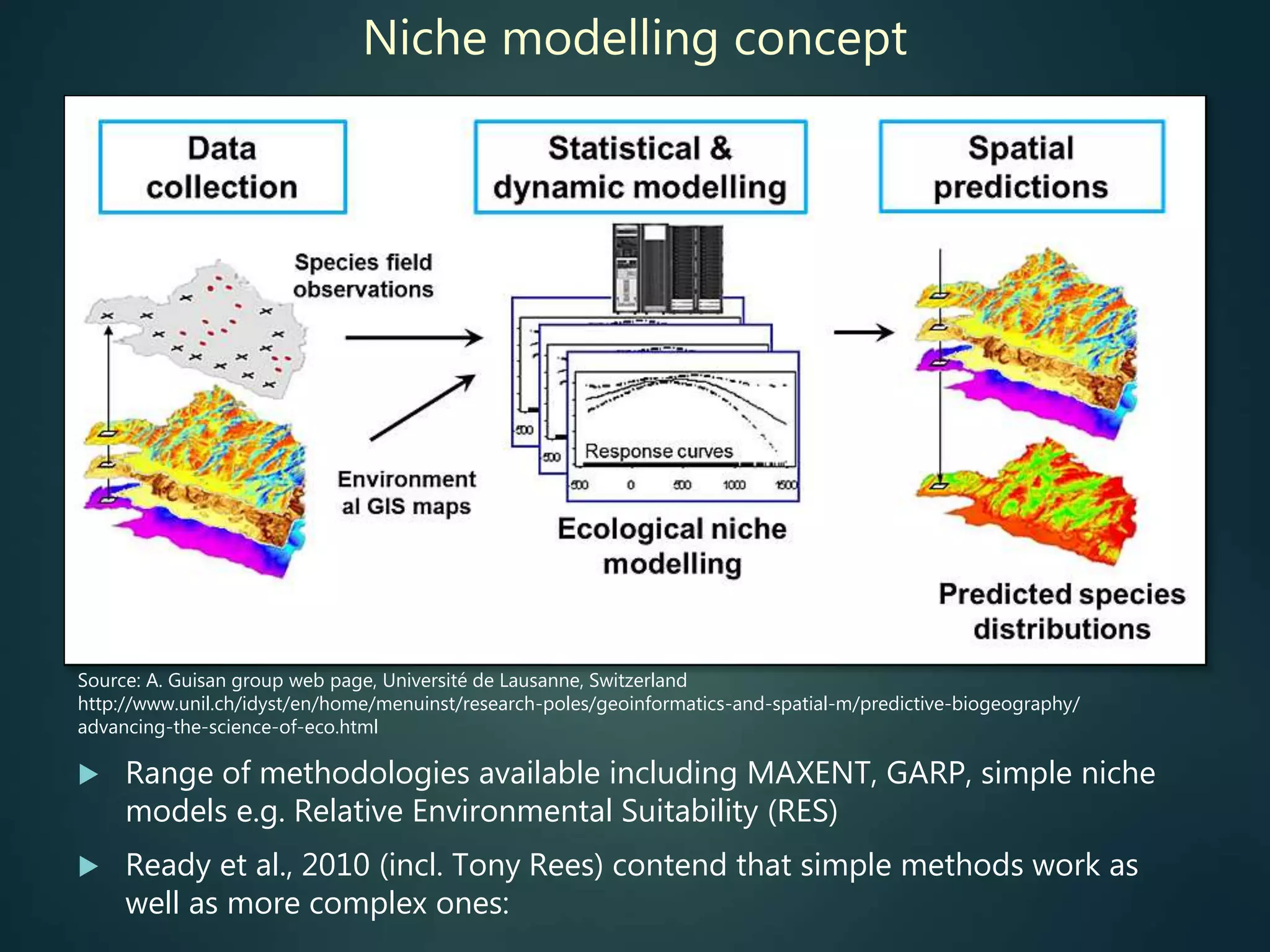
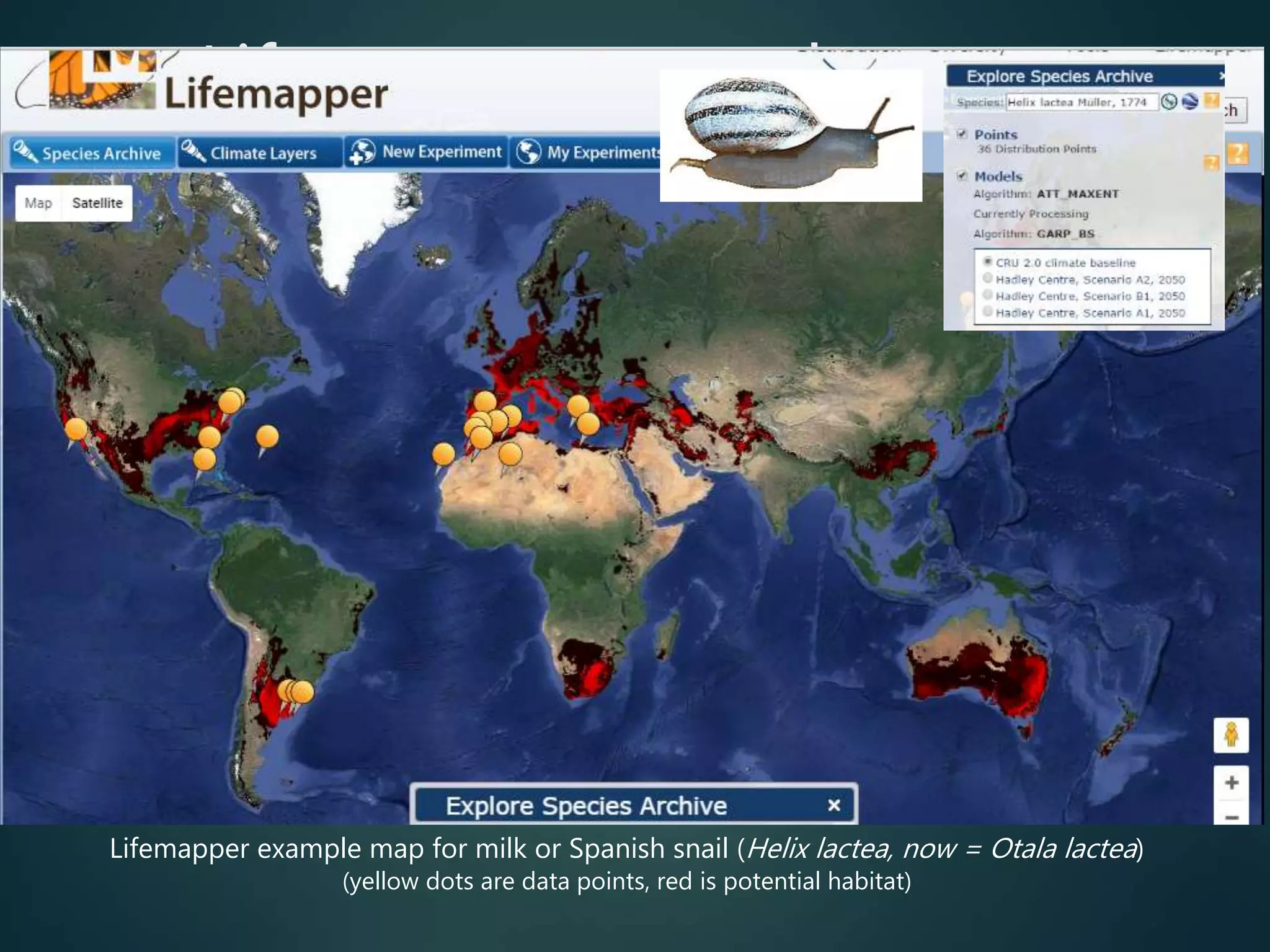
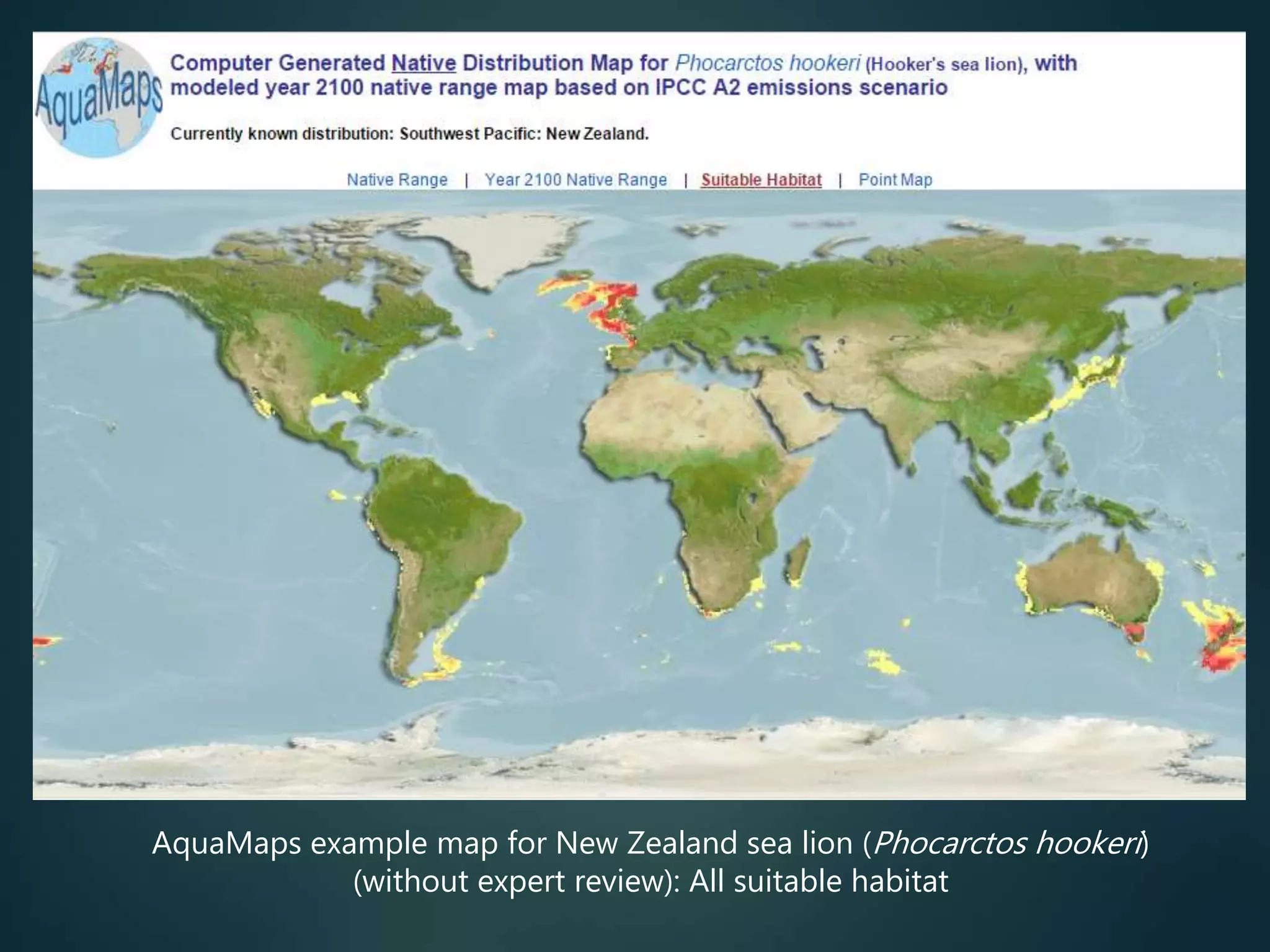
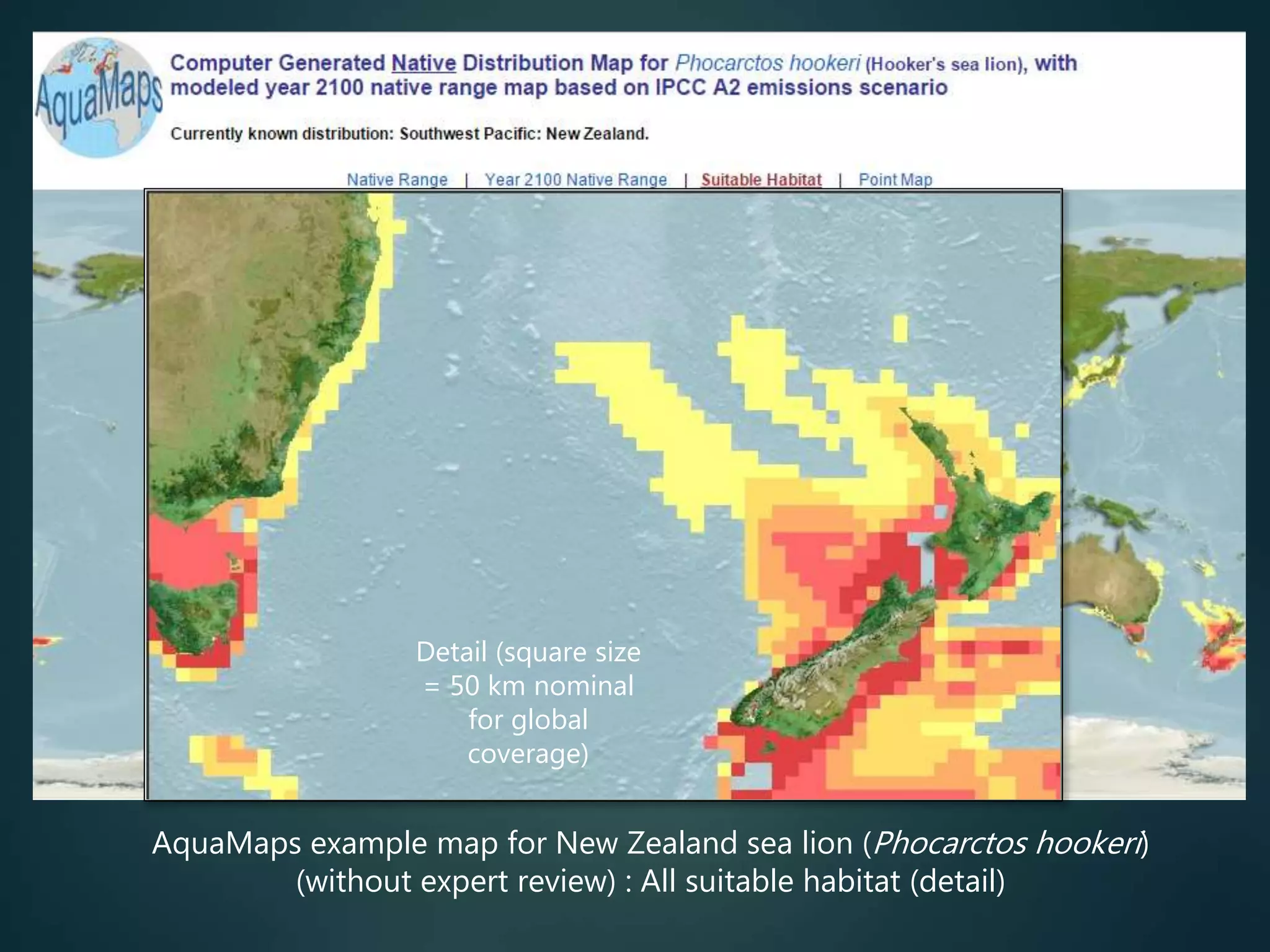
