Embed presentation
Download as PDF, PPTX






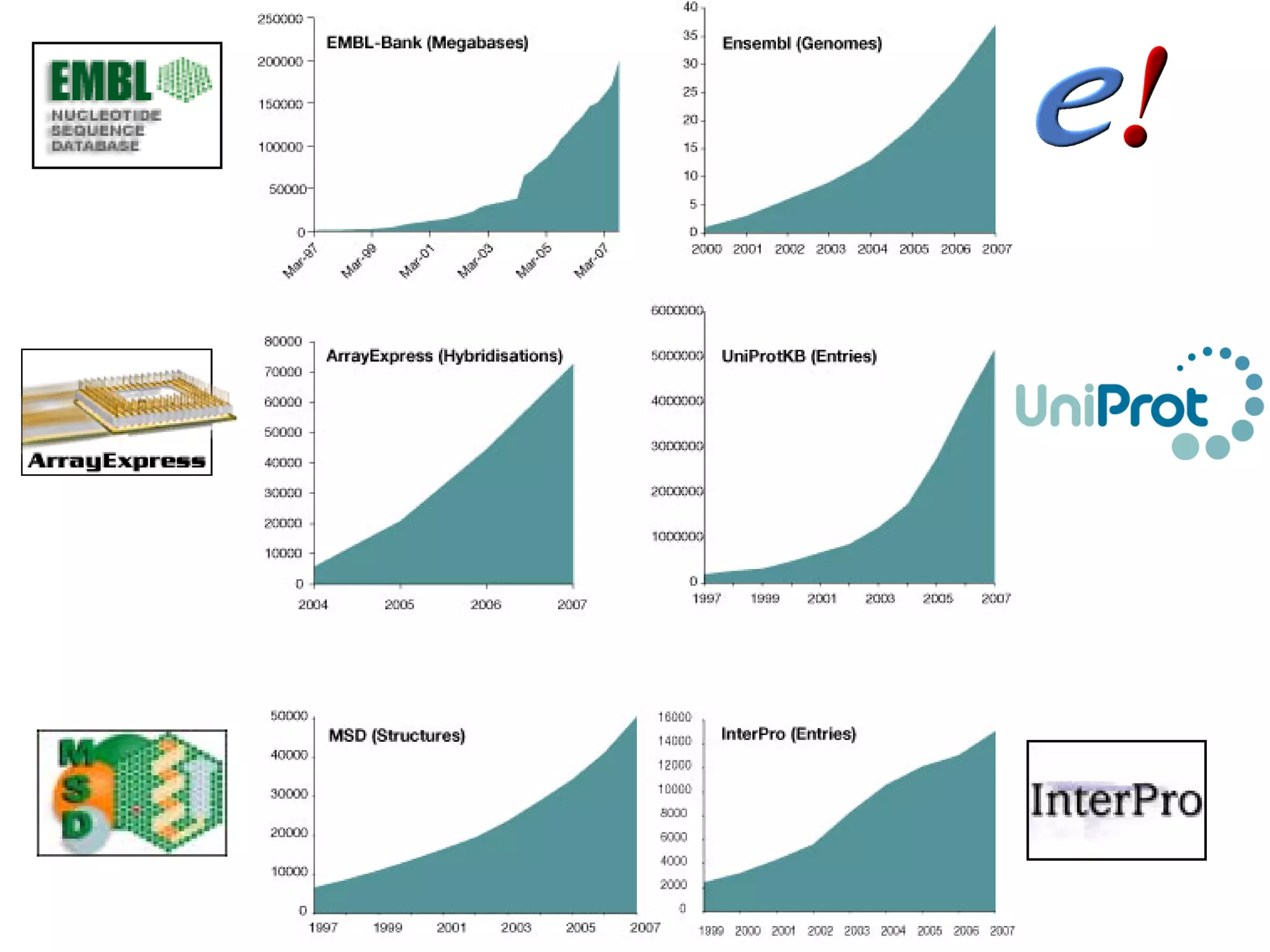

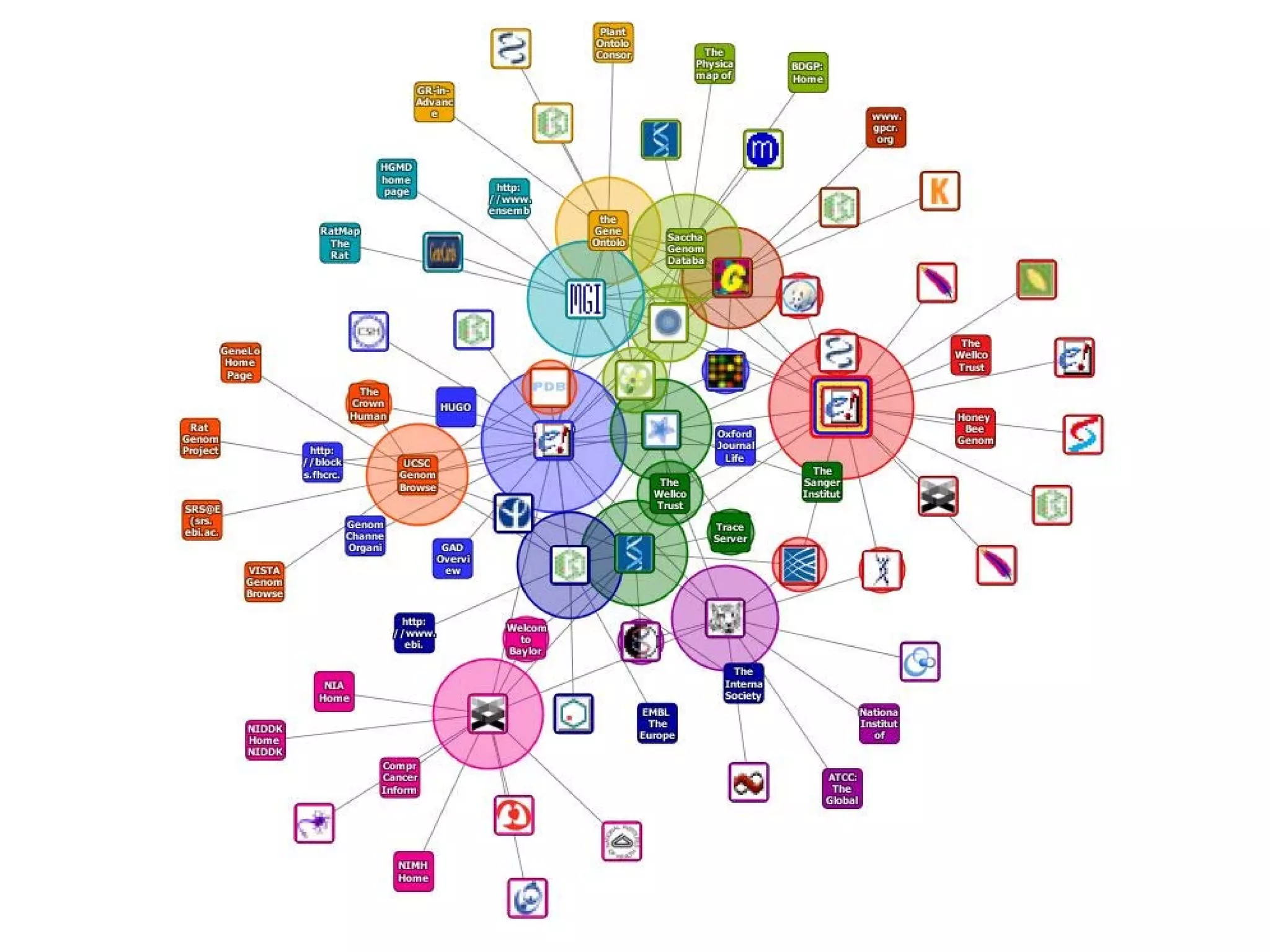





