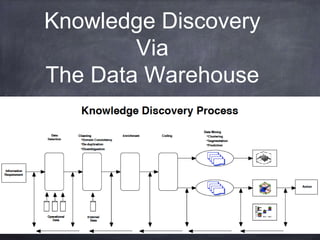
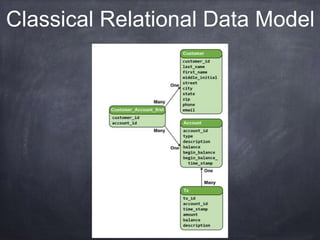
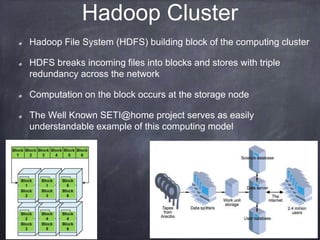
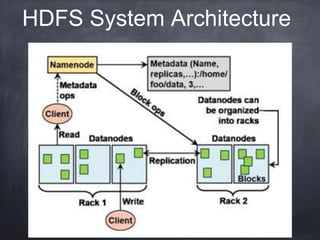
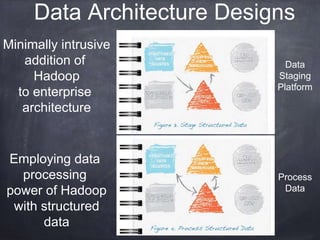
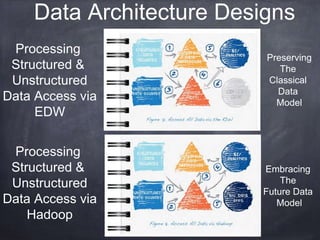
This document provides an introductory tutorial on big data in medicine and healthcare. It defines big data as large volumes of structured, semi-structured, and unstructured data that can be mined for information, often referring to sizes in petabytes and exabytes. The key dimensions of big data are described as volume, velocity, variety, and veracity. Hadoop is presented as an open-source framework for distributed storage and processing of large datasets across clusters of commodity servers. Examples of using Hadoop and MapReduce for medical applications like predictive modeling, genomic research, and data integration are also provided.