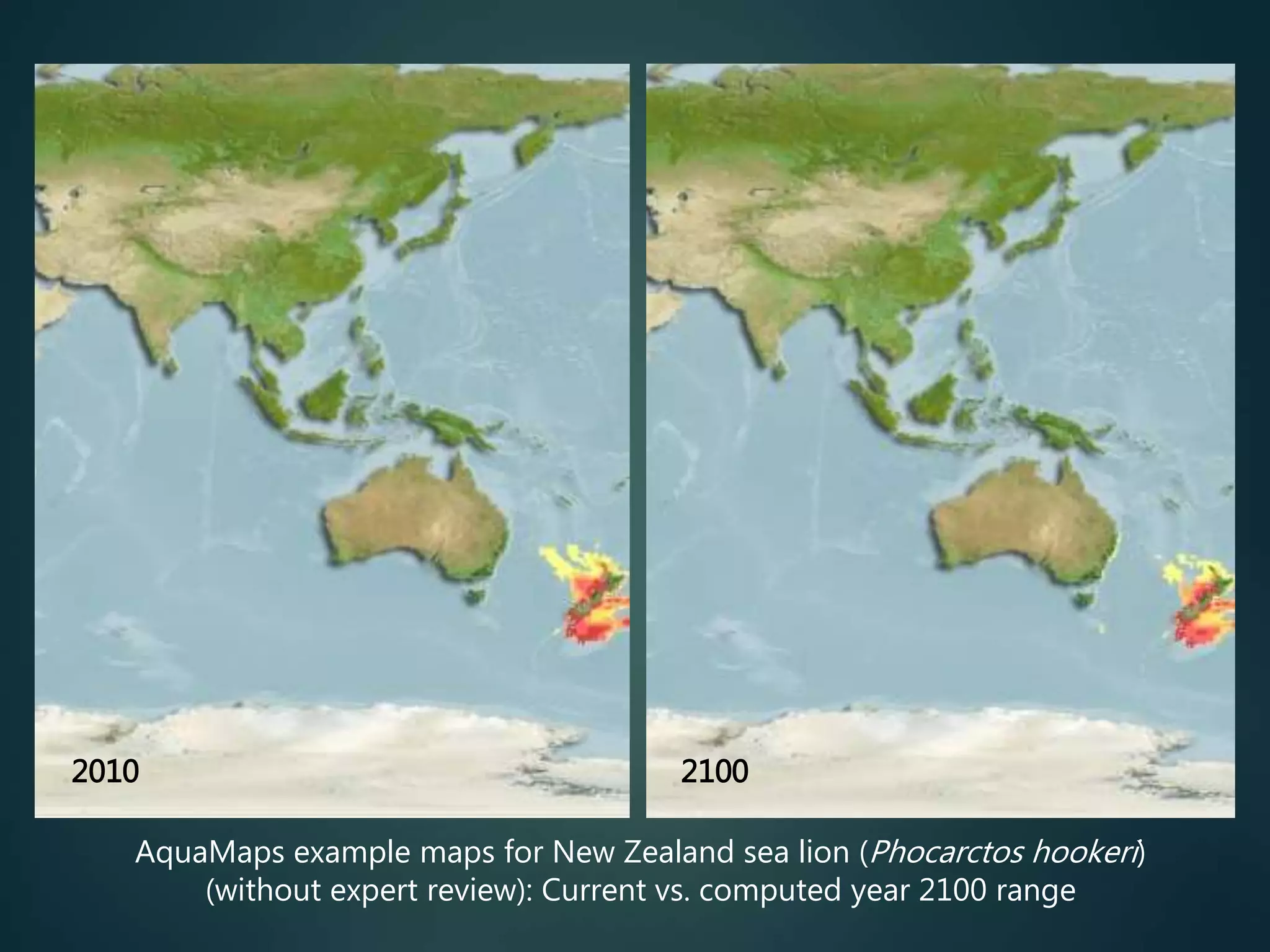
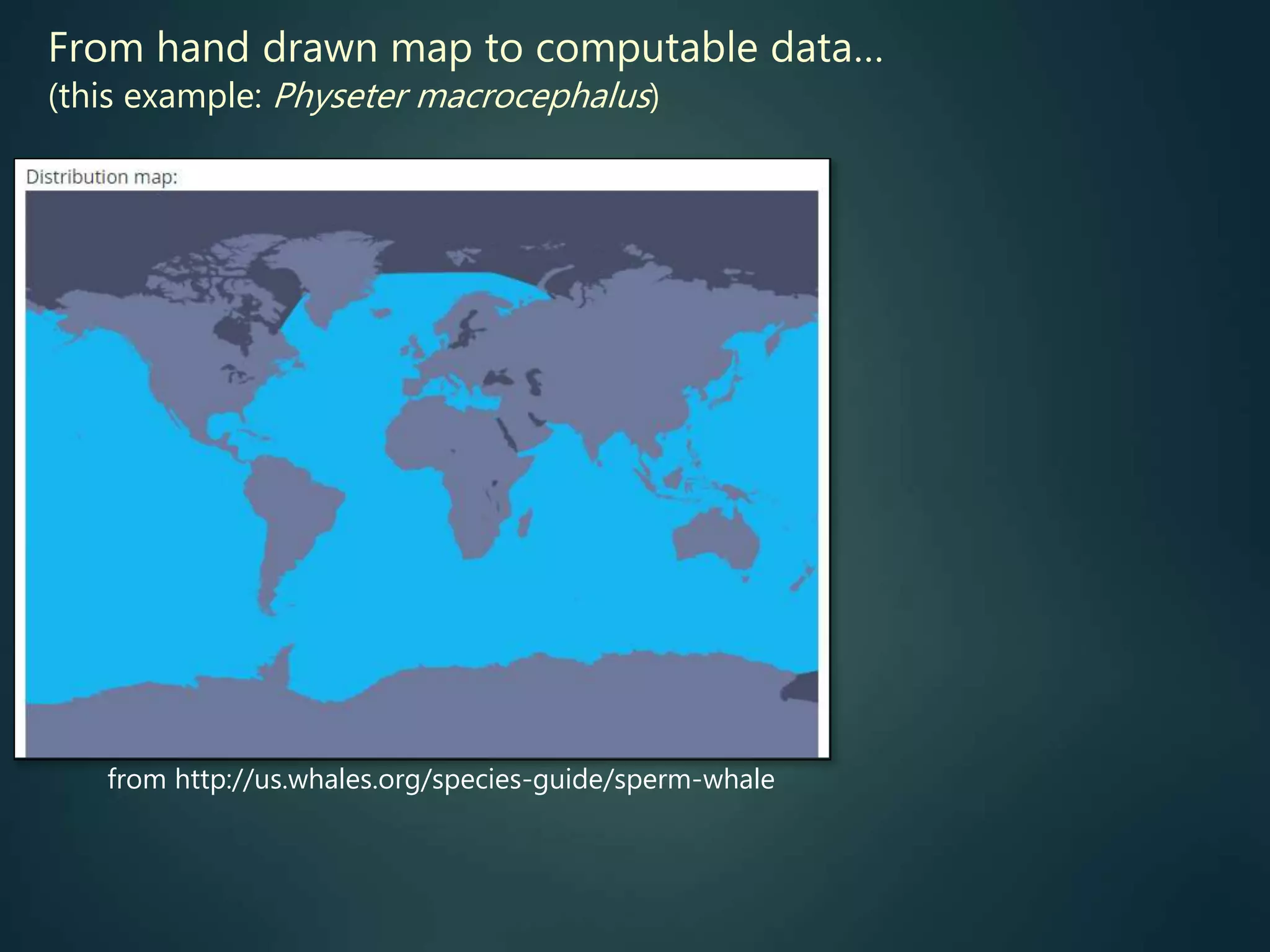
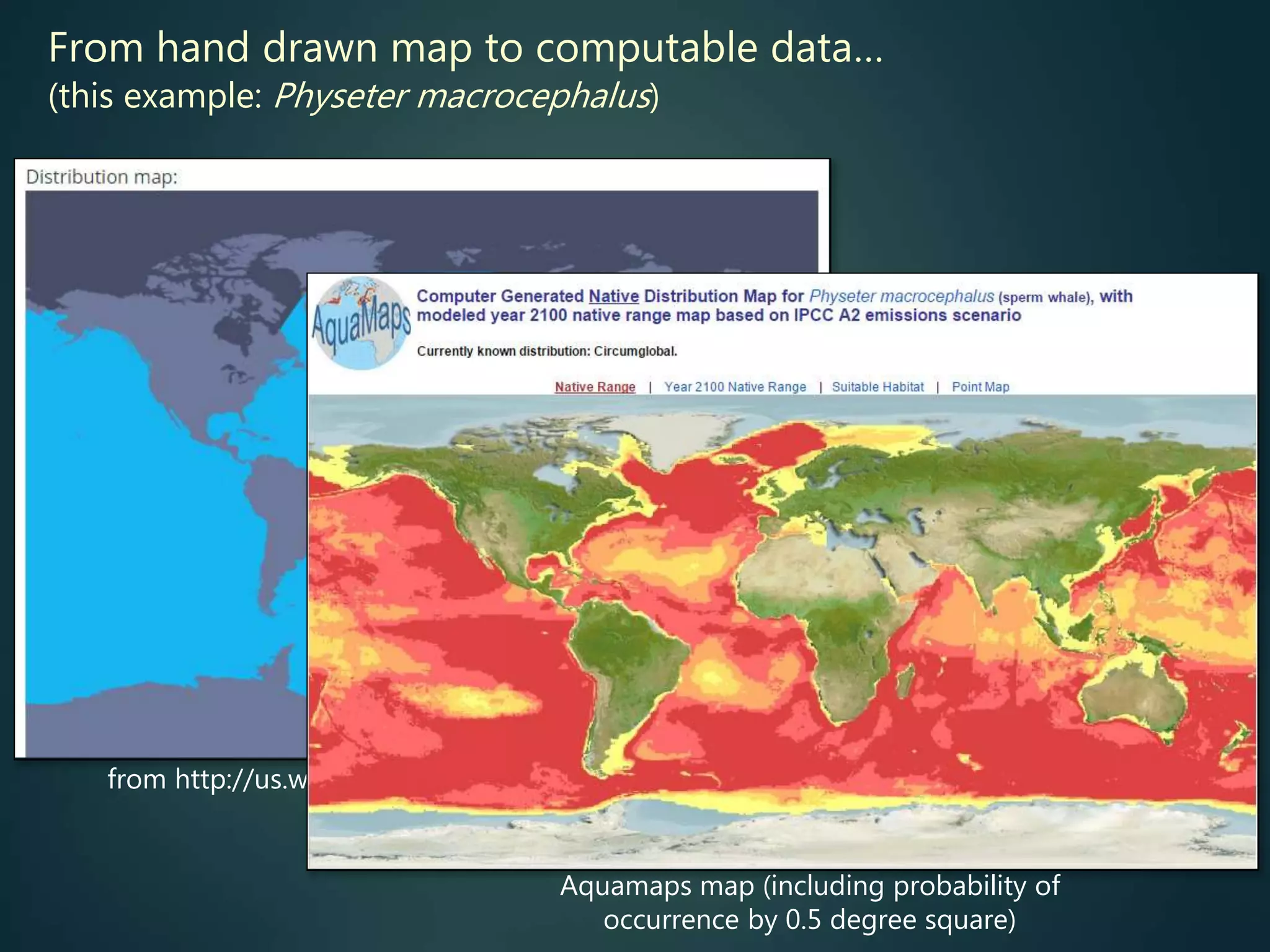
