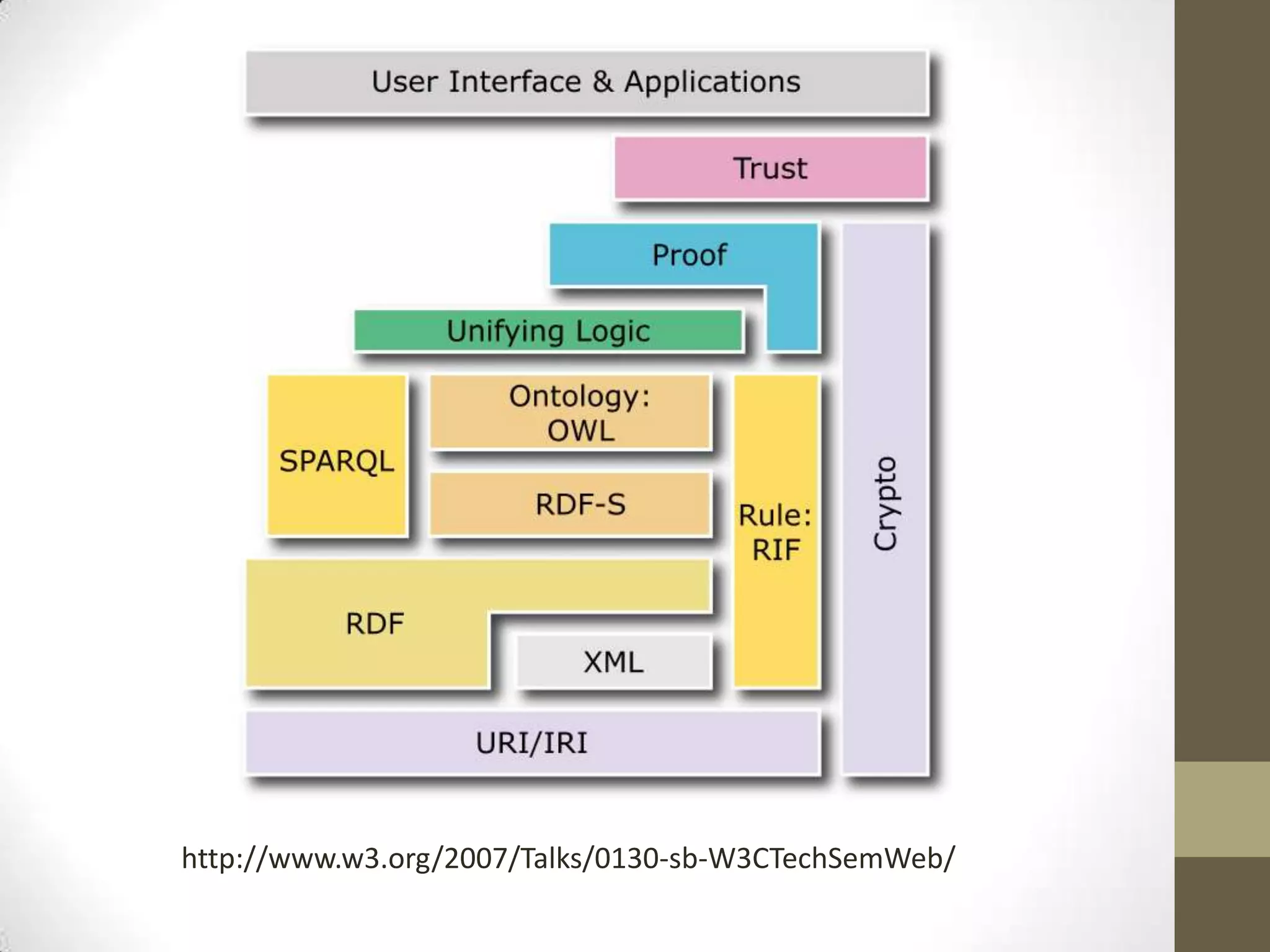
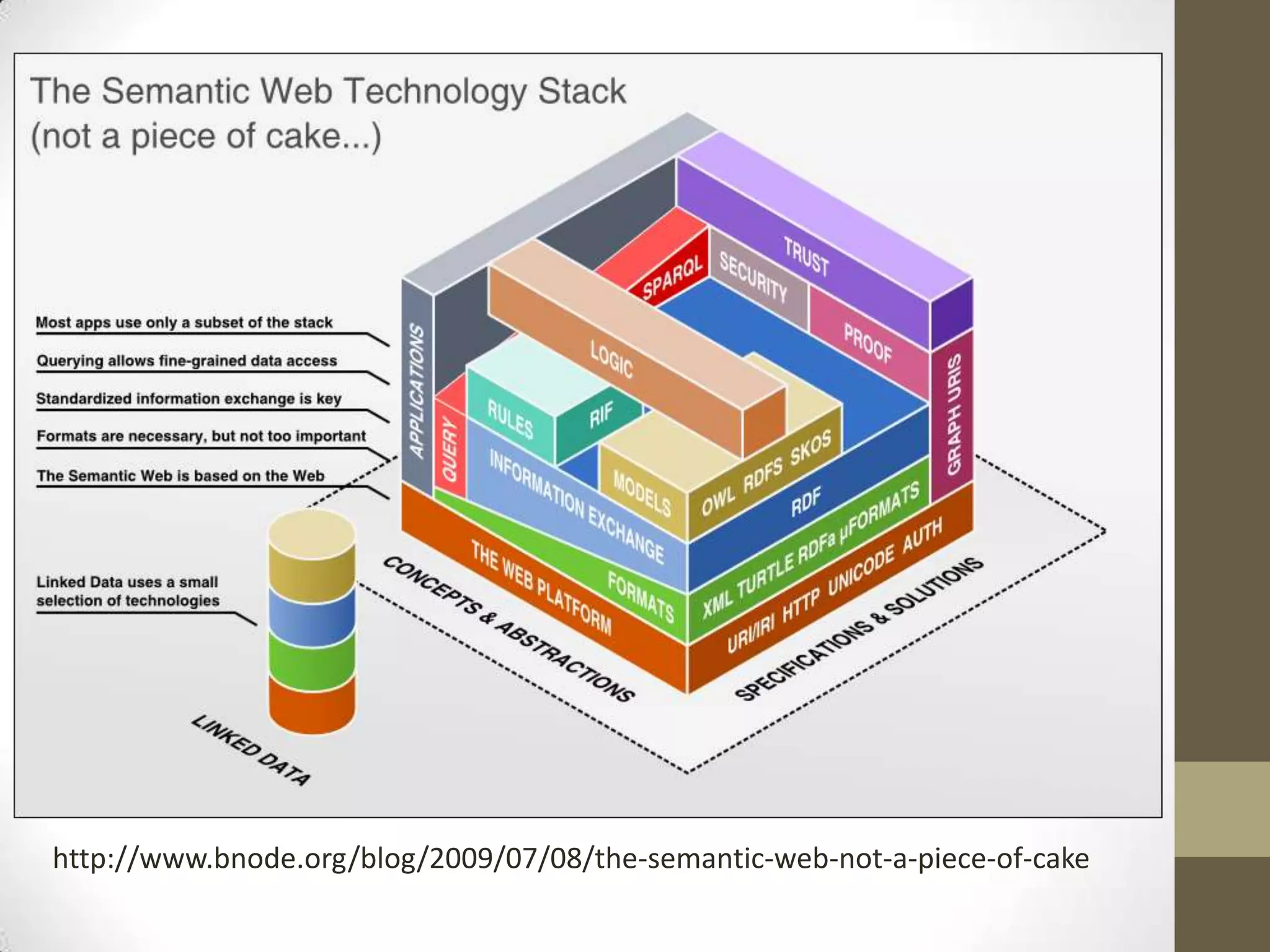
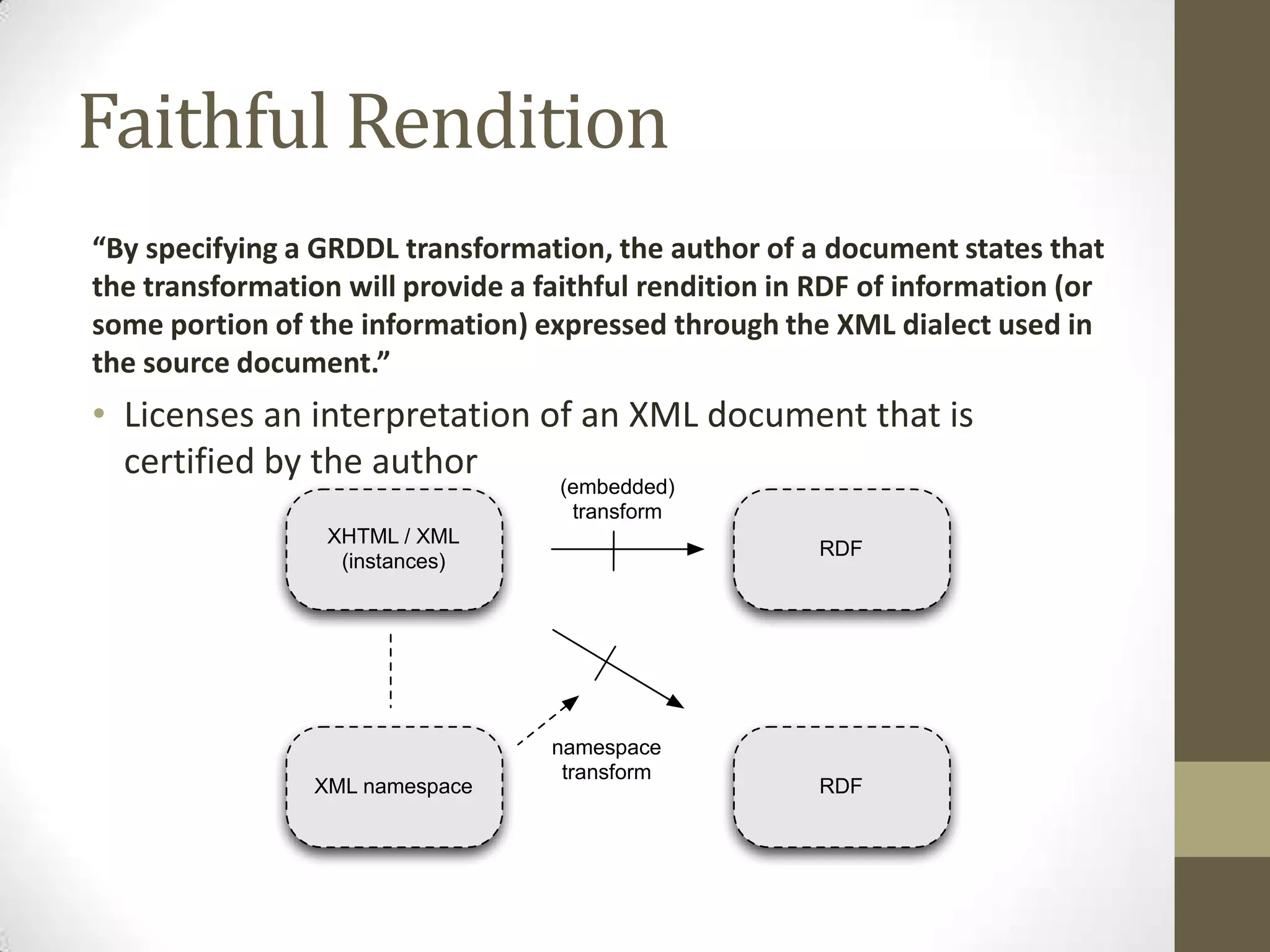
