Genetic Factors in Tracheal Development and Malformation
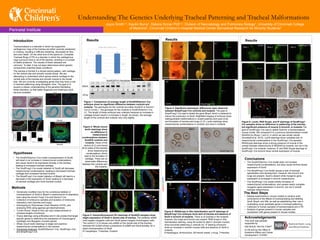
- 1. Results Figure 1: Comparison of average length of Sox9OXDermo1-Cre embryos show no significant difference between controls and mutants. The genotypes for the controls are either Sox9OX or Dermo1- Cre (2,3,4,etc.). The genotype for the mutants is Sox9OXDermo1-Cre (1). The length of these embryos was collected because an increase in cartilage should result in a increase in length. As shown, the average length of the controls and mutants vary only slightly. Figure 2: Whole mount lectin stainings show no difference in mesenchymal condensations between Sox9OX controls and mutants. Views of the ventral (A,C) and dorsal (B,D) sides of the trachea show mesenchymal condensations, which lead to the formation of cartilage. There are no observable differences between the controls and the mutants. Figure 3: Immunofluorescent (IF) stainings of Sox9OX samples show slight expression of Sox9 in dorsal side of trachea. The antibody aSMA (red) targets myogenic cells and Sox9 (green) targets chondrogenic cells. The control (1,2,3) shows only aSMA on the dorsal side of the trachea, whereas the mutant shows a prescence of aSMA and Sox9 dorsally, for a slight overexpression of Sox9. E= esopahgus, T=trachea, S=spine Understanding The Genetics Underlying Tracheal Patterning and Tracheal Malformations Introduction Tracheomalacia is a disorder in which the supportive cartilaginous rings of the trachea are either severely weakened or missing, resulting in difficulty breathing, decreased air flow, and even death. On the other end of the spectrum, Complete Tracheal Rings (CTR) is a disorder in which the cartilaginous rings surround most or all of the trachea, resulting in a number of health problems. The causes of these diseases are unknown. To date, it has not been determined which genetic components underline these conditions. The trachea is formed in a dorsal-ventral pattern, with cartilage on the ventral side and smooth muscle dorsal. We are attempting to understand which genes restrict cartilage to the ventral side of the trachea and smooth muscle to the dorsal side. We are currently investigating genes that may have a role in tracheal patterning using transgenic mice. The goal is to acquire a clearer understanding of the genetics facilitating these disorders, so that better diagnosis and treatment could become available. Results Results • The Sox9OXDermo1-Cre model does not increase mesenchymal condensations, but does cause minimal dorsal expression of Sox9. • The Bmp4Foxg1-Cre model causes a lack of tracheal specification and development. However, the bronchi and lungs are present. Sox9 is absent while myogenic gene expression is increased in bronchi mesenchyme . • The Sox9Foxg1-Cre model inhibits formation of mesenchymal condensations, and causes nearly complete myogenic gene expression in bronchi, but not in ventral tracheal mesenchyme. • The Sox9OXDermo1-Cre model (overexpression of Sox9) will result in an increase in mesenchymal condensations, and cause Sox9 to be expressed dorsally in the trachea, leading to increased tracheal cartilage. • The Sox9Foxg1-Cre model (deletion of Sox9) will decrease mesenchymal condensations, leading to decreased tracheal cartilage and increased tracheal muscle . • The Bmp4Foxg1-Cre model (deletion of Bmp4) will lead to a decrease in the expression of Sox9, leading to a decrease of tracheal cartilage and more tracheal muscle. Hypotheses . Joyce Smith1,3, Kaulini Burra1, Debora Sinner PhD1,2, Division of Neonatology and Pulmonary Biology1, University of Cincinnati College