Predictive mapping / computed
range maps for all taxa
fill sampling gaps via niche
modelling, to produce
comprehensive global species maps Plus more (phylogenies,
illustrations, genetics,
descriptions, keys…)](https://image.slidesharecdn.com/rees-10yrsglobalbiodiversitydatabases-151126225732-lva1-app6892/75/10-years-of-global-biodiversity-databases-are-we-there-yet-2-2048.jpg)







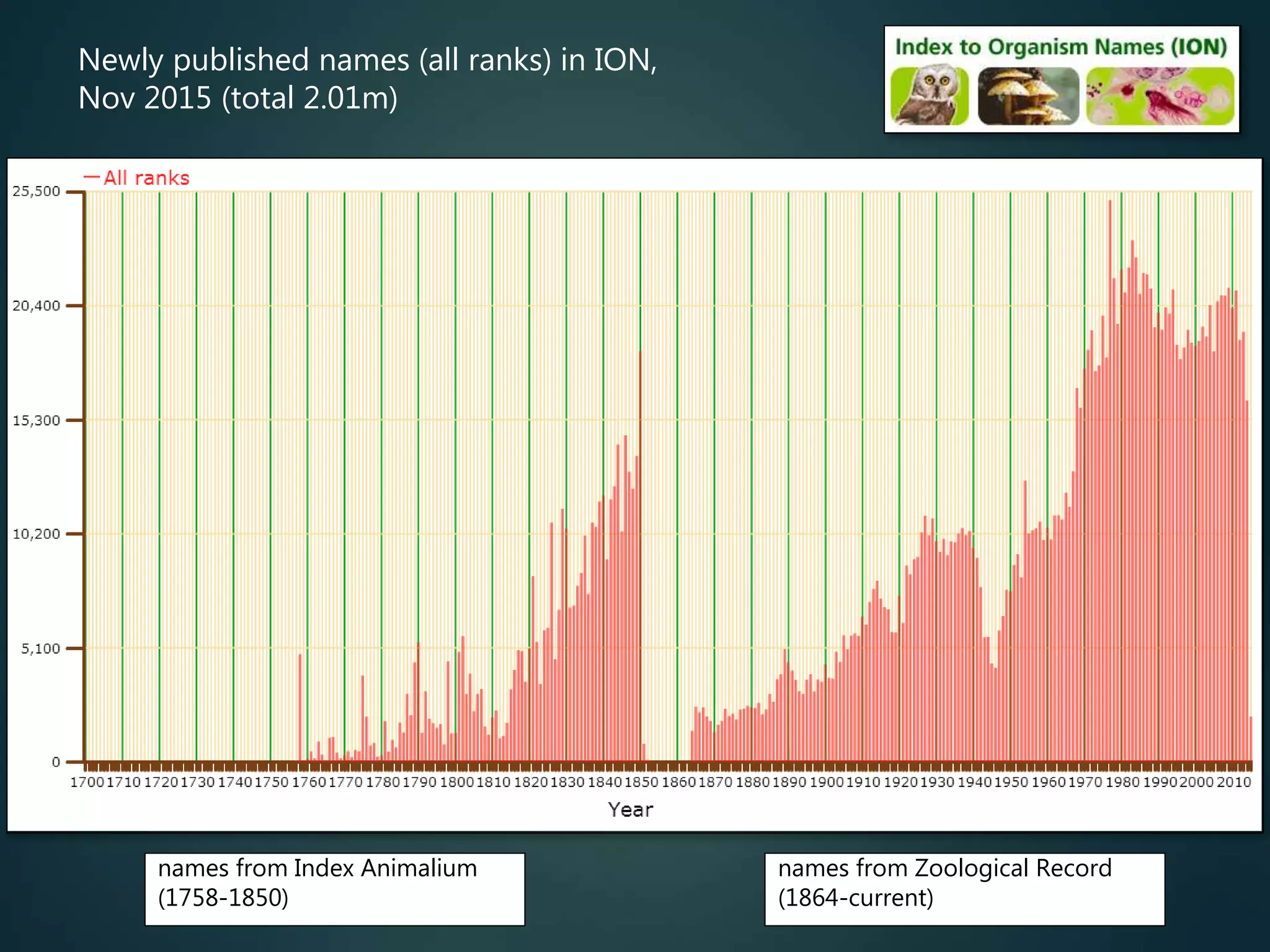

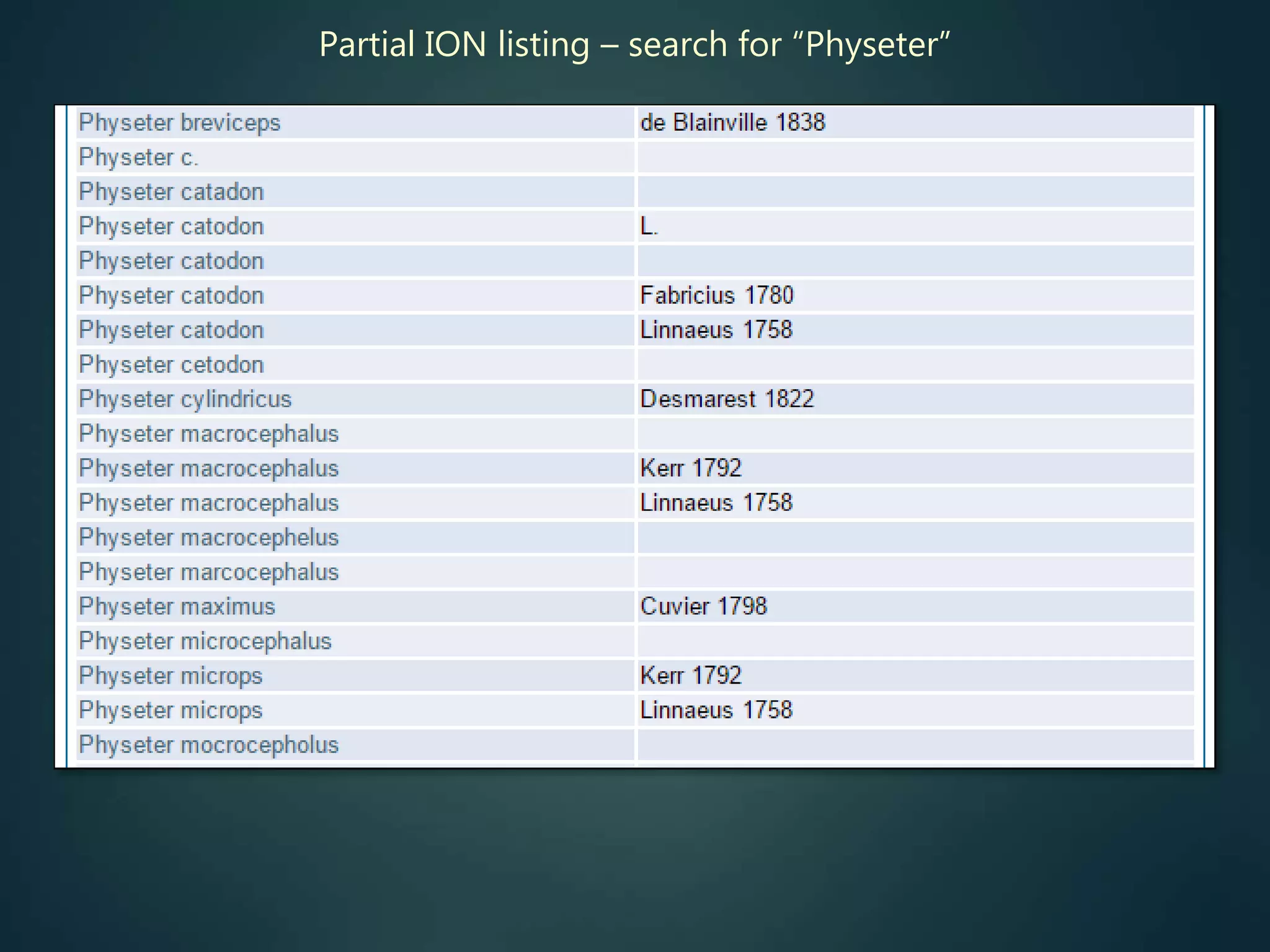