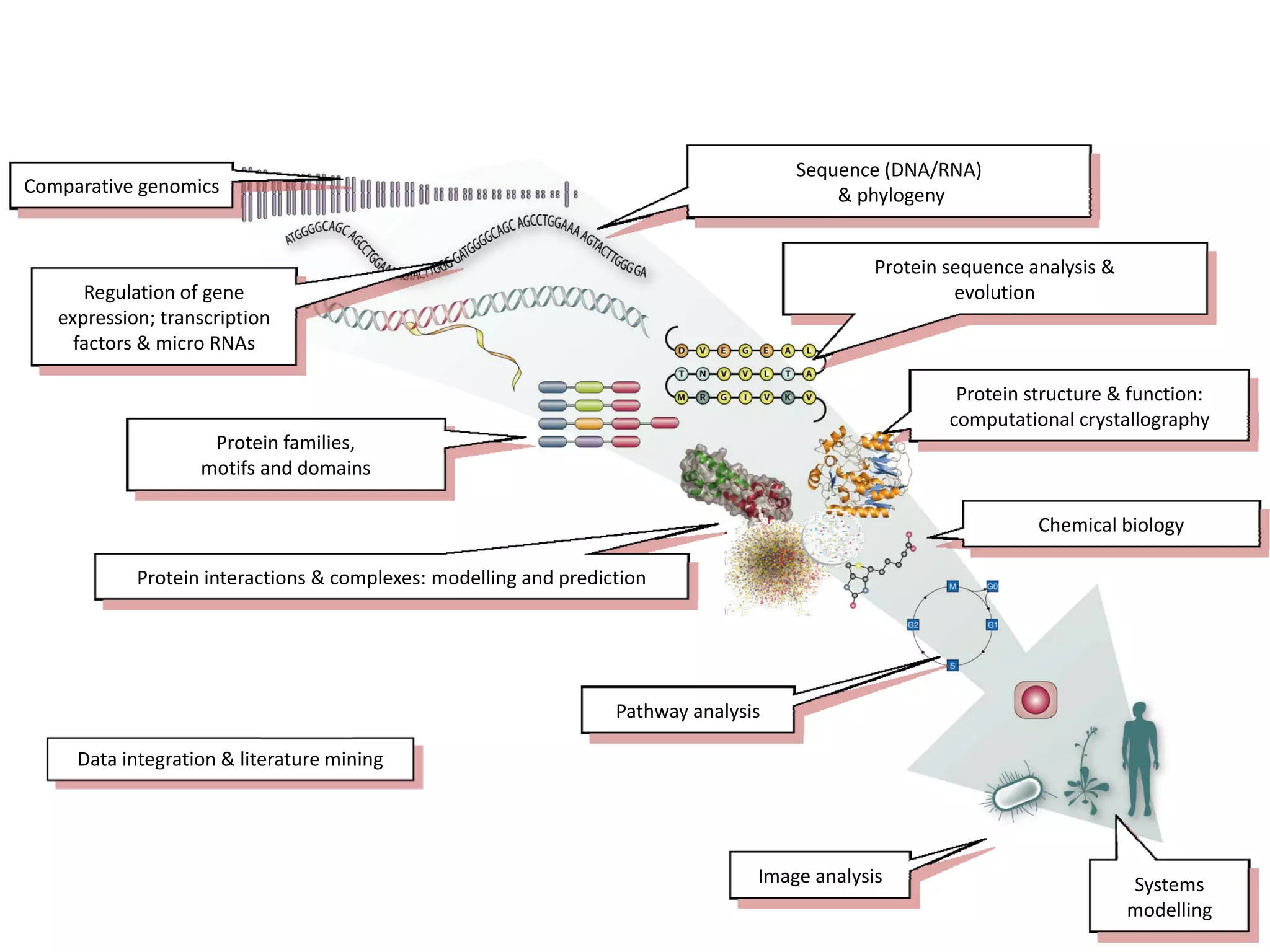




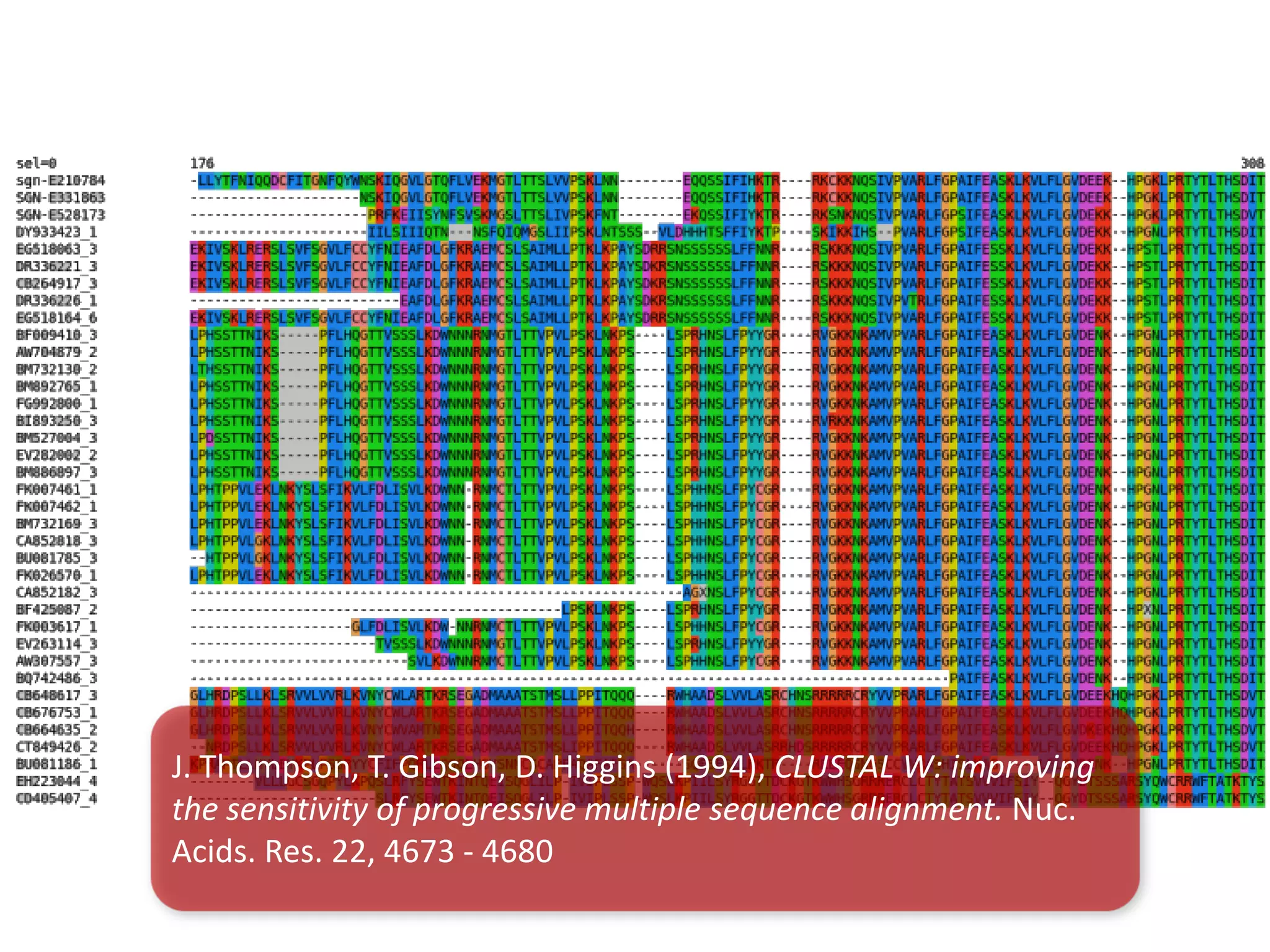



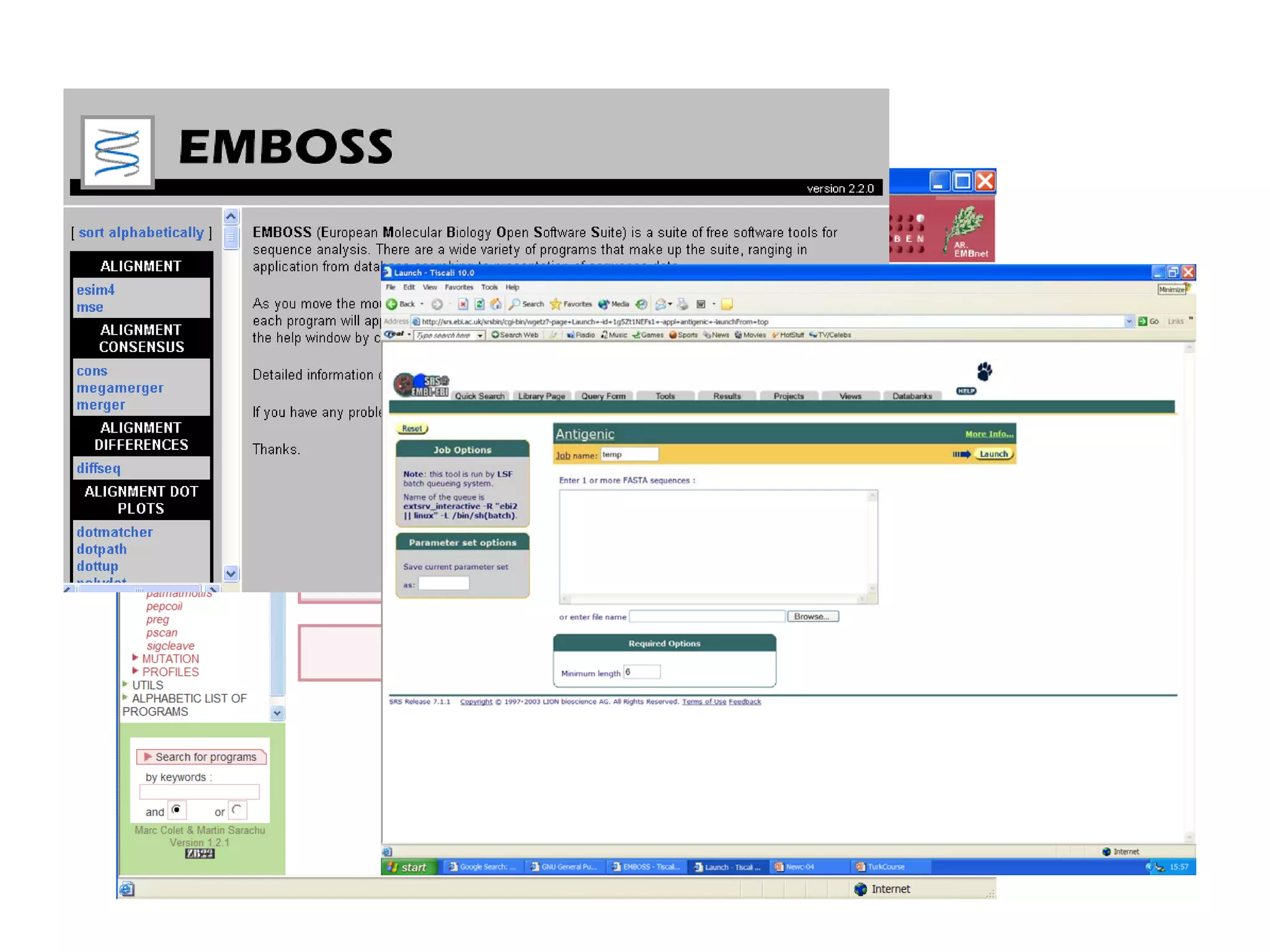



