Genetic Analysis of Limb Malformations Using Microsatellite Markers
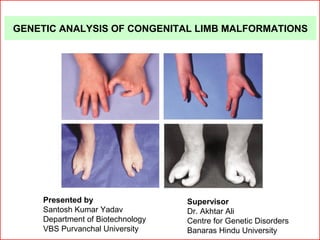
- 1. GENETIC ANALYSIS OF CONGENITAL LIMB MALFORMATIONS Supervisor Dr. Akhtar Ali Centre for Genetic Disorders Banaras Hindu University Presented by Santosh Kumar Yadav Department of Biotechnology VBS Purvanchal University
- 2. • Embryonic limb develop, as a limb "bud." Fibroblast growth factor (FGF) induces formation of an organizer, called the apical ectodermal ridge (AER) , which guide further development and controls cell death • Apoptosis is necessary to eliminate webbing between digits • Somites give to the axial skeleton, the lateral plate mesoderm generates the limb skeleton Normal Limb Development
- 3. (a) Progress zone model (b) Early specification model
- 5. Congenital Limb Malformations • genetic or environmental factor during embryogenesis • Clinically variable and genetically heterogeneous • Syndactyly, monodactyly, ectrodactyly, Brachydactyly
- 6. Split-Hand/Foot Malformation (SHFM) • Characterized by deep median cleft due to absent of central ray and fusion of remaining digits • Crab claw like hand (Ectrodactyly) • 1 in every 8,500 to 25,000 births accounting for 8-17% of all limb reduction defects • Clinically variable and genetically heterogeneous • syndromic (e.g. EEC) and non-syndromic forms(SHFLD) • autosomal dominant, autosomal recessive, X-linked incomplete penetrance, variable expressivity
- 7. SHFM Loci SHFM1 7q21 -- SHFM2 Xq26.3 -- SHFM3 10q24-q25 -- SHFM4 3q27 TP63 gene SHFM5 2q31 -- SHFM6 12q13.11 WNT10B gene
- 8. ObjectiveObjective Genetic Analysis of Congenital Limb Malformations Determination of Linkage of 10q25 locus with SHFM using microsatellite markers in a multiplex family Cytogenetic analysis of congenital limb malformations
- 9. Family based linkage study of congenital limb malformation DNA isolation from members of a multiplex family PCR of microsatellite markers and capillary electrophoresis Analysis of inheritance of alleles Determination of Linkage of 10q25 locus with SHFM using microsatellite markers in a multiplex family
- 10. II-1 I-2I-1 II-1 II-2 III-3 III-4III-2III-1 IV-2IV-1 V-1 V-2 V-3 IV-4IV-3 V-5V-4 IV-5 IV-6 IV-7 V-9V-8V-6 V-7 V-10 V-11 VI-1 VI-2 IV-8 IV-9 V-14 V-15V-12 V-13 IV-10 Pedigree of the Family Pedigree show autosomal dominant inheritance
- 11. Genomic DNA Quantification By agarose gel electrophoresis and Spectrophotometer (Nanodrop®) λ DNA 1 2 3 4 5 6 7 8 9 Photograph showing thegenomic DNA electrophoresisin 0.8% AgaroseGel .