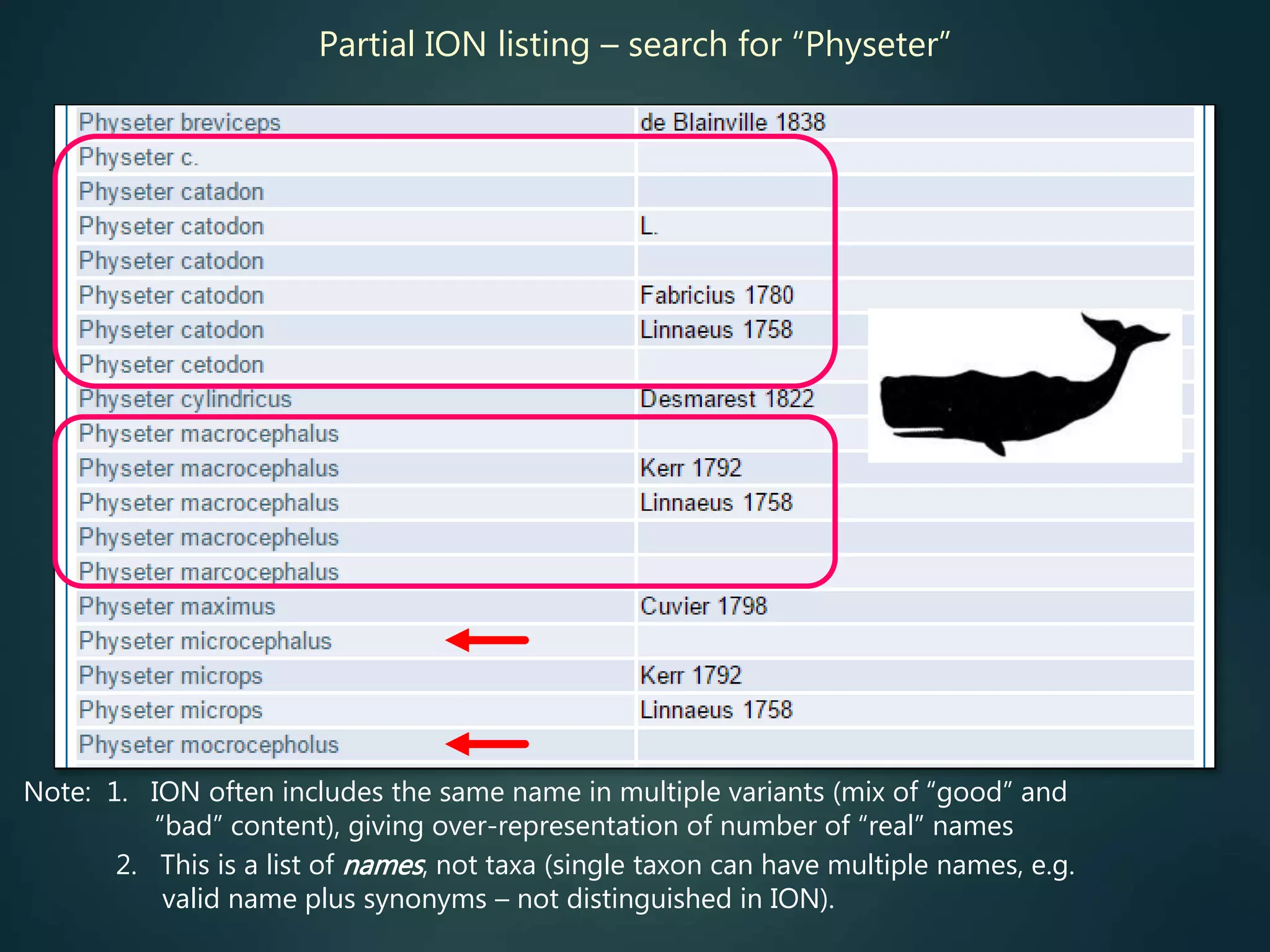





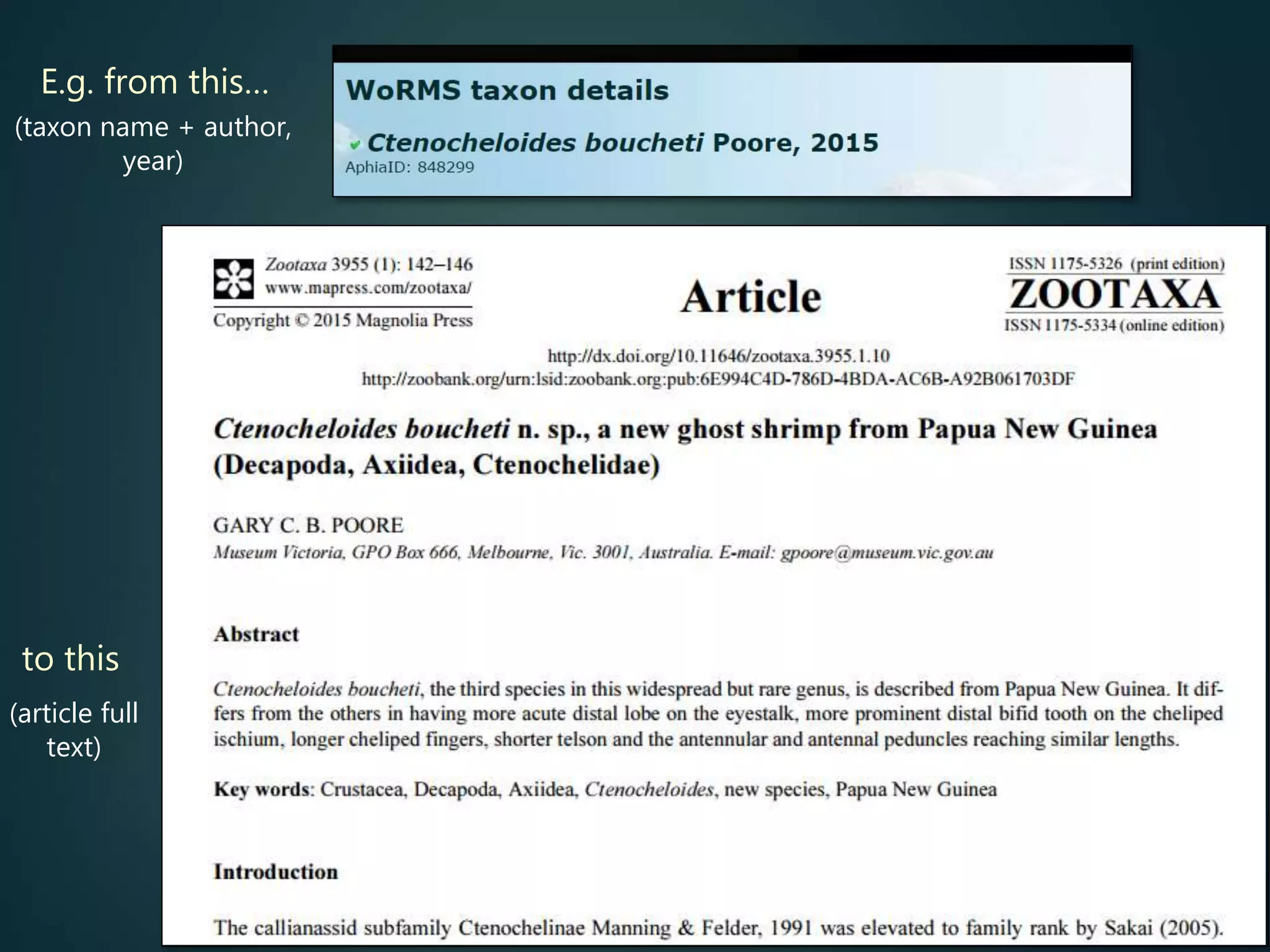


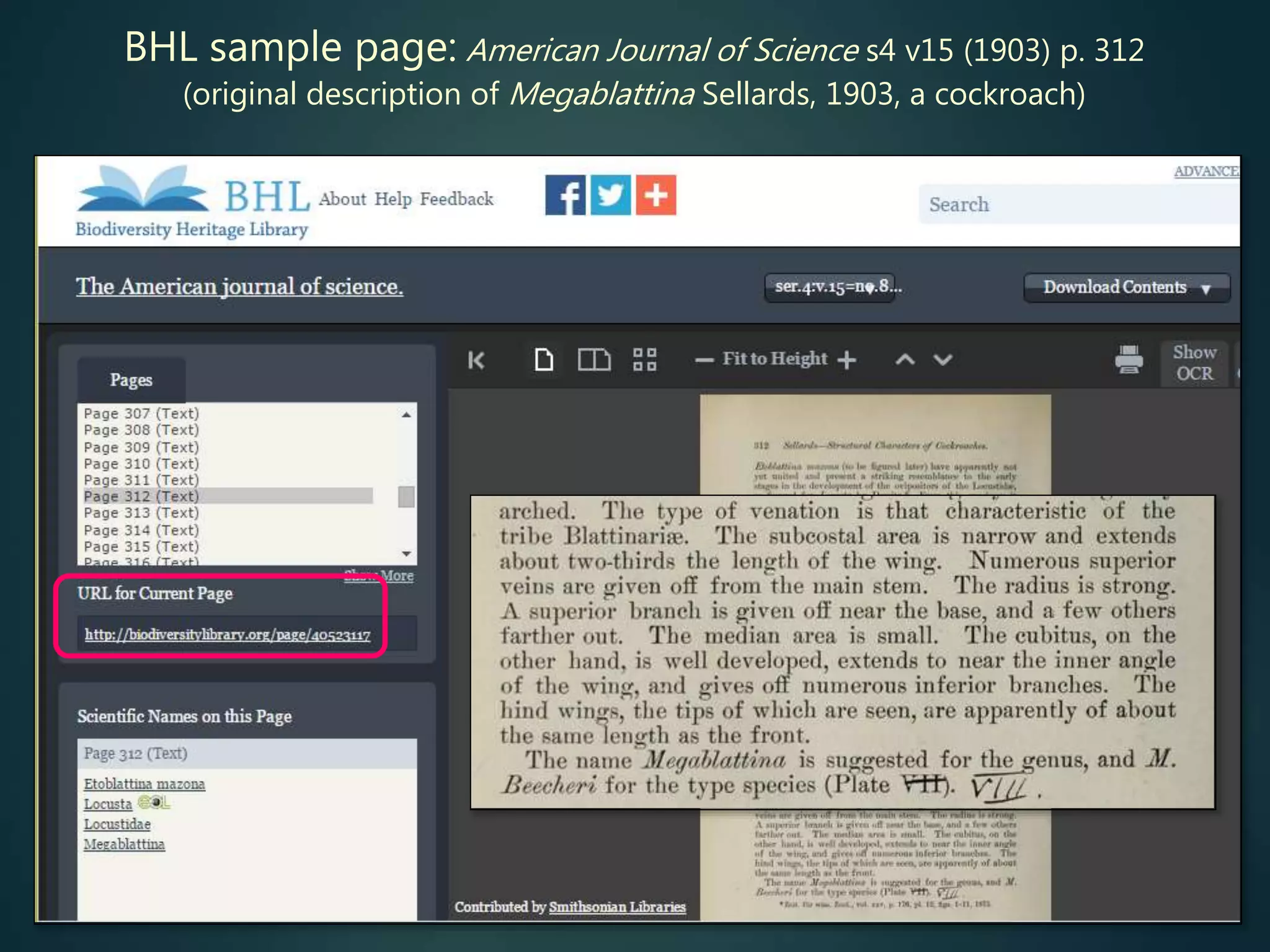
