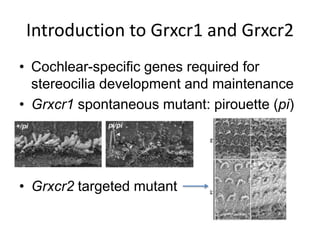
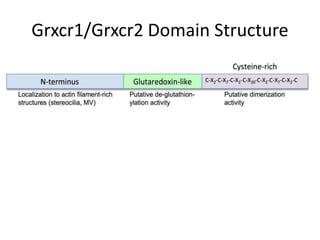
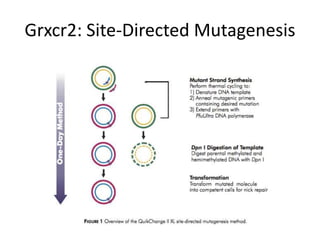
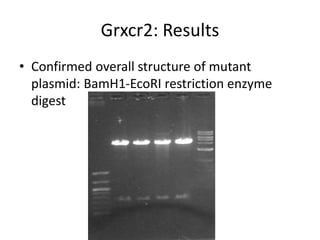
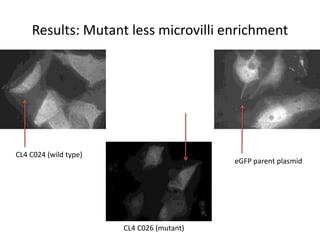
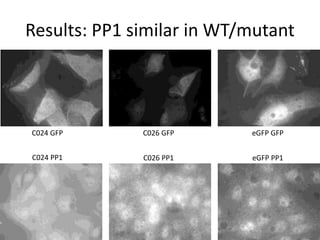
This presentation summarizes Keerthana Velappan's research on the cochlear-specific genes Grxcr1 and Grxcr2 during her Kresge Summer Research Internship. The genes are required for stereocilia development and maintenance in the inner ear. The intern generated mutant versions of the genes lacking a putative PP1 binding site to test the hypothesis that Grxcr1 and Grxcr2 control phosphorylation via PP1 inhibition. Preliminary results show the Grxcr2 mutant localizes differently than the wild type when expressed in epithelial cells. Future work will generate a Grxcr1 mutant and further examine mutant localization and effects on PP1.