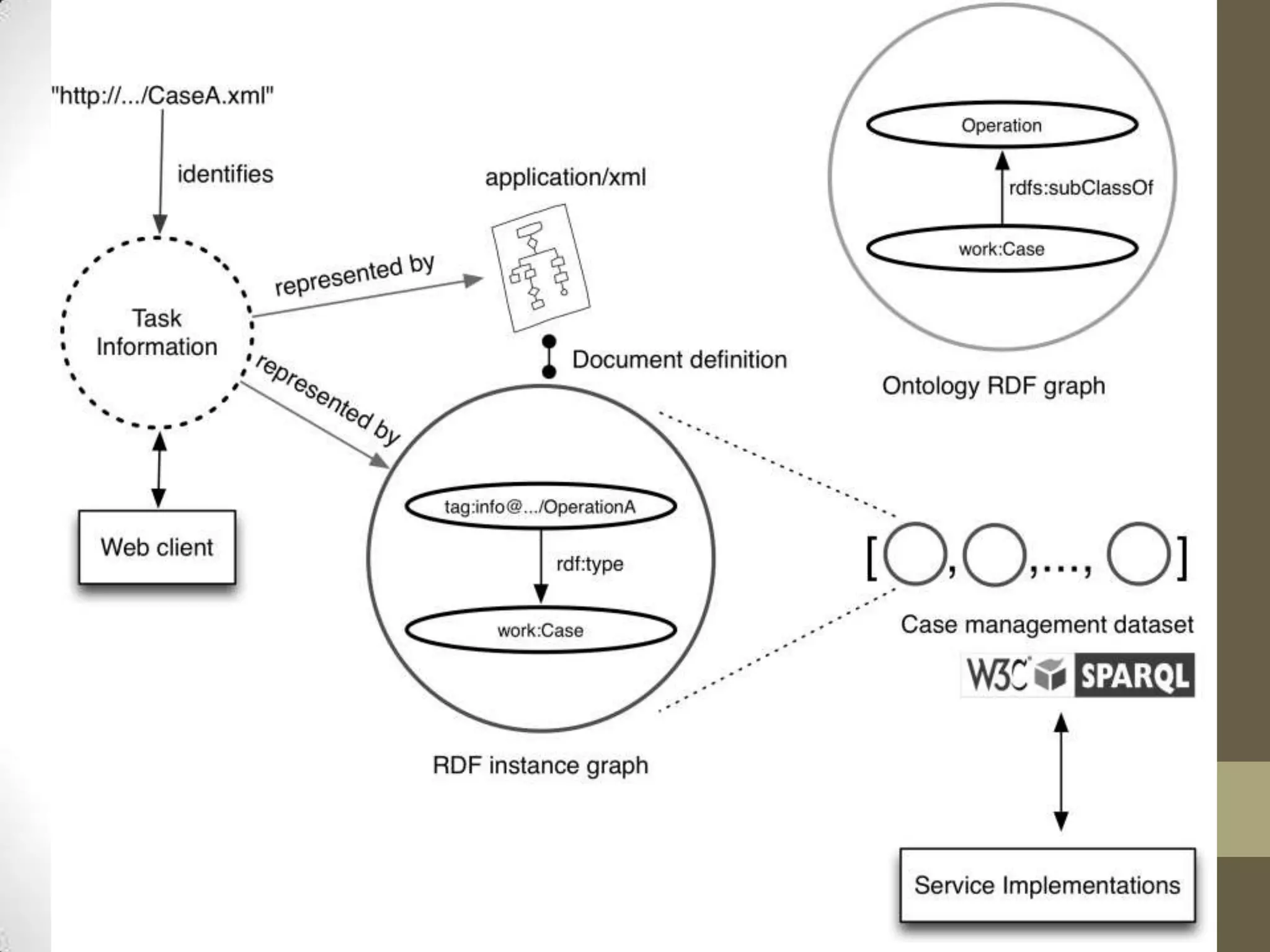
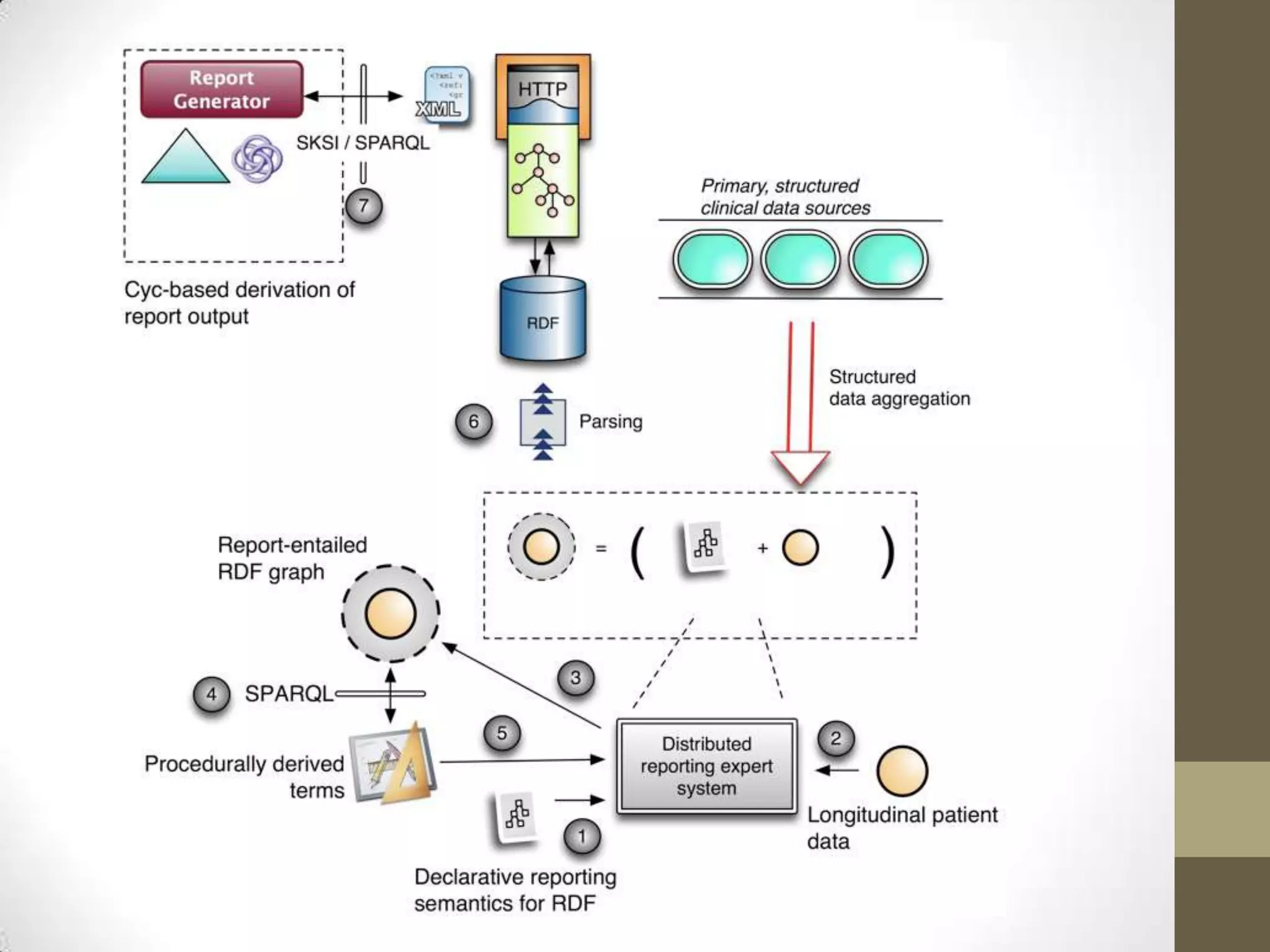
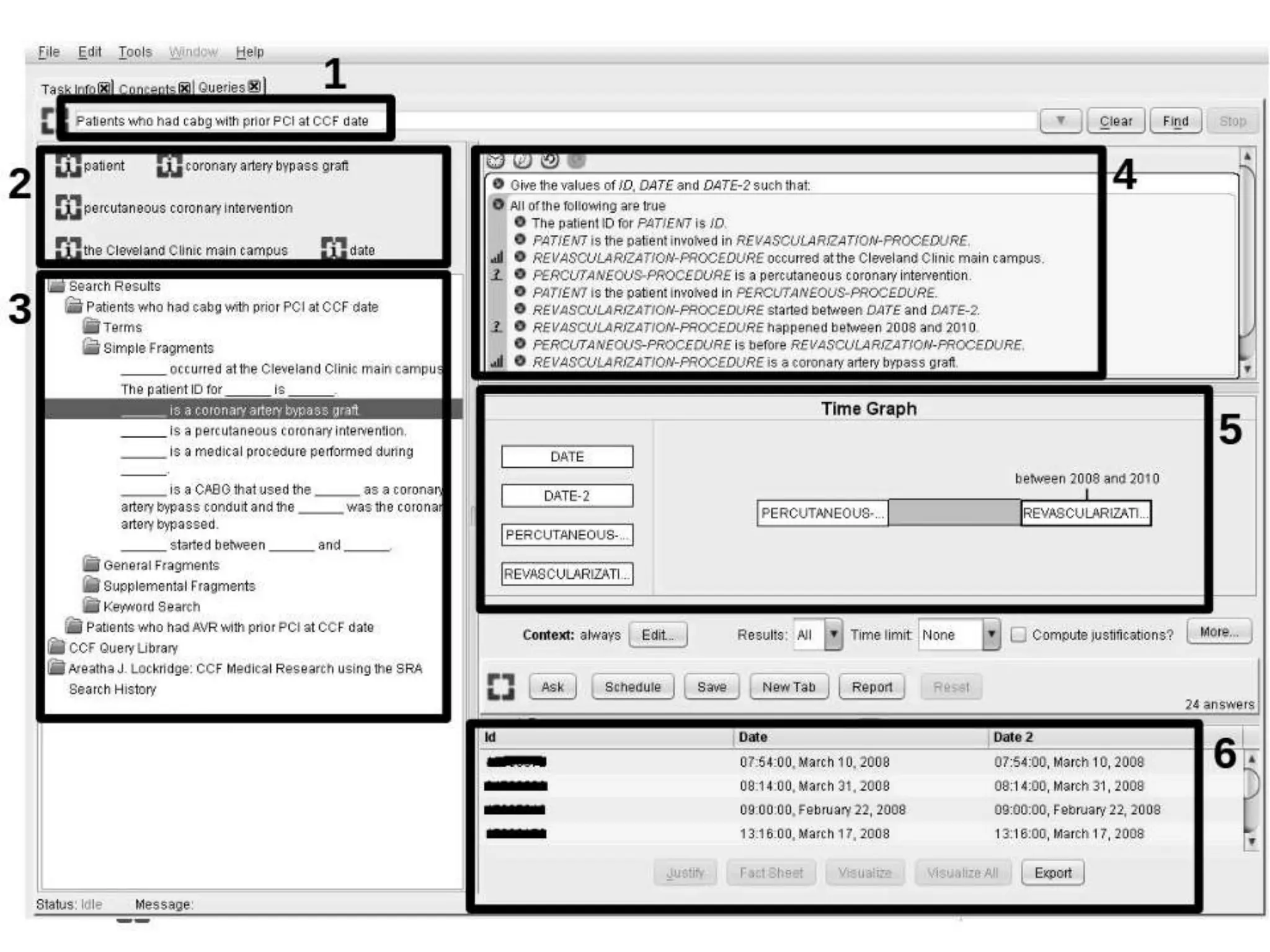
The presentation discusses the implementation of semantic web technologies in outcomes research, highlighting experiences from the Cleveland Clinic's semantic database project. Key components include RDF, SPARQL, and OWL to facilitate data management, reporting, and cohort identification for patient records. The system has enabled over 200 clinical investigations to utilize semanticdb for efficient cohort selection and data analysis.