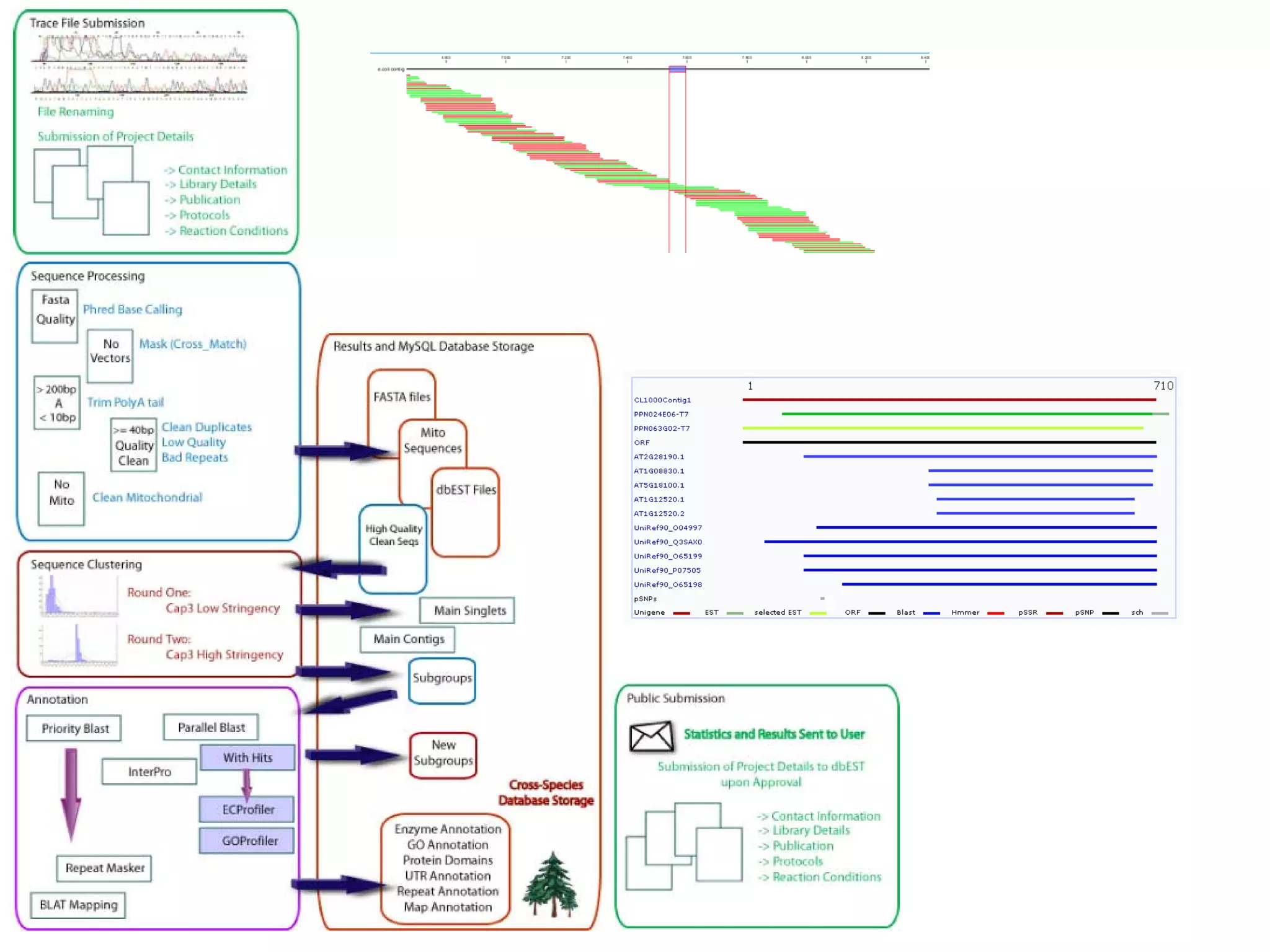
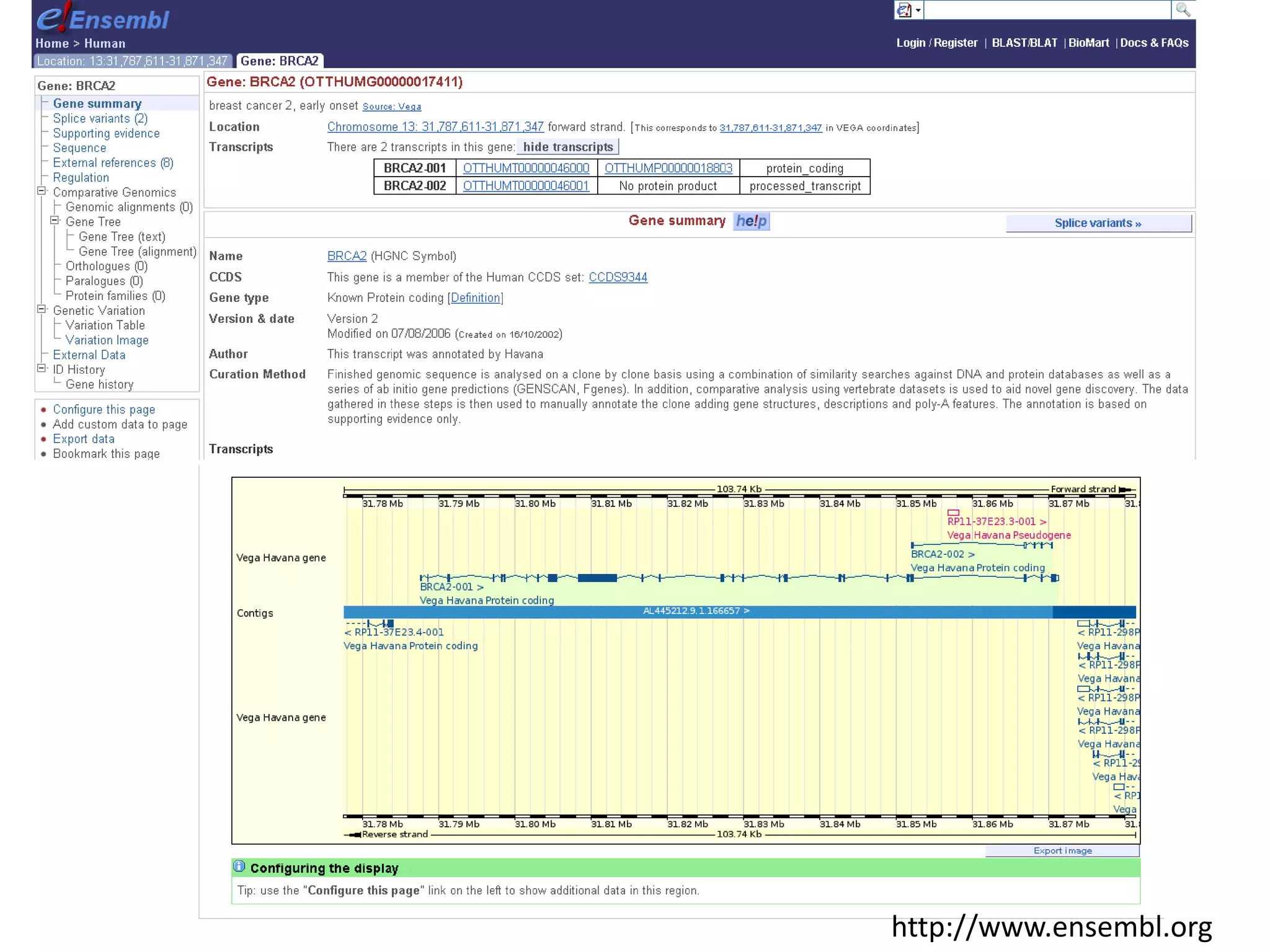
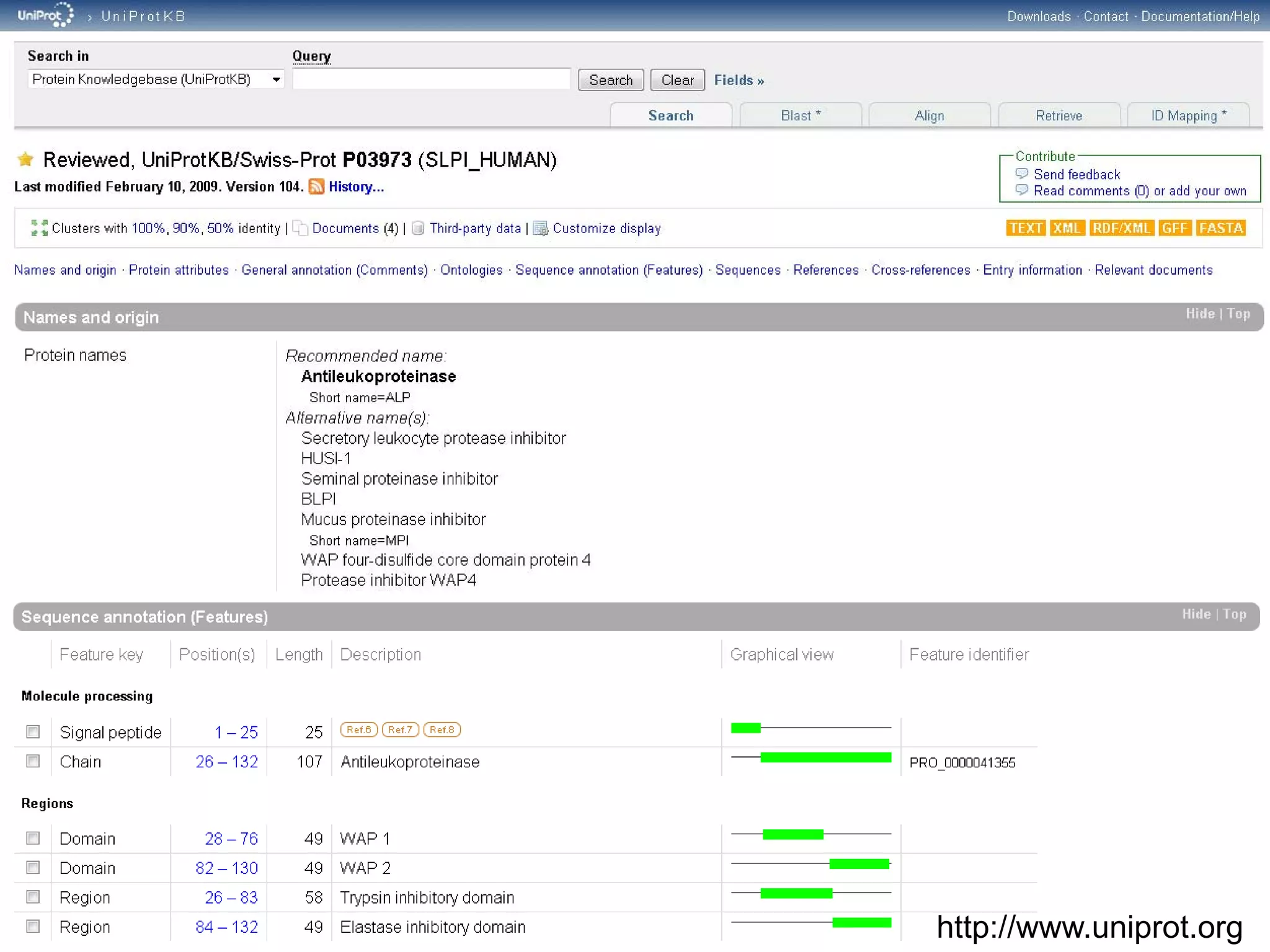
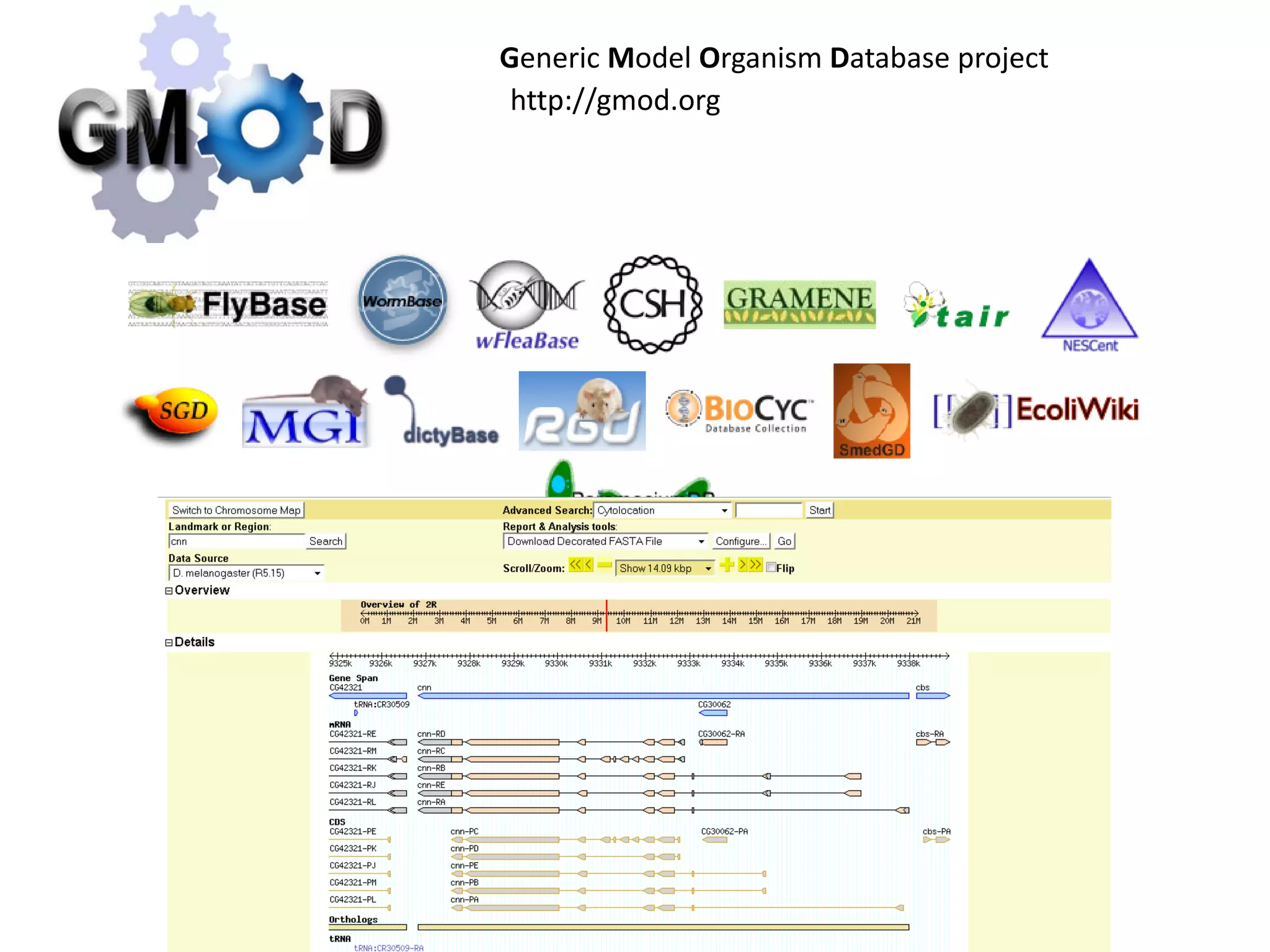