- 12. Microsatellite markerMicrosatellite marker • short tandem repeat sequence of nucleotide such as ‘CACACACA’ and occur in non-coding DNA • polymorphic and variable in population • applications in genetics such as mapping genes in inherited disease • detect microsatellite is to design PCR primer that is unique to one locus in genome and base pair on either side of repeated portion • Two PCR primer (F and R) are designed to microsatellite region
- 13. Microsatellite marker used DNA Marker Dye Colour D10S192 NED YELLOW D10S 185 FAM BLUE D10S 1686 VIC GREEN D10S 587 NED YELLOW D10S 537 NED YELLOW D10S 1693 VIC GREEN D10S 597 VIC GREEN D10S 1651 NED YELLOW D10S 1652 NED YELLOW
- 15. PCR reaction Components Vol per reaction (µl ) DNA (50 ng/µl) 0.6 True allel PCR mix (ABI, USA ) contains dNTPs, Polymerase, buffer, H2O 4.5 Sterile deionised water 1.9 Primer (F+R) 0.5 Total 7.5
- 16. PCR condition Cycle condition No. of cycles Initial denaturation 95ºC for 12 min 1 Denaturation 94 ºC for 15 sec 10 Anneal 55 ºC for 15 sec Extensions 72 ºC for 30 sec Melt 89 ºC for15 sec 20 Anneal 55 ºC for 15 sec Extensions 72 ºC for 30 sec Final extensions 72 ºC for 10 min 1 Hold 4 ºC hold
- 17. Capillary Electrophoresis • HiDi formamide and GS500LIZ DNA size standard (ABI, USA) • Take 1µl PCR product and 9µl Master mix in 96well sequencing plate • Then mixed spun, denatured at 95ºC for 5 minutes, snap-chilled and put in the 3130Genetic Analyzer (ABI, USA) for capillary electrophoresis. • After electrophoresis the data was analysed with Gene Mapper software (ABI, USA). HiDi + LIZ 9.0µl 0.5µlFAM : VIC : NED 1 : 1 : 2 1µl 9µl Mixed PCR product same panel Master mix
- 18. D10S1693, D10S192 , D10S1686, D10S597 RESULTS
- 19. D10S587 Heterozygous D10S537 Heterozygous
- 20. D10S185 D10S597
- 21. D10S192 250bp
- 22. Transmission of alleles of microsatellite markers 287 153 255 217 261 291 223 102 210 ---- ---- ----215 245 289 219 102 ---- D10S1652 D10S537 D10S1686 D10S185 D10S192 D10S597 D10S1693 D10S587 D10S1651 ---- 161 269 217 257 289 219 106 ---- ---- ---- ----217 259 289 221 98 ---- III-2 ----147 271 217 259 289 221 98 ---- IV-2 IV-3 273 273 291 210 258 291 219 106 216 291 147 255 206 254 291 219 98 210 IV-1 IV-4 IV-5 IV-6 IV-7 IV-8 IV-9 IV-10 291 147 259 217 259 289 221 102 216 289 161 259 217 259 291 221 106 212 291 147 273 217 259 289 221 98 212 289 147 273 217 259 289 221 98 216 289 159 255 210 259 285 223 102 210 V-1 V-2 V-3 10q21.2 10q23.2 10q24 10q24 10q24 10q25.1 10q24.2 10q25.3 10q26
- 23. • A haplotype alleles 217-259-289-221 of markers D10S185, D10S192, D10S597 and D10S1693 is co-transmitted with the disorder suggesting its linkage with the disorder. • A recombination happened between the markers D10S1693 and D10S587 in individual IV-8. • Therefore mutation/other genetic error causing split- hand/foot malformation in this family is likely to be present somewhere between D10S185 and D10S1693 markers.
- 24. • Heparinized blood was received from a patient with digital anomaly and other congenital malformations referred from the hospital for chromosome analysis . • The blood was cultured for 72 hours using RPMI1640 medium and 10% FBS. Phytohaemagglutinin (PHA-m) was added to induce cell division. • Colchicine was added 2 hour before harvesting of the culture to arrest cells in metaphase stage. • This culture was harvested and chromosome slides were prepared . • The slides were incubated for overnight followed by G-banding . • G-banded metaphase plates were observed under the microscope and karyotyping was performed using iKaryos® software (Carl Zeiss, Germany) Cytogenetic analysis of congenital limb malformations
- 25. No gross chromosomal anomaly was detected in this patient RESULT
- 26. G-banding Karyotype of the patient
- 27. ReferencesReferences • Alison M. Elliott, Martin H. Reed, and Jane A. Evans, (2007). Triphalangeal Thumb in Association with Split Hand/Foot Malformation : A Phenotypic Marker for SHFM3 Birth Defects Research (Part A) 79:58–61 • Alison M. Elliott and Jane A. Evans Genotype–Phenotype Correlations in Mapped Split Hand Foot Malformation (SHFM) Patients Department of Biochemistry and Medical Genetics, University of Manitoba, Winnipeg, Manitoba, Canada American Journal of Medical Genetics Part A 140A:1419–1427 (2006). • Christian Babbs Raoul Heller David B. Everman Mark Crocker Stephen R. F. Twigg Charles E. Schwartz Henk Giele Andrew O. M. Wilkie A new locus for split hand/foot malformation with long bone deWciency (SHFLD) at 2q14.2 Wed from a chromosome translocation Hum Genet (2007) 122:191–199.
