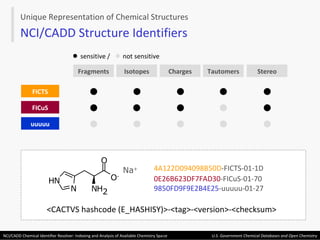
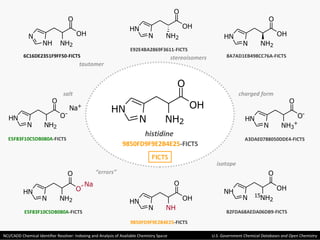
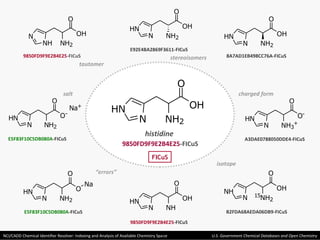
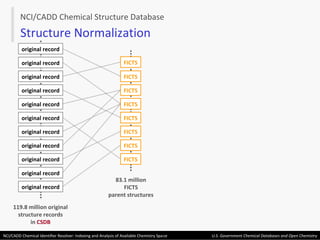
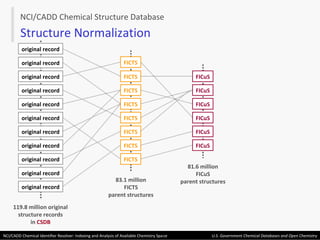
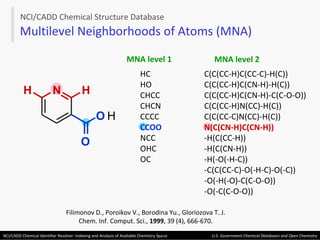
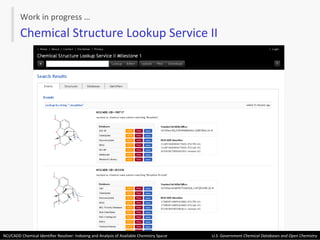
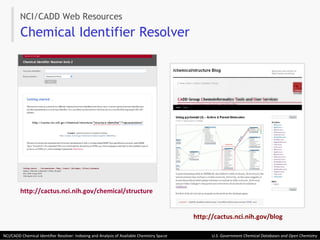
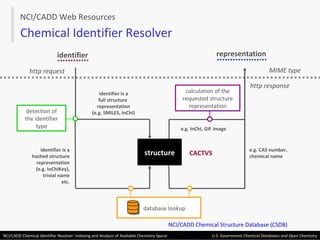
The document describes the NCI/CADD Chemical Identifier Resolver, which works as a resolver for different chemical structure identifiers. It allows the conversion of a given structure identifier into another representation or identifier. The resolver indexes over 150 chemical structure databases containing over 120 million structures. It is accessible via a web API that detects the identifier type and performs the requested conversion via calculation or database lookup.
![Markus Sitzmann 1 , Wolf-Dietrich Ihlenfeldt 2 , and Marc C. Nicklaus 1 [1] Computer-Aided Drug Design Group, Chemical Biology Laboratory, NCI-Frederick, NIH, DHHS [2] Xemistry GmbH, Auf den Stieden 8, D-35094 Lahntal, Germany NCI/CADD Chemical Identifier Resolver: Indexing and Analysis of Available Chemistry Space](https://image.slidesharecdn.com/markussitzmann-usdatabasemeeting-public-110831145516-phpapp02/85/5th-Meeting-on-U-S-Government-Chemical-Databases-and-Open-Chemistry-Talk-1-320.jpg)






C(O)=O </item> </data> <data id=" 2 " resolver=" name_by_opsin " string_class=" IUPAC Name (OPSIN) "> <item id=" 1 "> C[C@H](N)C(O)=O </item> </data> <data id=" 3 " resolver=" name_by_cir " string_class=" Chemical Name (CIR) "> <item id=" 1 “> C[C@H](N)C(O)=O </item> </data> </request> http://cactus.nci.nih.gov/chemical/structure/ L-alanin /smiles/xmls ?resolver= name_by_chemspider , name_by_opsin , name_by_cir Chemical Identifier Resolver NCI/CADD Web Resources](https://image.slidesharecdn.com/markussitzmann-usdatabasemeeting-public-110831145516-phpapp02/85/5th-Meeting-on-U-S-Government-Chemical-Databases-and-Open-Chemistry-Talk-9-320.jpg)