![Online access to scientific literature – 3
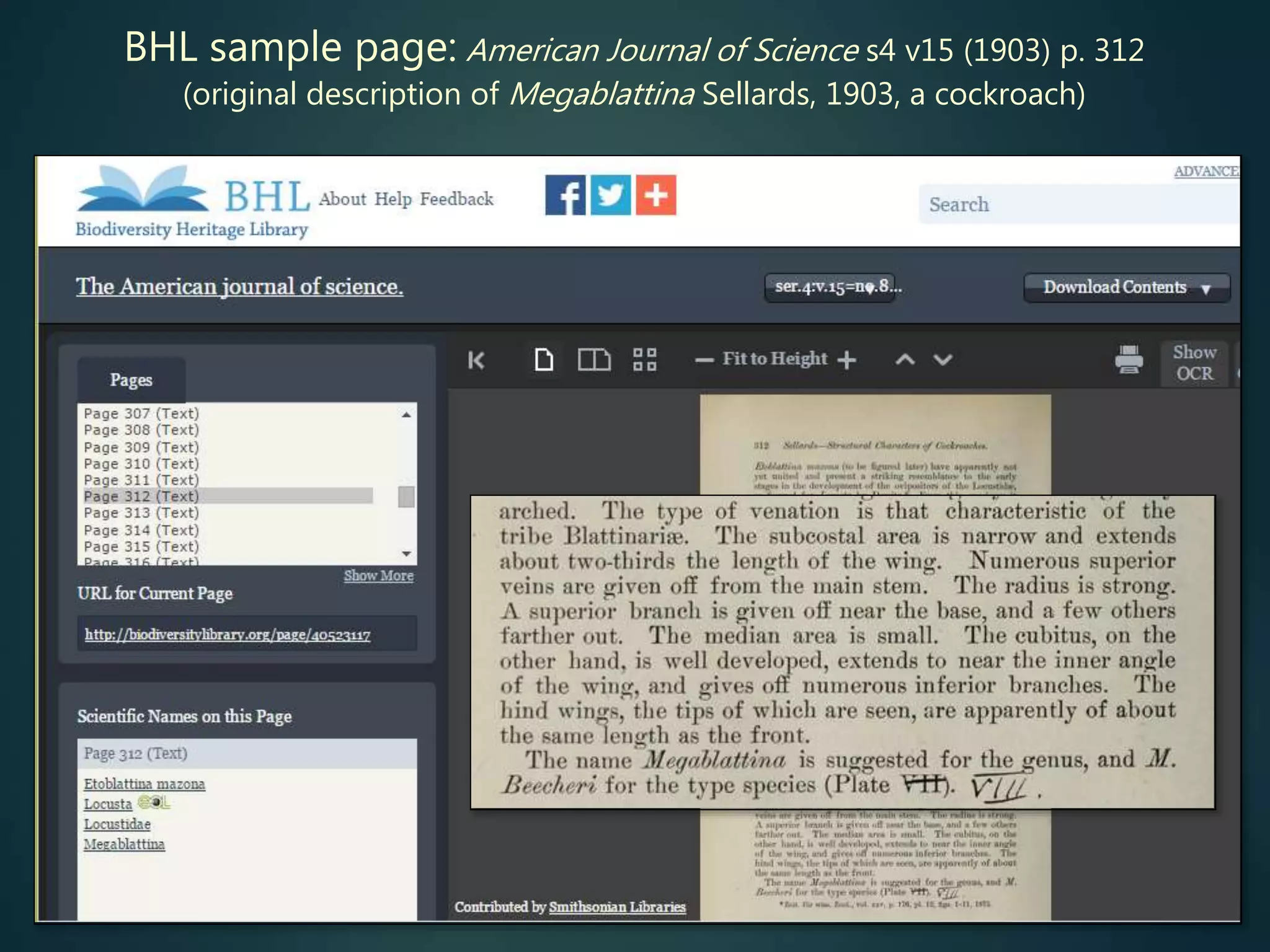
Biodiversity Heritage Library (BHL) is scanning older literature (esp.