- 28. • David B. Everman, Chad T. Morgan, Robert Lyle, Mary E. Laughridge, Michael J. Bamshad, Frequency of Genomic Rearrangements Involving the SHFM3 Locus at Chromosome 10q24 in Syndromic and Non-Syndromic Split-Hand/Foot Malformation1Center for Molecular Studies, J.C. Self Research Institute of Human Genetics, Greenwood Genetic Center, Greenwood, South Carolina American Journal of Medical Genetics Part A 140A:1375–1383 (2006). • Hans van Bokhoven ,Ben C.J.Hamel.Mike Bamshad,Eugenio Sangiorgi,fiorella Gurrieri,P63 Gene Mutation in EEC Syndrome, Department of Human Genetics, University Medical Centre, Nijmegen, The Netherlands Am. J. Hum. Genet. 69:481– 492, 2001). • Mark E.Nunes ,Gregory Schutt,Raj p.Kapur ,Frederick Luthardt ,Mary Kkolich ,Peter Byers and Jams P.Evans (1995 ) A second autosomal split hand /split foot locus maps to chromosome 10q24-q25 Human Molecular Genetics, 1995 Vol.4 , No. 11 2165-2170. • Robert Lyle, Uppala Radhakrishna, Jean-Louis Blouin, Sarantis Gaos, David B.Everman, Corinne Gehrig, Nath ,Fiorella Gurrieri ,Giovanni Neri V.Solanki, Charles E.Schwartz,and Stylianos E. Antonarakis (2005 ) Split-Hand /Split Foot Malformation 3 (SHFM3) at 10q24,Development of Rapid Diagnostic Methods and Gene Expression From the Region American General of Medical Genetics part A 140A:1384-1395.
- 29. • Rydvan S. Ozen, Bora E. Baysal, Bernie Devlin , Joan E. Farr ,Michael D.Ehrlich, and Charles W. Richard, III (1999) Gorry, and Garth Fine Mapping of the Split- hand/Split-Foot locus (SHFM3) at 10q 24: Evidence for Anticipation and Segregation Distortion Am.j. Human. Genetic.64:1646-1654 • Mohammed Naveed, Swapan K. Nath, Mathew Gaines, Mahmoud T. Al-Ali, Genomewide Linkage Scan for Split–Hand/Foot Malformation with Long-Bone Deficiency in a Large Arab Family Identifies Two Novel Susceptibility Loci on Chromosomes 1q42.2-q43 and 6q14.1 American Journal of Medical Genetics Part A 140A:1375–1383 (2006). • Tony Roscioli, Peter J. Taylor, Andrew Bohlken, Jennifer A. Donald, John Masel, Ian A. Glass, and Michael F. Buckley, The 10q24-linked Split Hand/Split Foot Syndrome (SHFM3): Narrowing of the Critical Region and Confirmation of the Clinical Phenotype American Journal of Medical Genetics 124A:136–141 (2004). • Raas-Rothschild, S Manouvrier, M Gonzales, J P Farriaux, S Lyonner, A Munich Refined mapping of a gene for split hand-split foot malformation on chromosome 10q25 France medical genetic 1996;33;996-1001
