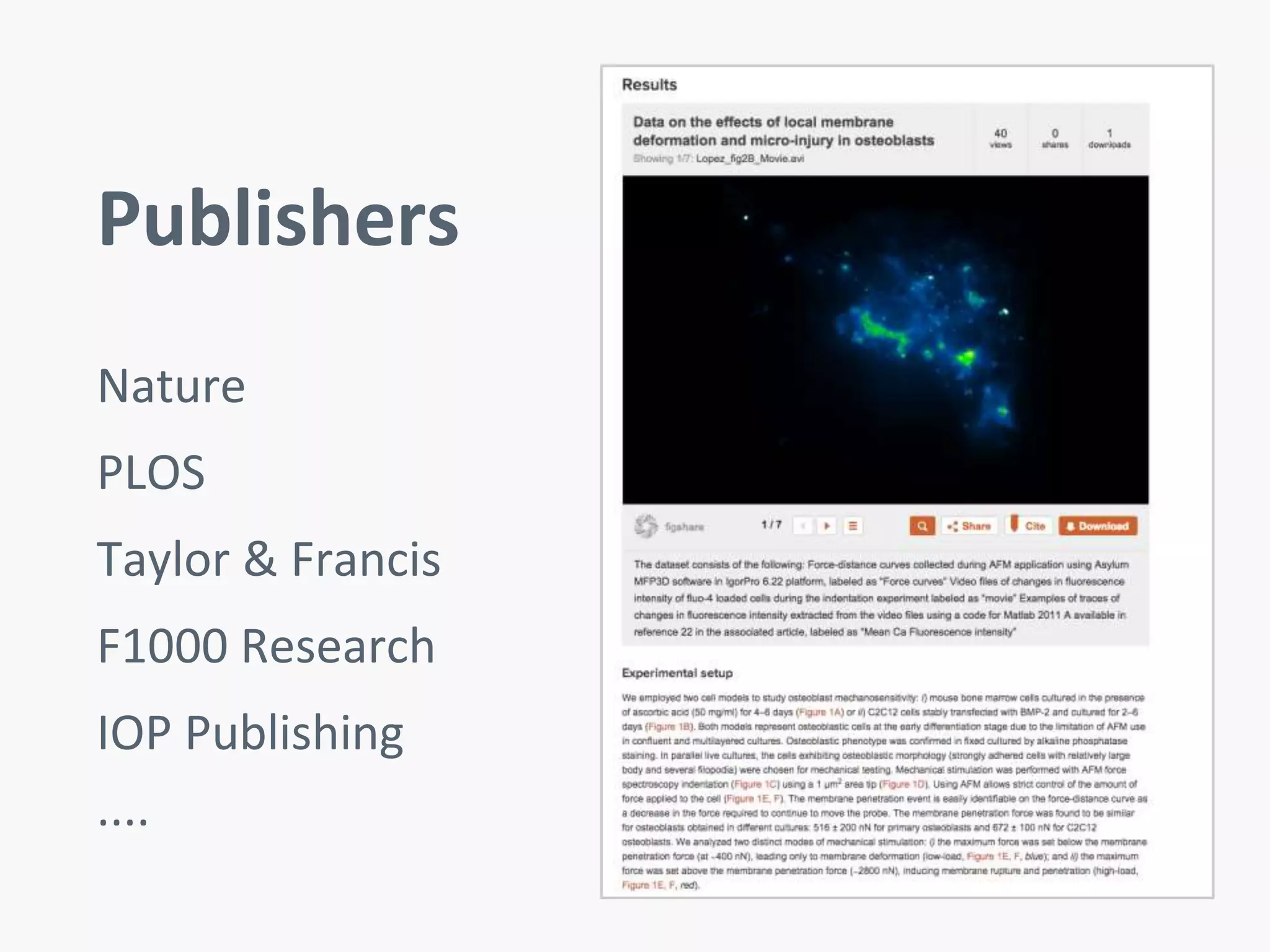
Figshare is an open data repository that allows researchers to publish all of their research data online for public access within minutes. This speeds up the publishing process compared to traditional journals, which can take years and only share a small portion of the data. Figshare aims to improve collaboration and reproducibility of research by making all data citable and discoverable. It has over 1.5 million public research articles and is used by many universities worldwide to help students and researchers more easily access and share data.