TaguibaoCandice_Binucleation in Preimplantation Embryos Poster
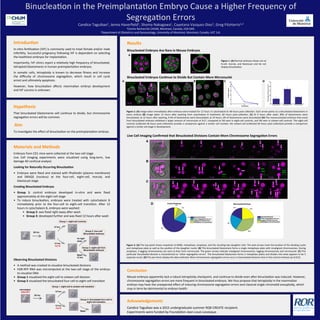
- 1. Introduc)on In-‐vitro fer,liza,on (IVF) is commonly used to treat female and/or male infer,lity. Successful pregnancy following IVF is dependent on selec,ng the healthiest embryos for implanta,on. Importantly, IVF clinics report a rela,vely high frequency of binucleated, tetraploid blastomeres in human preimplanta,on embryos. In soma,c cells, tetraploidy is known to decrease fitness and increase the difficulty of chromosome segrega,on, which result in cell cycle arrest and ul,mately apoptosis. However, how binuclea,on affects mammalian embryo development and IVF success is unknown. Materials and Methods Embryos from CD1 mice were collected at the two-‐cell stage. Live Cell Imaging experiments were visualized using long-‐term, low damage 4D confocal analysis Looking for Naturally Occurring Binuclea)on • Embryos were fixed and stained with Phalloidin (plasma membrane) and DRAQ5 (nucleus) at the four-‐cell, eight-‐cell, morula, and blastocyst stage Crea)ng Binucleated Embryos • Group 1: control embryos developed in-‐vitro and were fixed approximately at the eight-‐cell stage • To induce binuclea,on, embryos were treated with cytochalasin B immediately prior to the four-‐cell to eight-‐cell transi,on. AYer 12 hours in cytochalasin B, embryos were washed: • Group 2: was fixed right away aYer wash • Group 3: developed further and was fixed 12 hours aYer wash Observing Binucleated Divisions • A method was created to visualize binucleated divisions • H2B-‐RFP RNA was microinjected at the two-‐cell stage of the embryo to visualize DNA • Group 1 visualized the eight-‐cell to sixteen-‐cell division • Group 2 visualized the binucleated four-‐cell to eight-‐cell transi,on Results Binucleated Embryos Are Rare in Mouse Embryos Binucleated Embryos Con)nue to Divide But Contain More Micronuclei Live Cell Imaging Confirmed that Binucleated Divisions Contain More Chromosome Segrega)on Errors Binuclea,on in the Preimplanta,on Embryo Cause a Higher Frequency of Segrega,on Errors Candice Taguibao1, Jenna Haverfield1, Shoma Nakagawa1, Cayetana Vazquez-‐Diez1, Greg FitzHarris1,2 1Centre Recherche CHUM, Montreal, Canada, H2X 0A9. 2Department of Obstetrics and Gynaecology, University of Montreal, Montreal, Canada, H3T 1J4. Hypothesis That binucleated blastomeres will con,nue to divide, but chromosome segrega,on errors will be common. Aim To inves,gate the effect of binuclea,on on the preimplanta,on embryo. Conclusion Mouse embryos apparently lack a robust tetraploidy checkpoint, and con,nue to divide even aYer binuclea,on was induced. However, chromosome segrega,on errors are more frequent in binucleated embryos. We thus propose that tetraploidy in the mammalian embryo may have the unexpected effect of inducing chromosome segrega,on errors and classical single-‐chroma,d aneuploidy, which may in term be detrimental to embryo health. Figure 2. (A) Image taken immediately aYer embryos were treated for 12 hours in cytochalasin B; 48 hours post-‐collec,on. Each arrow points to a binucleated blastomere in every embryo (B) Image taken 12 hours aYer washing from cytochalasin B treatment; 60 hours post-‐collec,on. (C) At 0 hours aYer wash, 90% of blastomeres were binucleated; at 12 hours aYer washing, 9.4% of blastomeres were binucleated; at 24 hours, 0% of blastomeres were binucleated (D) The mononucleated embryos that arose from binucleated embryos exhibited a larger amount of micronuclei at %17, compared to %6 seen in eight-‐cell controls, and %8 seen in sixteen-‐cell controls. The eight-‐cell controls (collected 48 hours post-‐collec,on) provide a comparison against a similar cell number; the sixteen-‐cell (collected 60 hours post-‐collec,on) provide a comparison against a similar cell-‐stage in development. Old Acknowledgements Candice Taguibao was a 2015 undergraduate summer RQR CREATE recipient. Experiments were funded by Founda,on Jean-‐Louis Levesque. A B D C A B Figure 3. (A) The top panel shows snapshots at NEBD, metaphase, anaphase, and the resul,ng two daughter cells. The pink arrows mark the loca,on of the dividing nuclei and metaphase plate as well as the posi,on of the daughter nuclei. (B) This binucleated blastomere forms a single metaphase plate with misaligned chromosomes. During anaphase, 2 lagging chromosomes are seen to then form micronuclei. The green arrows indicate misaligned chromosomes, lagging chromosomes and micronuclei. (C) This par,cular binucleated division is characterized as “other segrega,on errors’. The binucleated blastomere forms 2 metaphase plates and divides into what appears to be 5 separate nuclei. (D) The pie charts display the data collected. More chromosome segrega,on errors occur in binucleated divisions than in the control embryos (p=0.012) C D Figure 1. (A) Normal embryos shown are at 8-‐cell, morula, and blastocyst and do not display binuclea,on A 36 hrs 12 hrs Cyt B 12 hrs Wash: 12 hrs fix right away Group 1: eight-cell controls Group 3: eight-cell from Binucleated embryos Group 2: four-cell Binucleated embryos microinject H2B-RFP 36 hrs 12 hrs 12 hrs Cyt B Wash Group 1: eight-cell to sixteen-cell transition Group 2: binucleated four-cell to eight-cell transition 48:00 72:00 96:00 48:00 48:00 60:00 Misaligned Chromosomes Lagging Chromosomes Micronuclei Series 001 z=13/27 Series 001 z=9/27 Series 002 z=14/28 48:00 49:83 50:17 50:50 51:08 51:17 51:83 49:83 50:08 51:8351:2551:1751:0849:83 49:00 51:9151:7551:5051:3349:33 52:08 CONTROL BINUCLEATED BINUCLEATED