pre-1923) and placing online
limited subset indexed by article title, otherwise (all) indexed by journal
and page no. (then has BHL page ID – can link to that)
search can be initiated by journal title, volume + page (if already
known)
can also search by taxon scientific name – but some instances will be
missed (BHL OCR [optical character recognition] is less than 100%
reliable)
this author’s experience looking for initial publication instances of
older names – success in around 1/3 of cases (not too bad), however
requires manual search (time consuming)
ideally, original description page links should be compiled somewhere
for others to re-use (not currently done on any scale)](https://image.slidesharecdn.com/rees-10yrsglobalbiodiversitydatabases-151126225732-lva1-app6892/75/10-years-of-global-biodiversity-databases-are-we-there-yet-27-2048.jpg)

![In summary: online [open] access available to subsets of article titles
> abstracts > full text in decreasing proportions
No single comprehensive source of online refs. available at
this time, users must “mix and match” sources as available
Few direct links in current tax. databases to literature that is
online (some noteworthy exceptions)
Over 95% of taxonomic literature pre-dates year 2000 starting
point for DOIs
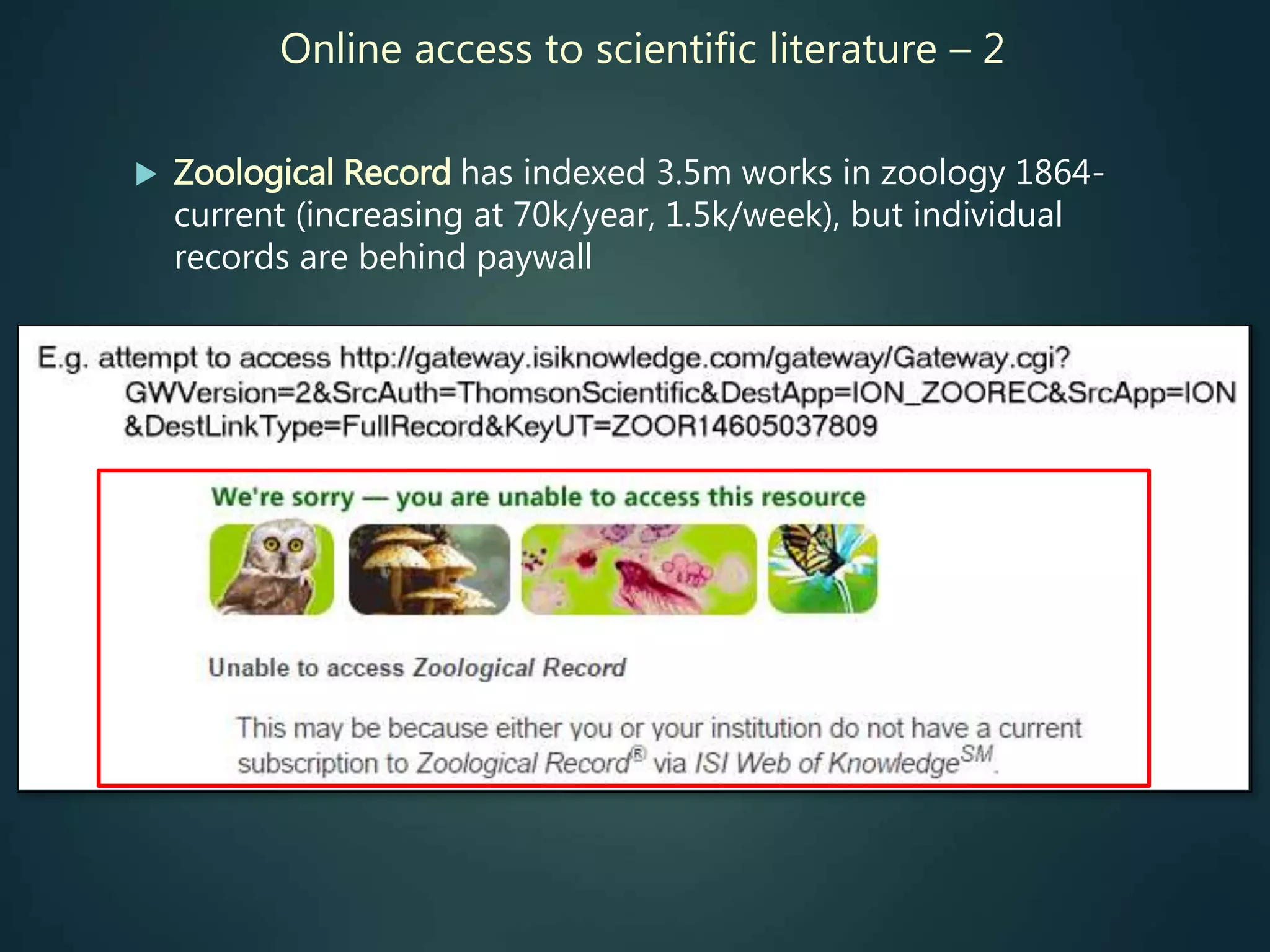
Most comprehensive indexes are currently commercial
products (behind paywalls), not much traction in “community
/ open access” equivalents as yet.](https://image.slidesharecdn.com/rees-10yrsglobalbiodiversitydatabases-151126225732-lva1-app6892/75/10-years-of-global-biodiversity-databases-are-we-there-yet-31-2048.jpg)