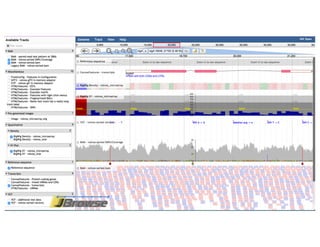
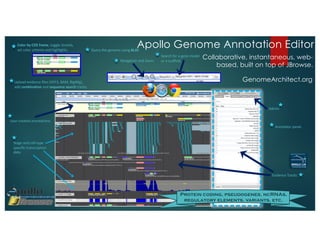
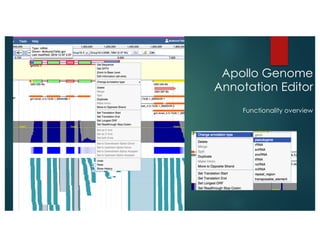
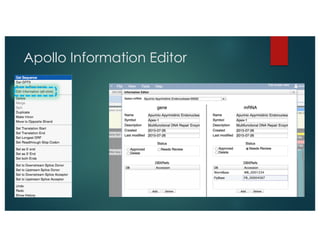
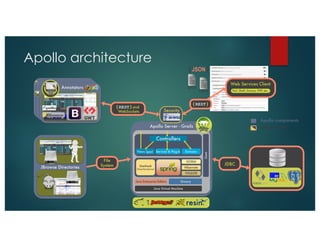
The document outlines the features and future developments of the JBrowse and Apollo genome annotation tools, detailing their capabilities in handling large genomes and supporting various data formats. It emphasizes JBrowse's transition to a more versatile JavaScript client and the enhancements planned for Apollo, including better coordination transformations and generalized annotation features. Both tools are supported by several NIH grants and are utilized by numerous institutions globally.