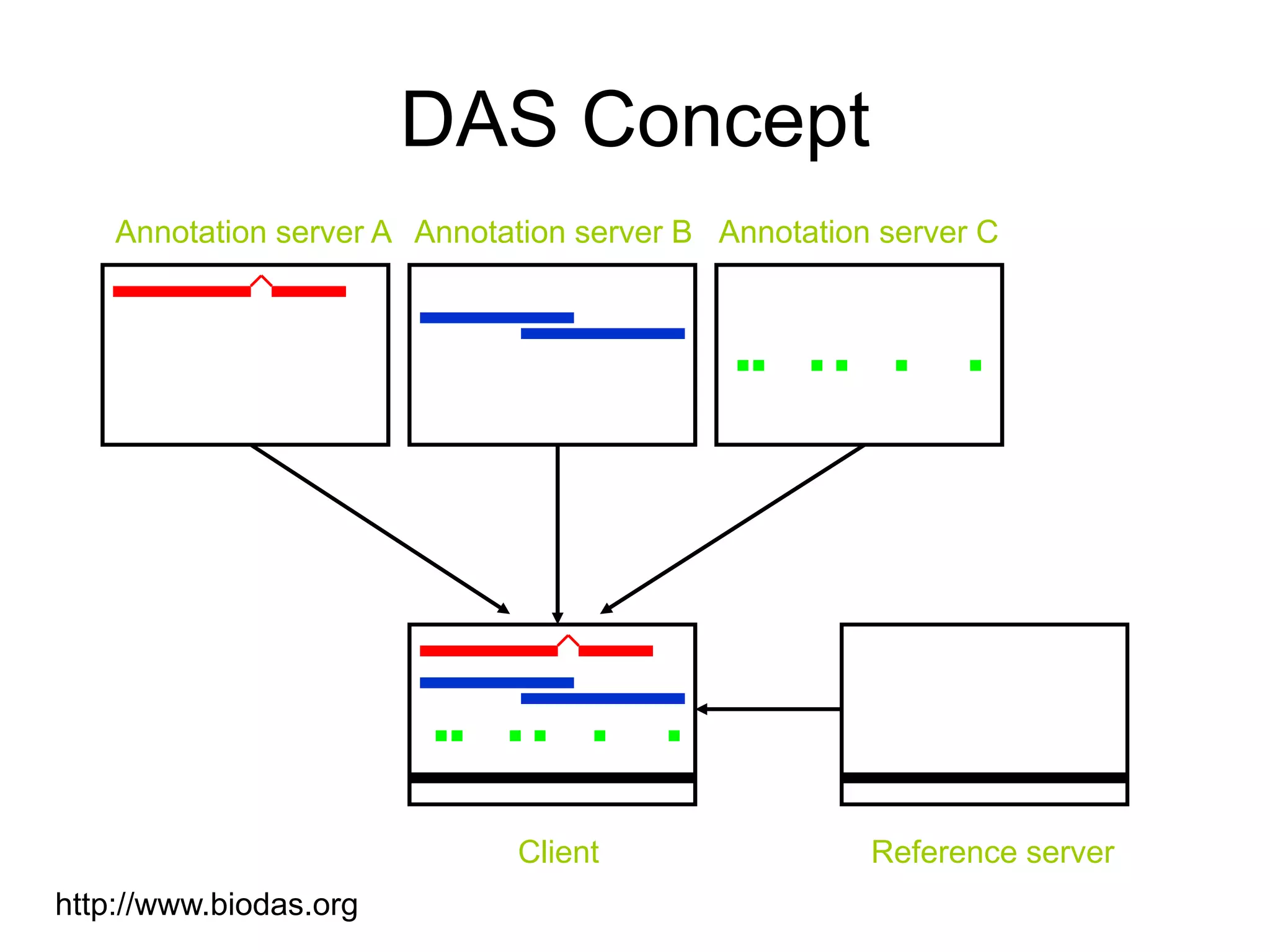

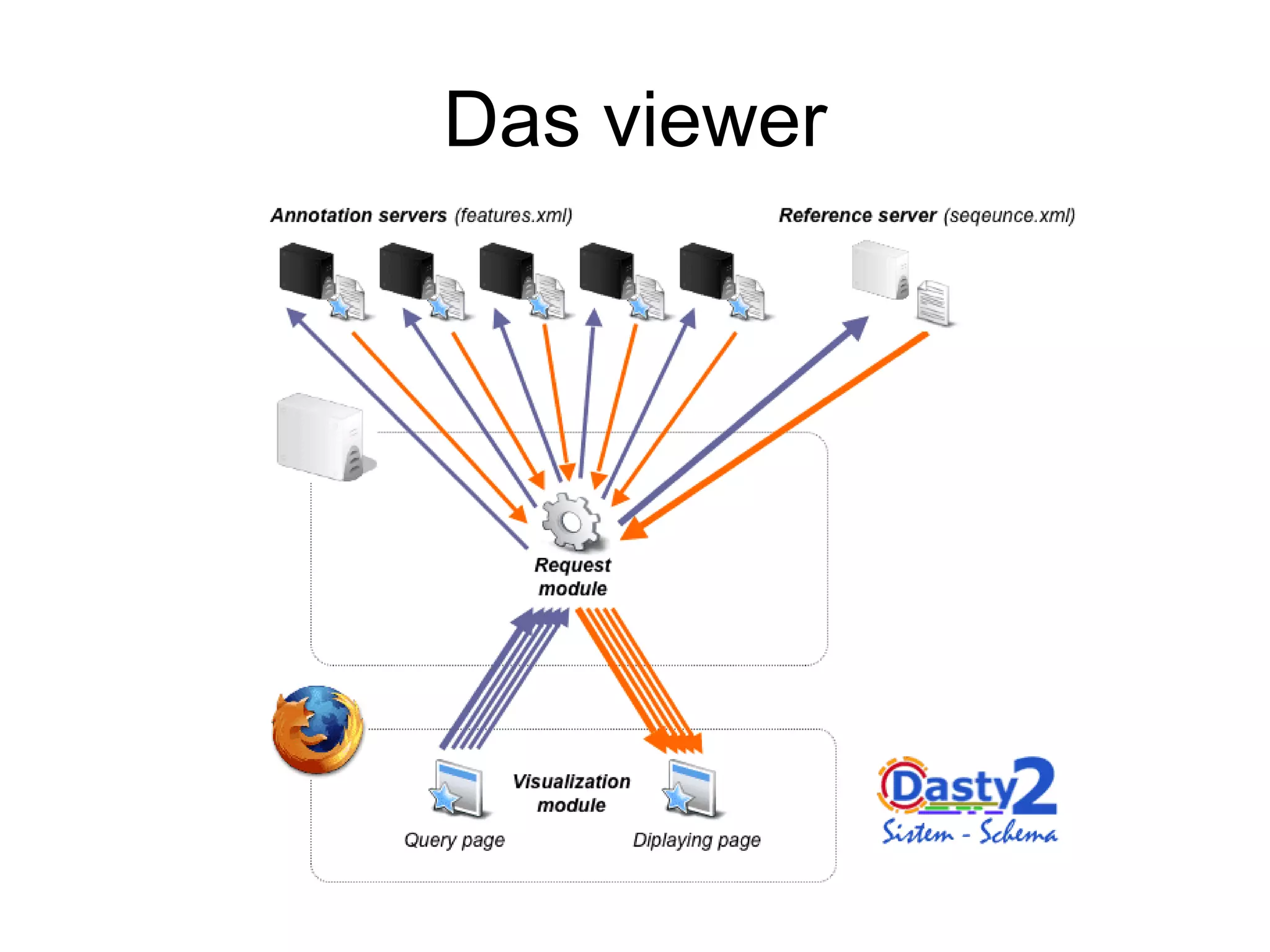



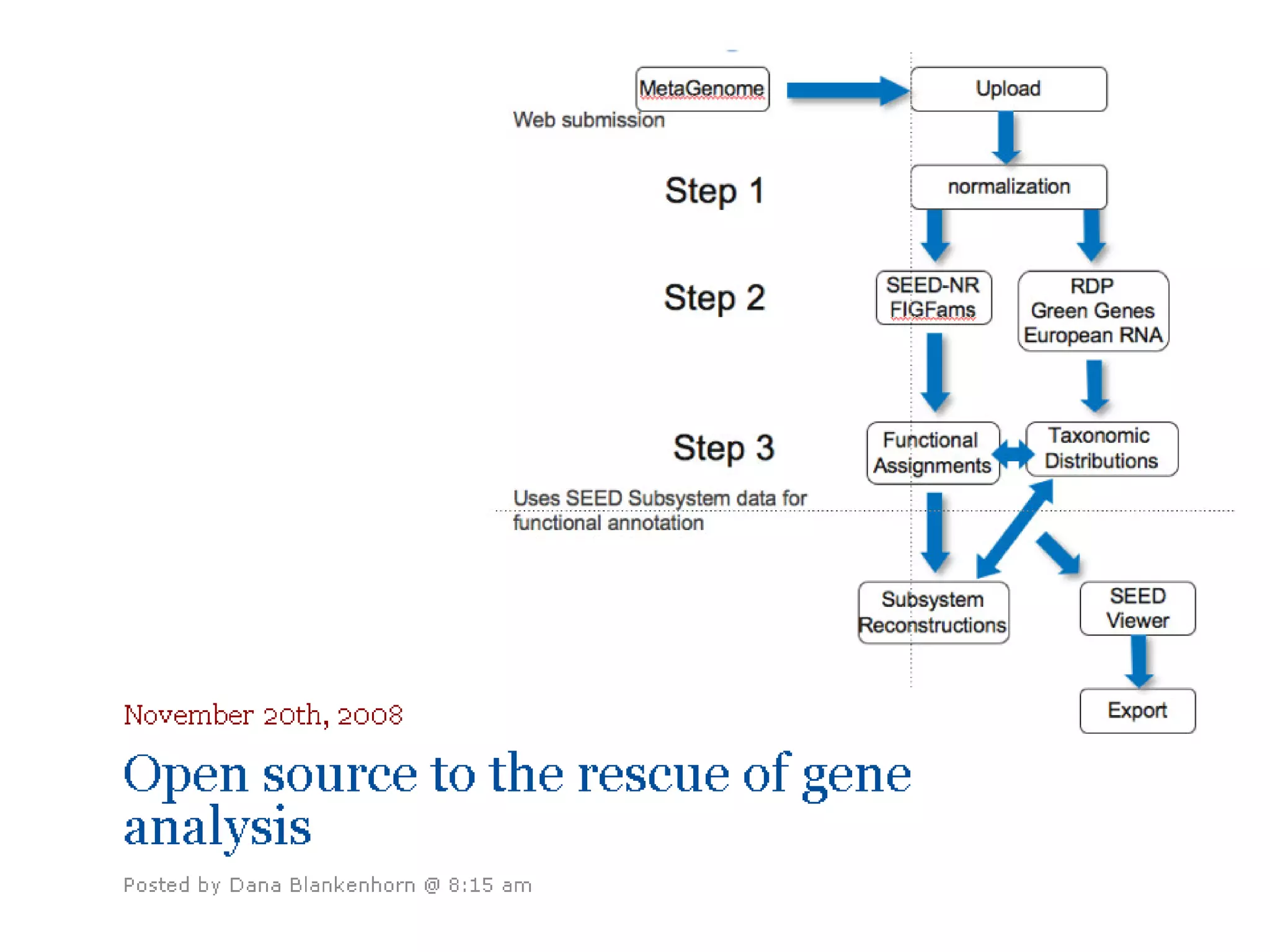
