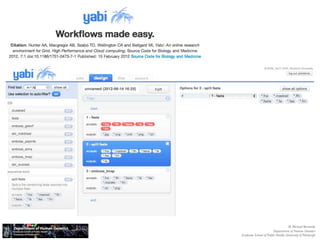
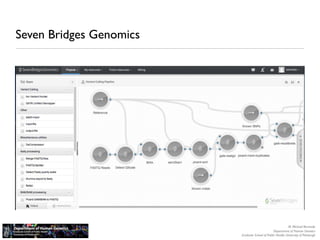
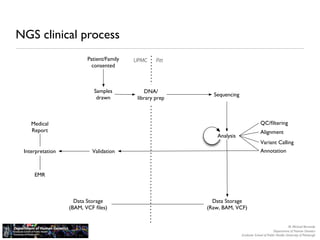
The document discusses barriers to centralizing clinical next-generation sequencing (NGS) data analysis at the University of Pittsburgh. It describes how the university's geographically dispersed campus and hospital system initially led to many small, localized computing clusters lacking sufficient resources for NGS work. It then outlines how the university established a larger central computing cluster and upgraded storage and networking capabilities to help address these issues. However, software interfaces and data sharing across resources remained challenges. The document proposes several solutions being implemented, including research gateways and cloud computing, to further facilitate centralized analysis while improving access and data management.