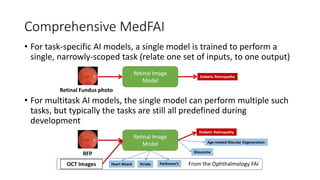
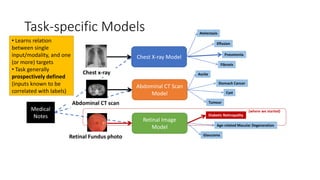
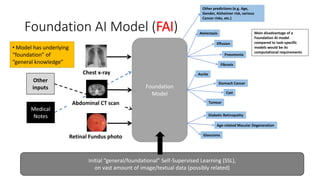
The document discusses foundation models versus task-specific models. It notes that foundation models are trained on vast amounts of diverse, unlabeled data using self-supervised learning, often based on large language models. They can then be fine-tuned for specific tasks. In contrast, task-specific models are intended for a single task and are fragile, not easily repurposed. It provides examples of task-specific medical models and discusses how a foundation AI model could integrate different medical inputs and outputs instead of being limited to a single task. Developing a comprehensive medical foundation model could allow for more holistic, predictive, and sparse capabilities beyond what individual physicians can achieve.






![Masked Autoencoder (Generative SSL)
(trained) ViT
Encoder
FAI Task Adaptation
Test CFP
Encoder
high-level
features
Multilayer
Perceptron
Prediction
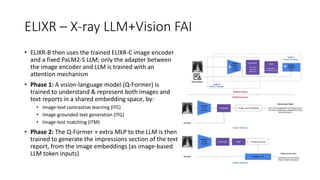
• RetFound uses a masked
autoencoder, with the training
objective being to reconstruct
input images from a randomly-
masked version of the image
• Once the autoencoder is trained,
only the (ViT) encoder is used to
generate the high-level features
for task-specific classification
• Comparison against established
SL DCNN models (trained directly
on the [augmented] images,
instead of high-level features)
would have been interesting
Main value of FAI here
lies in producing good
high-level features?
Self-supervised since no
labels are required in
training the autoencoder,
only the (randomly
masked) image itself
Note that actually still
need the task labels, to
train the task-specific
classifiers!
Standard Classification](https://image.slidesharecdn.com/foundationmultimodels-231127075449-a76cebb2/85/Foundation-Multimodels-pptx-9-320.jpg)


![General FAI for Medicine (MedFAI)
• Obvious extension: from “ophthalmology FAI” to “medical FAI”
• Several desirable attributes going “beyond human physicians”:
• Holistic
Modern medicine is necessarily fragmented into specialties
(too much for any single physician to know/learn)
Cross-specialty boundaries difficult to cross
(e.g. eye [images] as window to heart/brain/cardiovascular health etc.)
• Comprehensive
Can in theory query implications of Variable(s) A → Condition/Outcome B, for
any A and B, with evidence-based justification
• Predictive
Physicians generally can only diagnose current conditions, not future](https://image.slidesharecdn.com/foundationmultimodels-231127075449-a76cebb2/85/Foundation-Multimodels-pptx-13-320.jpg)
![Holistic MedFAI
• Previously, AI models in medicine are generally designed to replicate
existing capabilities, or at least prospectively
• For example, it is known that retinal fundus photographs can be used
to diagnose diabetic retinopathy (DR)
• So we plan to train an AI model to classify DR from retinal photos
• Then just a matter of collecting sufficient labelled data
(both for model development and [external] validation)
• Often encounter delays with data acquisition, model robustness
(if insufficient data)](https://image.slidesharecdn.com/foundationmultimodels-231127075449-a76cebb2/85/Foundation-Multimodels-pptx-14-320.jpg)