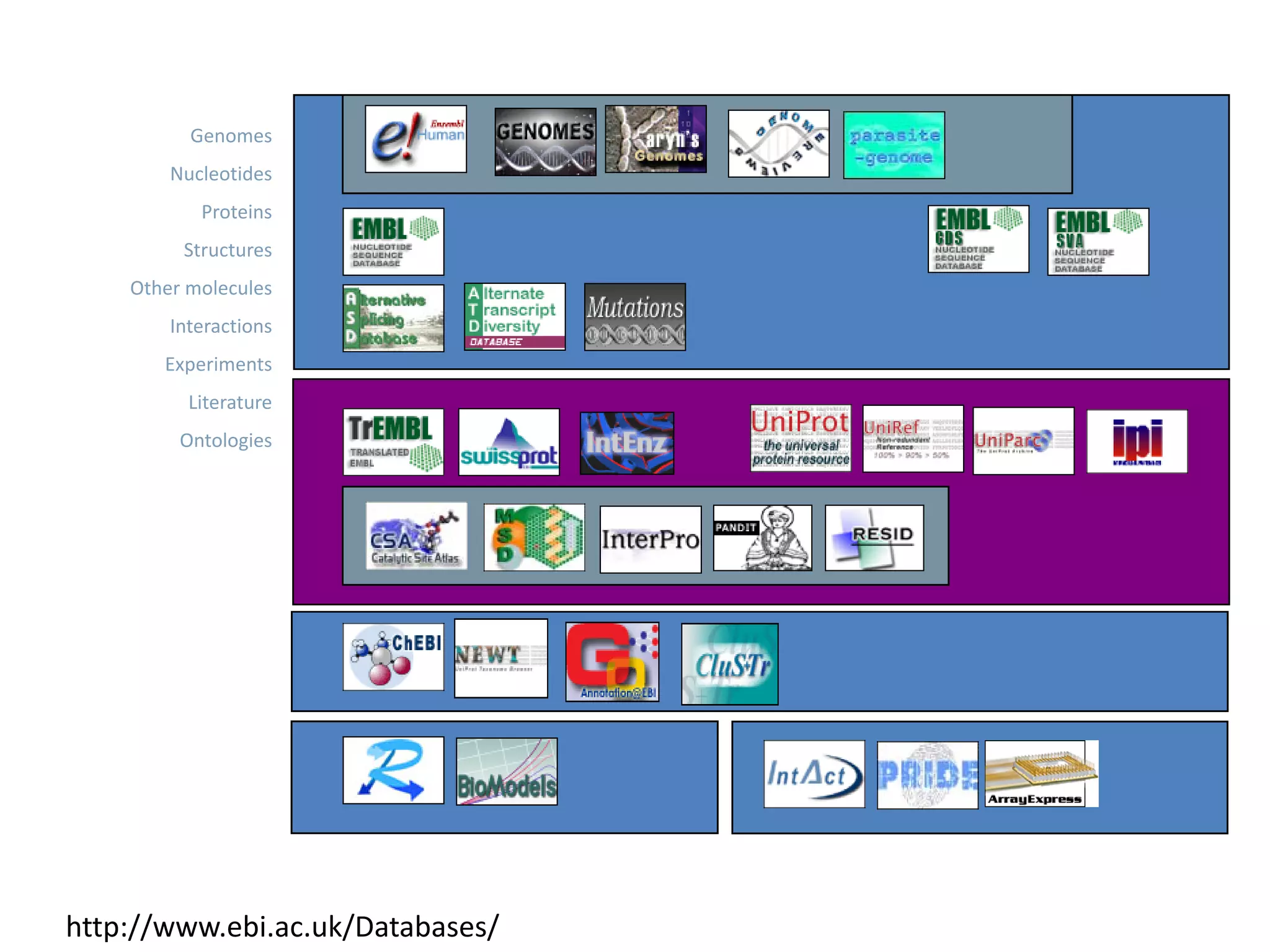
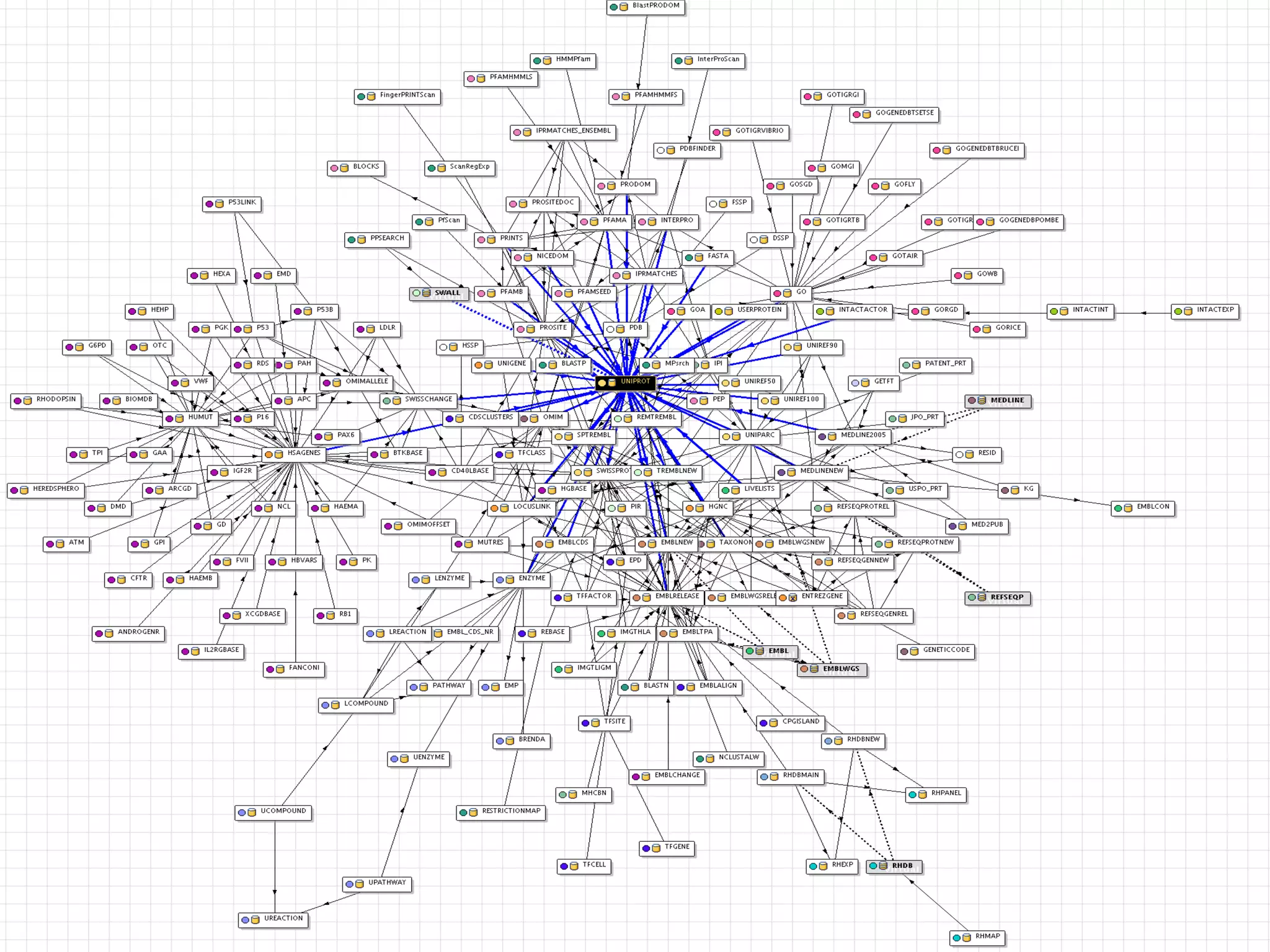





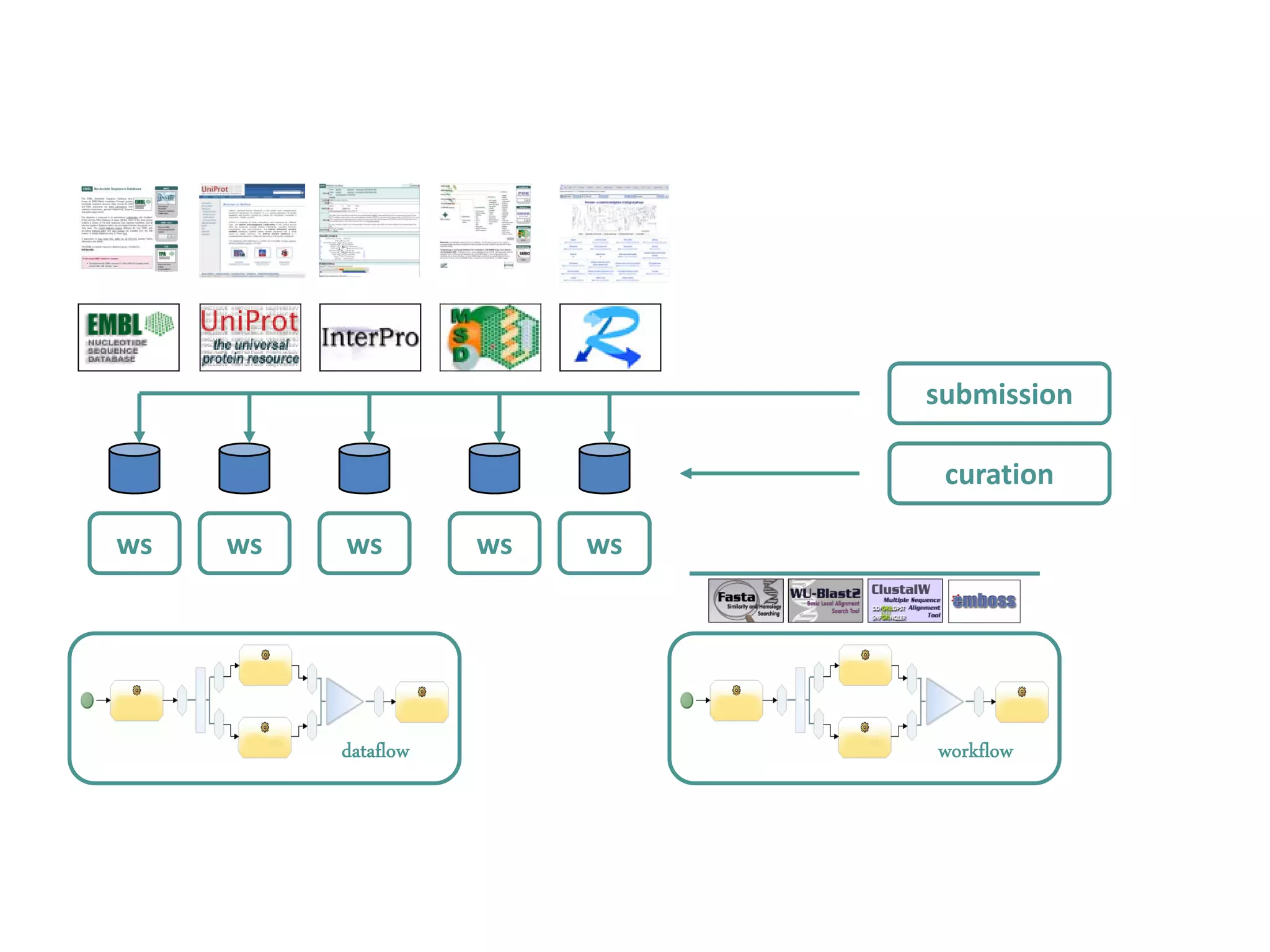

