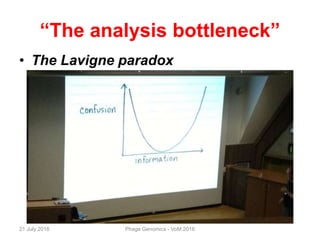
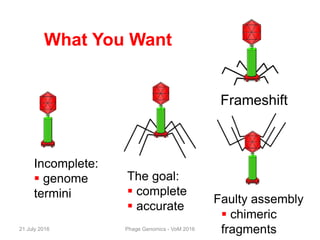
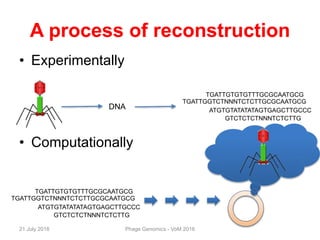
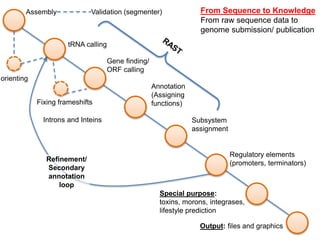
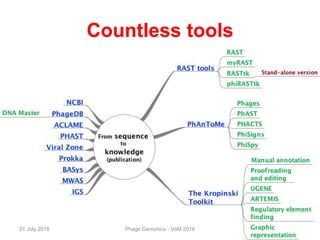
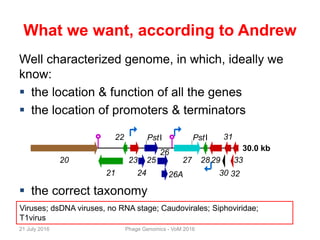
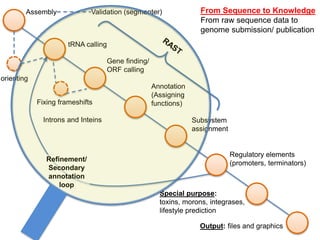
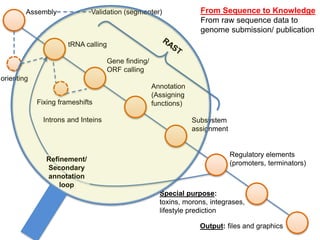
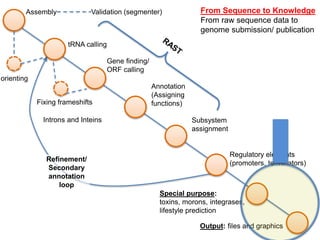
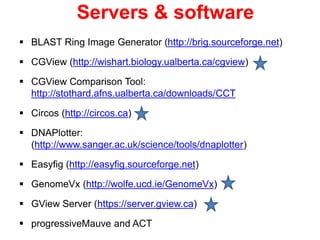
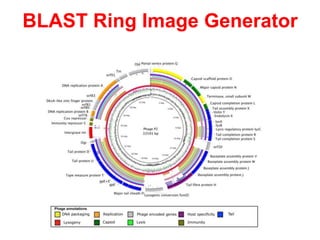
The document discusses the challenges of phage genome annotation, highlighting the analysis bottleneck due to rapid data generation. It introduces various tools and frameworks, including the Kropinski Toolkit and RAST, to enhance genome reconstruction and annotation accuracy. It emphasizes the importance of defining specific goals and employing both automated and manual processes for reliable genome analysis.