of Medicine2, Cincinnati Children’s Hospital Medical Center Biomedical Research for Minority Students3 Perinatal Institute Methods • Genetically modified mice for the conditional deletion or overexpression of Sox9 or Bmp4 in mesenchyme of respiratory tract using the drivers Foxg1-Cre and Dermo1-Cre. • Collection of embryonic samples and isolation of embryonic respiratory tract (trachea and lungs). • Genotyping by Polymerase Chain Reaction (PCR), and analyzing DNA using agarose gel electrophoresis • “Crown-Rump” (head to foot, per say) measurements to compare lengths of embryos. • Tissue stainings using antibodies and In situ probes that target specific genes to determine the expression of Chondrogenic (cartilage) and Myogenic (muscle) genes. • Whole mount lectin stainings to target and examine mesenchymal condensations in the trachea. • Genotypes Analyzed: Sox9OXDermo1-Cre, Sox9Foxg1-Cre, and Bmp4Foxg1-Cre. A B C D control Sox9OXDermo1-Cre CTR mutant control Figure 4: Significant phenotypic differences were observed between Bmp4Foxg1-Cre controls and mutants. The goal of Bmp4Foxg1-Cre was to delete the gene Bmp4, which is believed to induce the production of Sox9. Brightfield imaging of embryos show distinguishable malformations in mutant jawlines and eyes (A,B), and formation of trachea and lungs (C,D). Lectin stainings show mesenchymal condensations in controls, but none in mutants. Figure 5: RNAScope In Situ Hybridization and IF stainings of Bmp4Foxg1-Cre embryos show lack of trachea and absence of Sox9 in bronchi of mutants. There is no trachea in the mutants; however, the lungs and bronchi are present. RNA Scope In Situ Hybridization staining, using the probe for Myh11 (smooth muscle) show bronchial patterning in controls (A). IF staining of bronchi show an increase in smooth muscle cells and absence of Sox9 in mutants. E=esophagus, Br=bronchus, BV=blood vessel, L=lung, T=trachea The Next Steps • Repeat experiments in mouse models to achieve solid conclusions of the effects of overexpressing and deleting Sox9, Bmp4, and Wls, as well as establishing their role in modulating novel candidate genes in tracheal patterning. • Collection of human samples of tracheomalacia and CTR. • Analysis of genes present in human samples and comparison with genes present in mouse models. This work was partially supported by NIH-NHLBI 1K01HL115447 to DS and by the Office of Academic Affairs and Career Development, CCHMC. Acknowledgements control Sox9OXDermo1-Cre Sox9OXD10 Genotypes 1 S9OX 2 3 4 5 6 7 8 9 10 C1 C2 DCre Figure 6: Lectin, RNA Scope, and IF stainings of Sox9Foxg1- Cre samples show no difference in patterning of the trachea, but significant presence of muscle in bronchi, in mutants.The goal of Sox9Foxg1-Cre was to delete Sox9 for a tracheomalacia mouse model. We compared it to a previous tracheomalacia model, WlsShhCre (Rows 1 and 2), in which we see similar results (Snowball et.al. 2015). Lectin stainings show complete lack of mesenchymal condensations in the Sox9Foxg1-Cre samples. RNAScope stainings show a strong presence of muscle in the ventral tracheal mesenchyme of WlsShhCre mutants, but not in the Sox9Foxg1-Cre mutants; however, IF and RNA Scope stainings of Sox9Foxg1-Cre bronchi show ventral expression of muscle. Br Br BV BV E L control Bmp4Foxg1-Cre E Row 1 Row 2 Row 3 control WlsShhCre Sox9Foxg1-Cre A B E Sox9/aSMA T E T E T BV BV BV Br Br Br Br E BV aSMA/Sox9 A B Myh11 aSMA/Sox9/TTF1 E Br Br E Br Br Br control Bmp4Foxg1-Cre 1 2 3 4 5 6 E T E T S control Sox9OXDermo1-Cre Conclusions Bmp4Foxg1-Cre control control Bmp4Foxg1-Cre A B C D E1 F1 E2 F2 Sox9Foxg1-Cre Sox9Foxg1-Cre