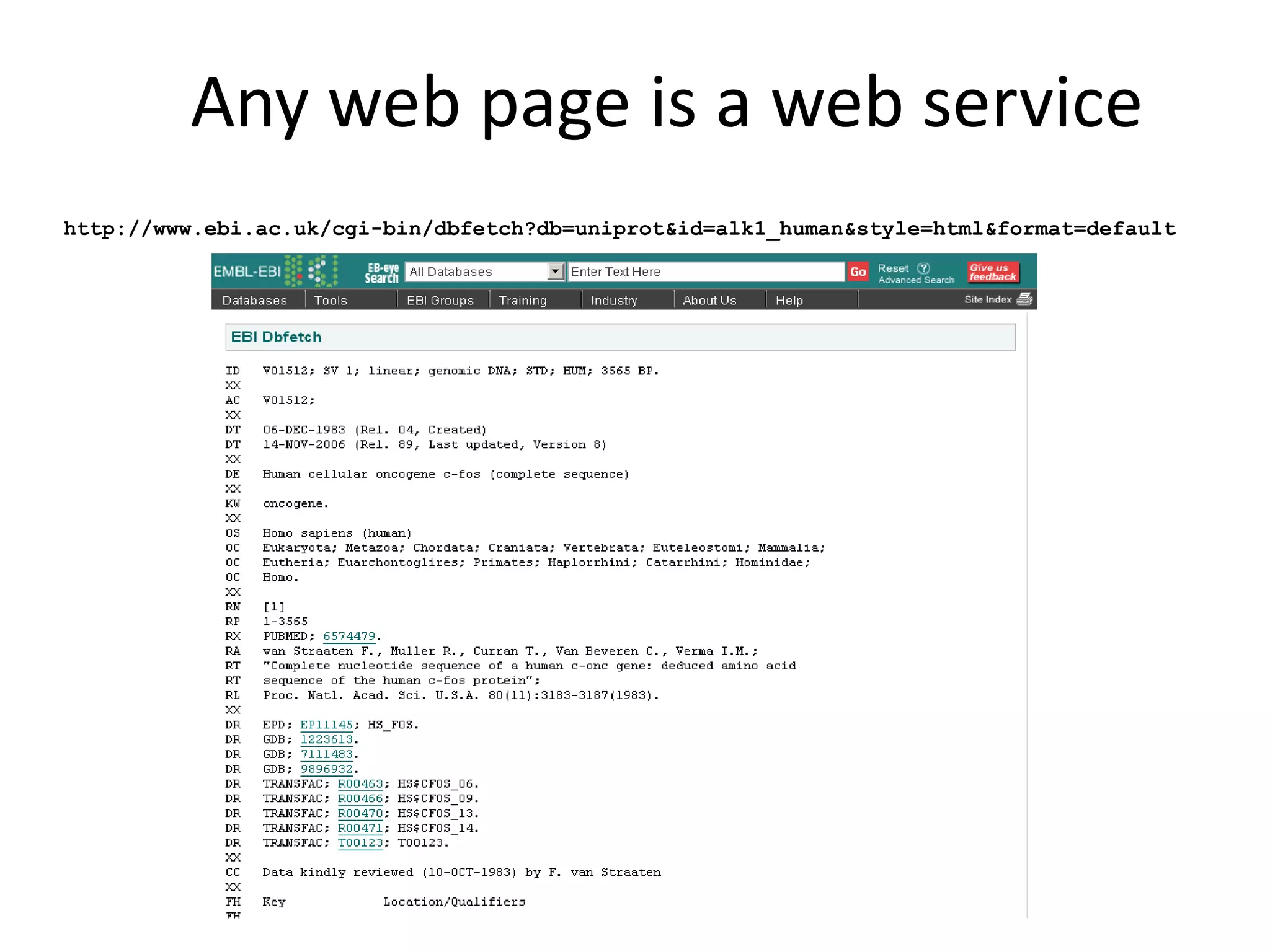
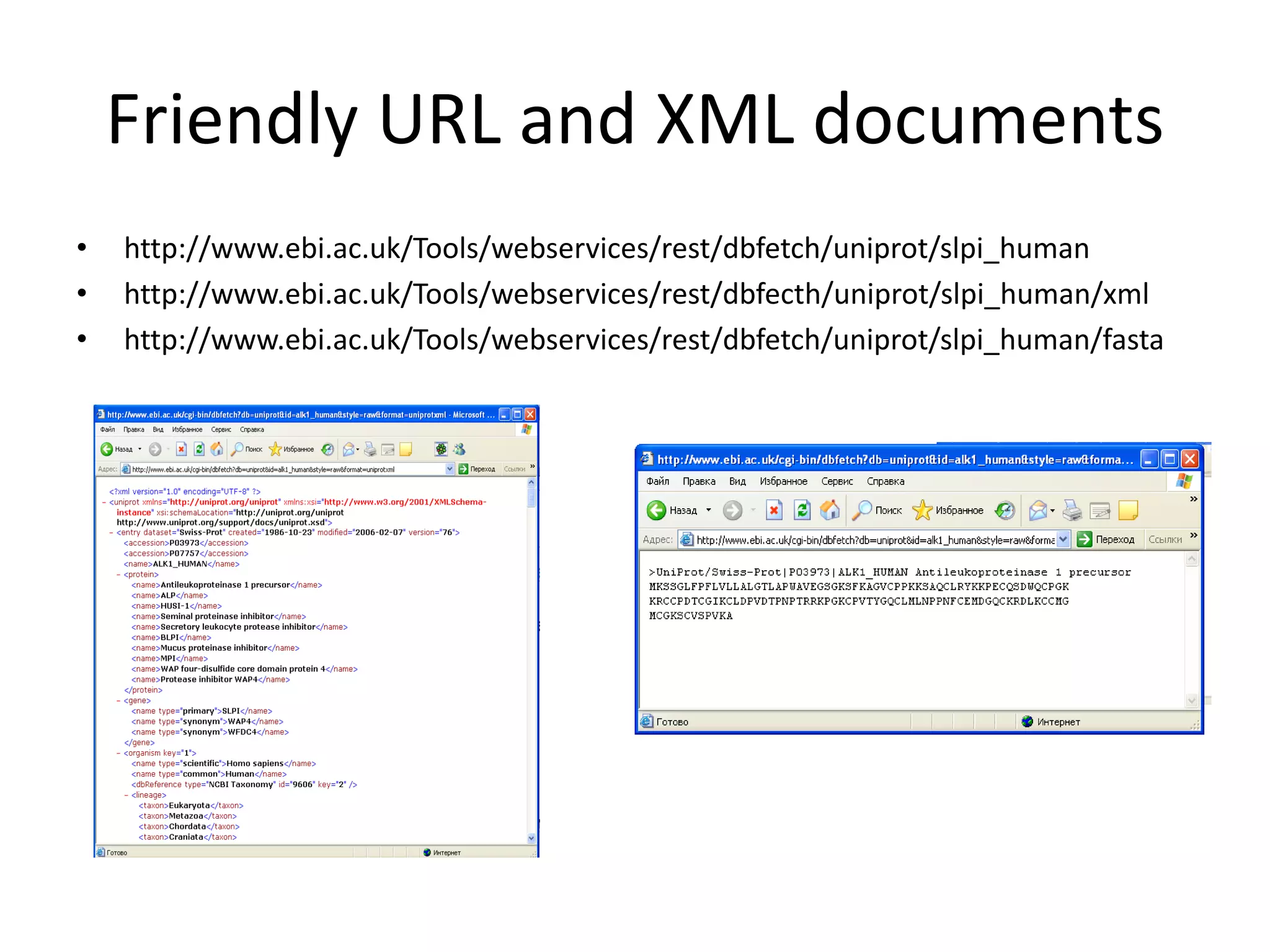

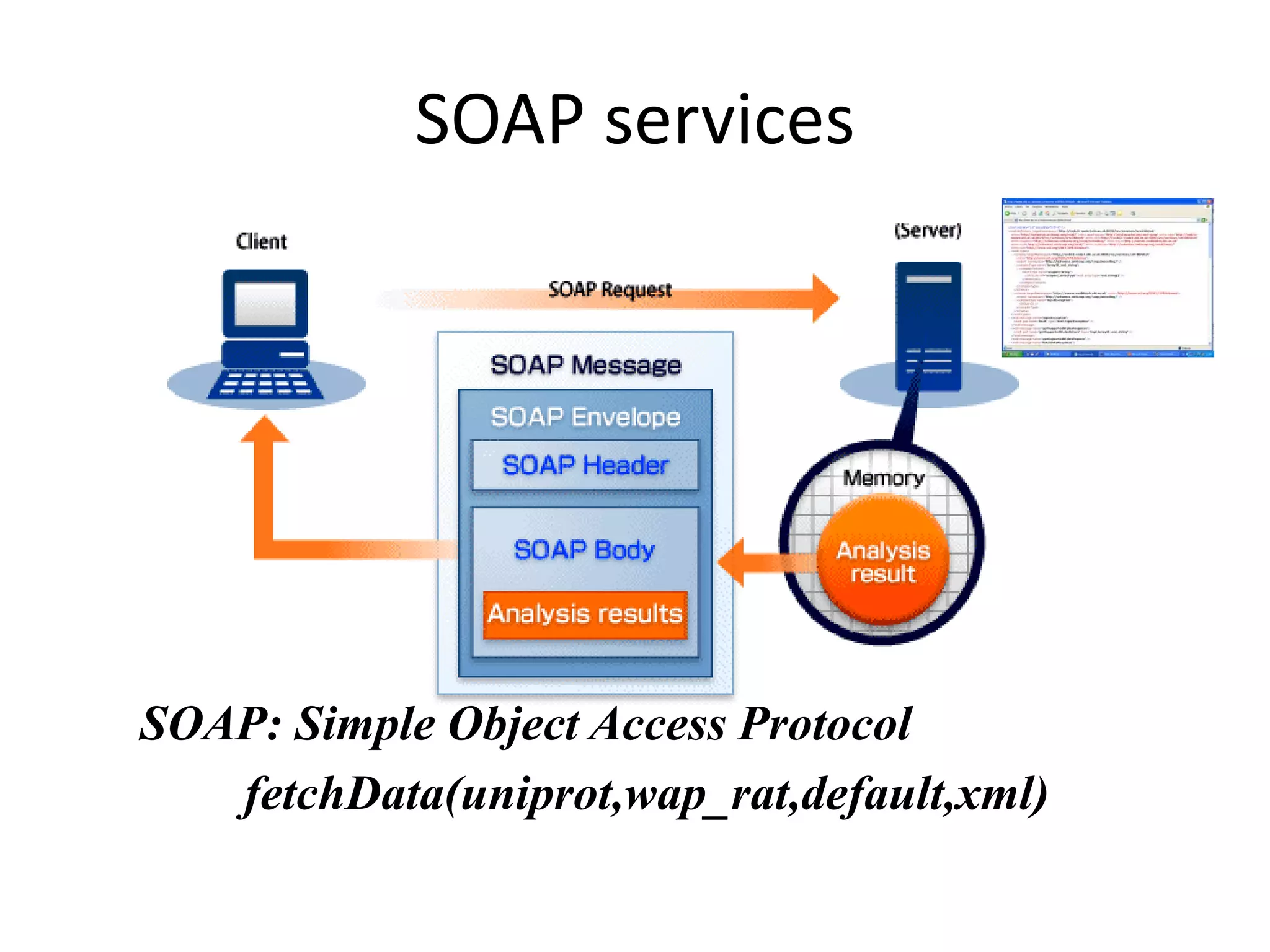
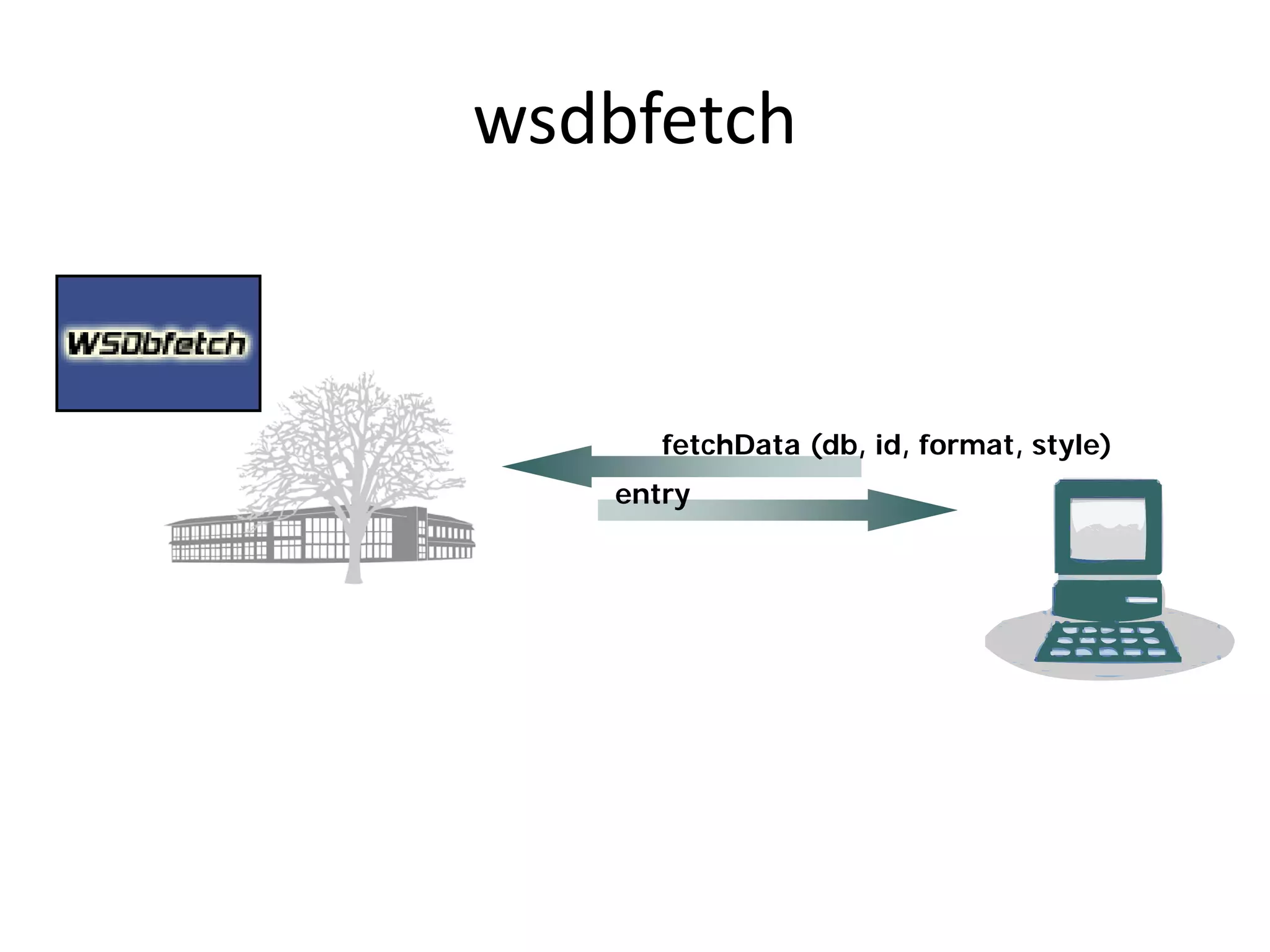

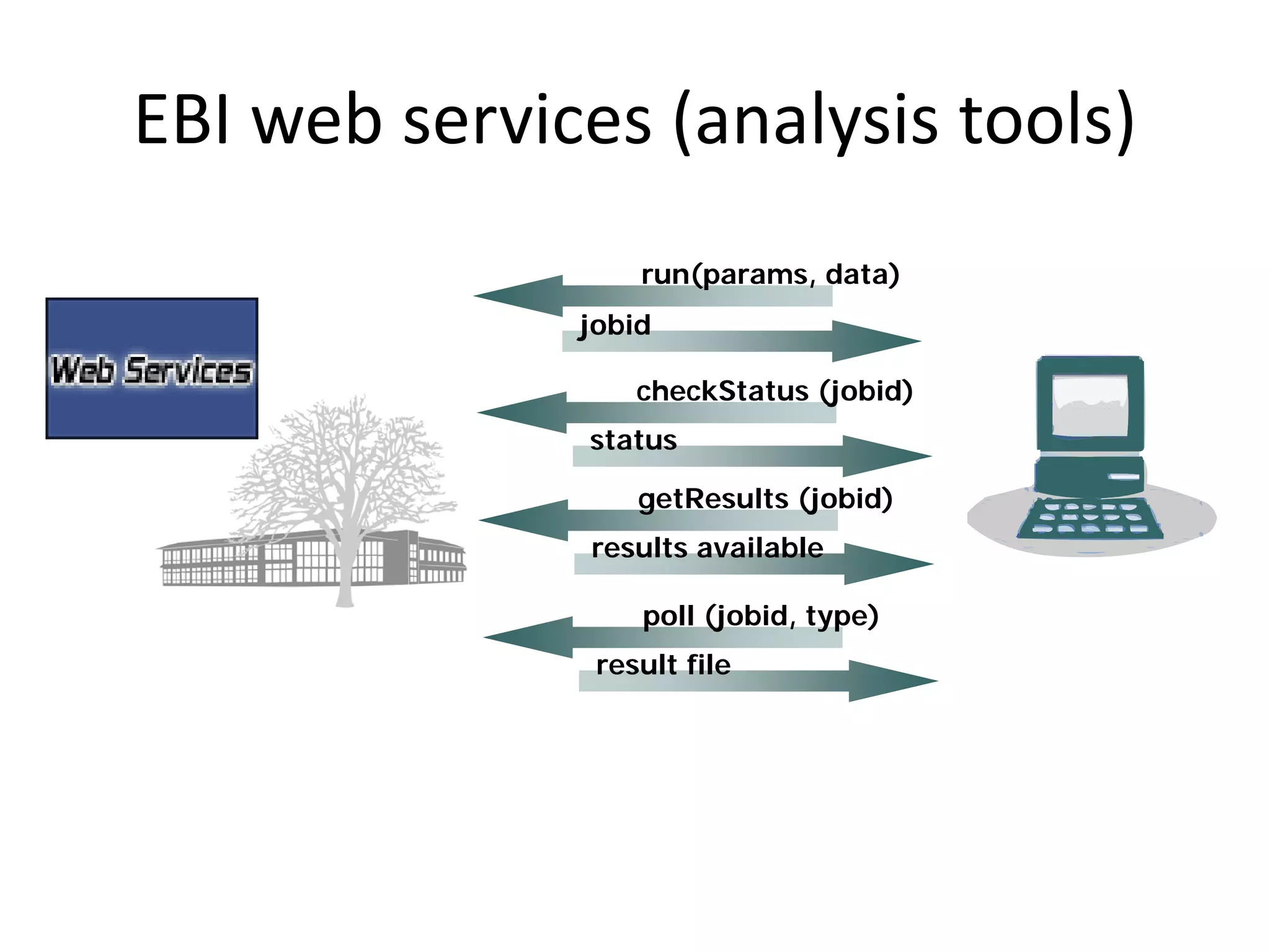


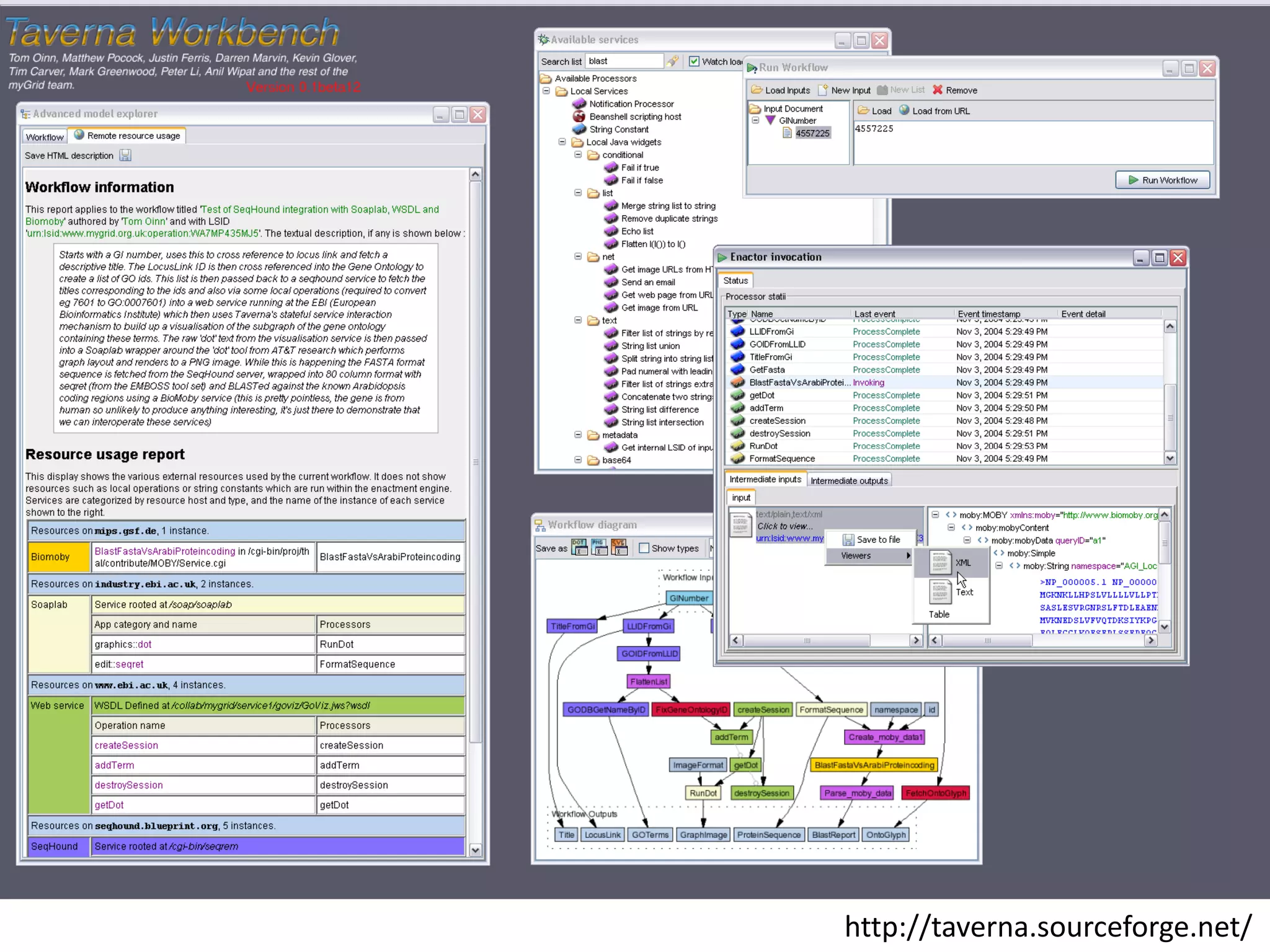
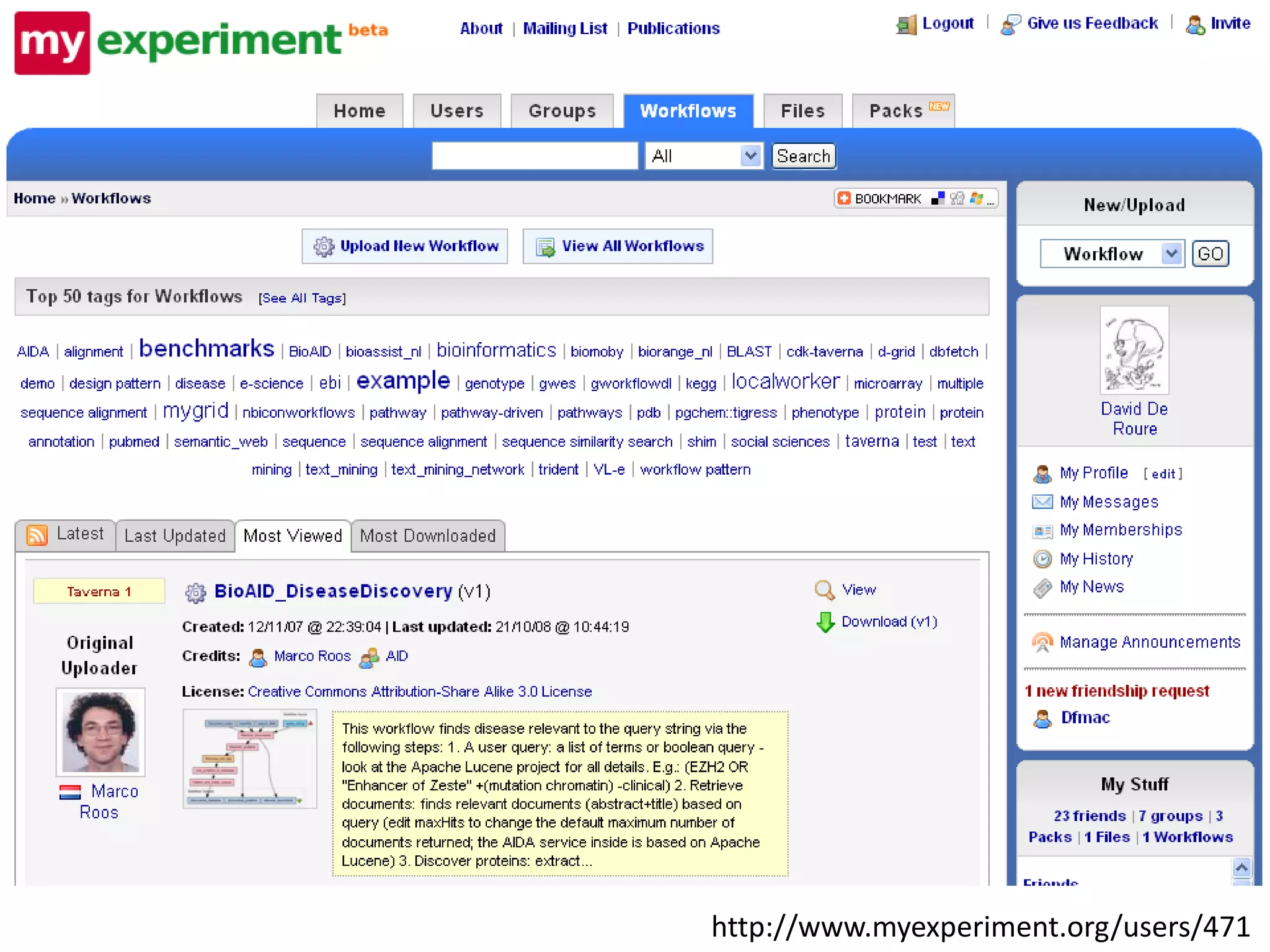
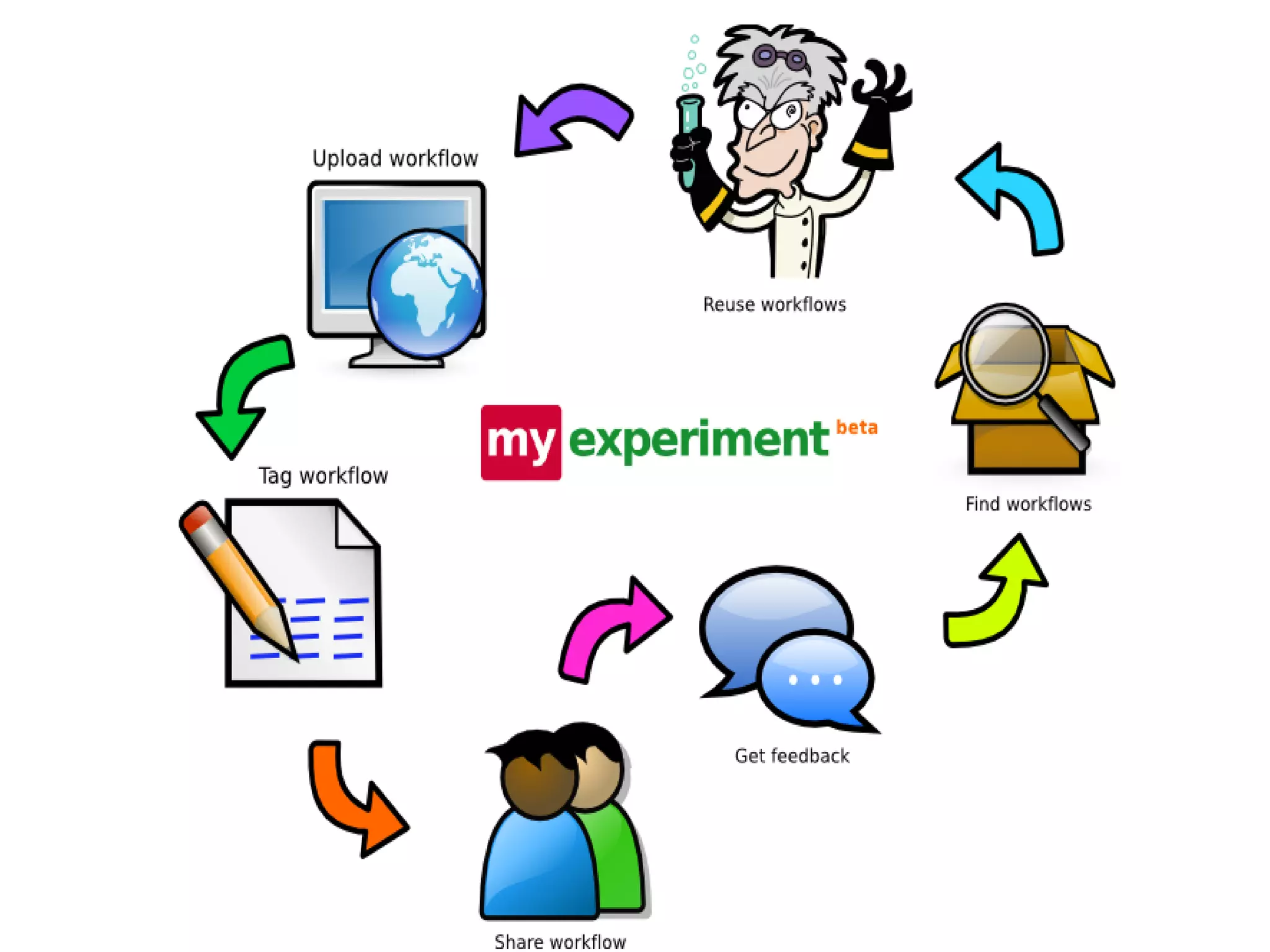

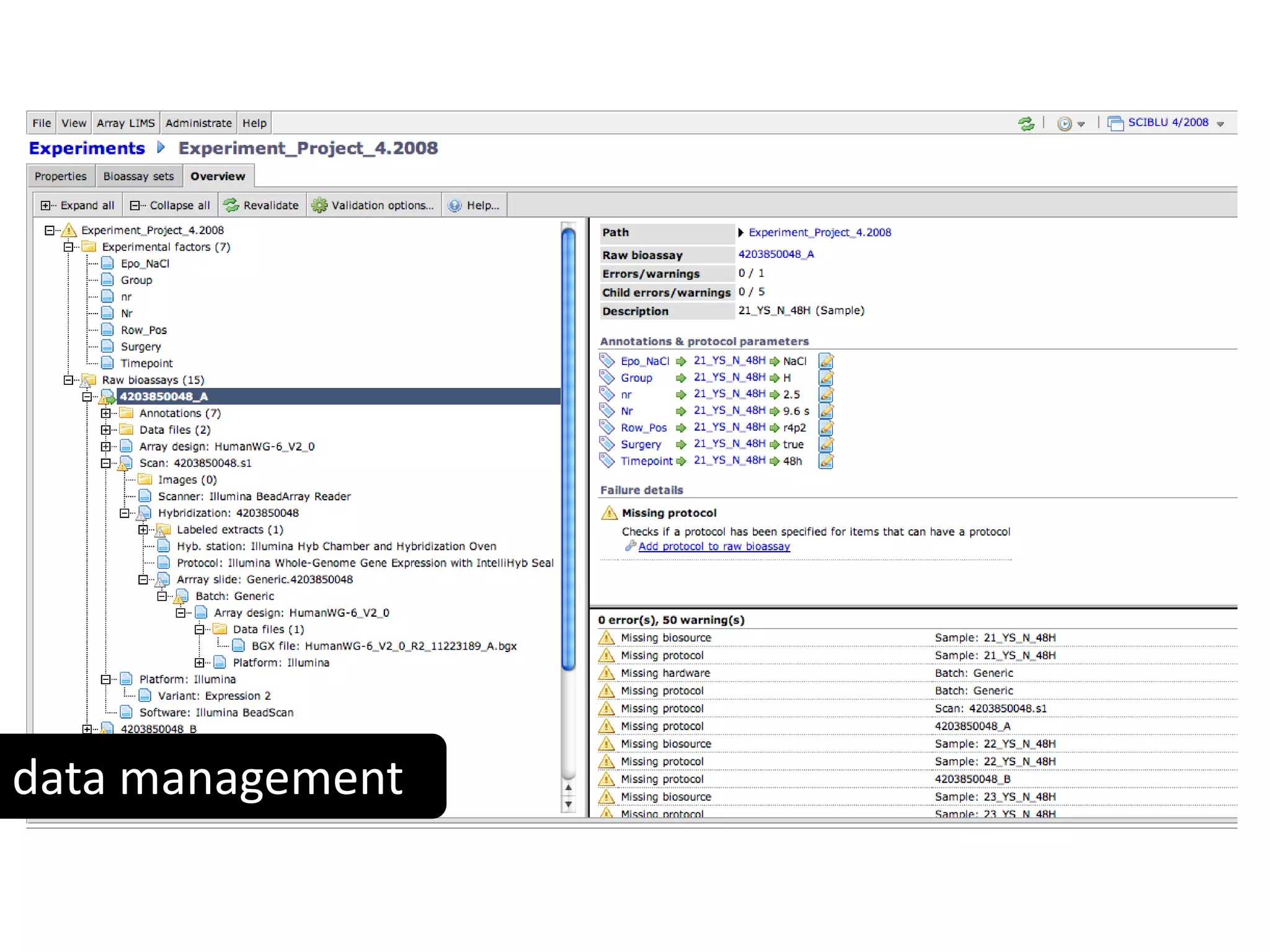
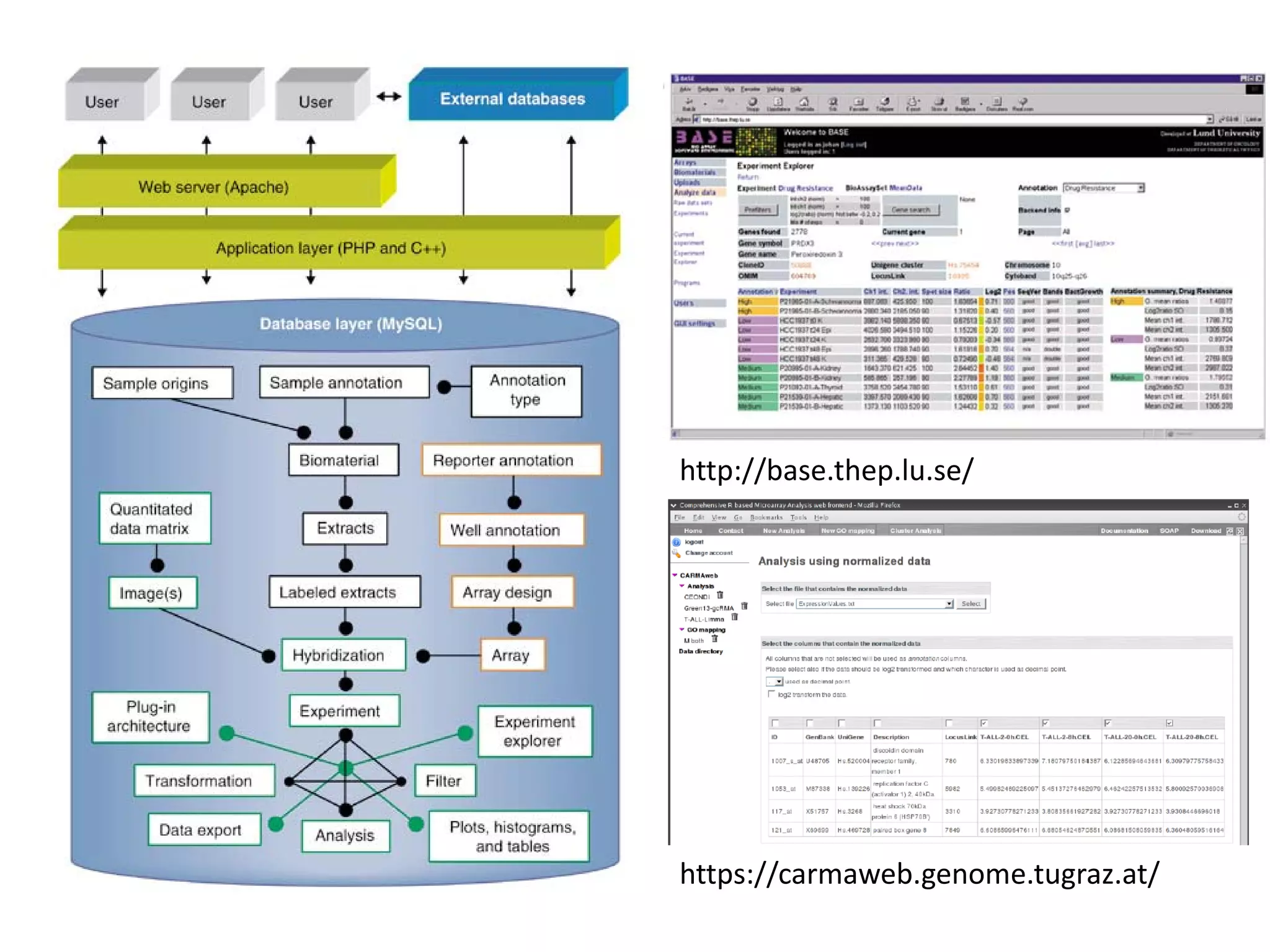



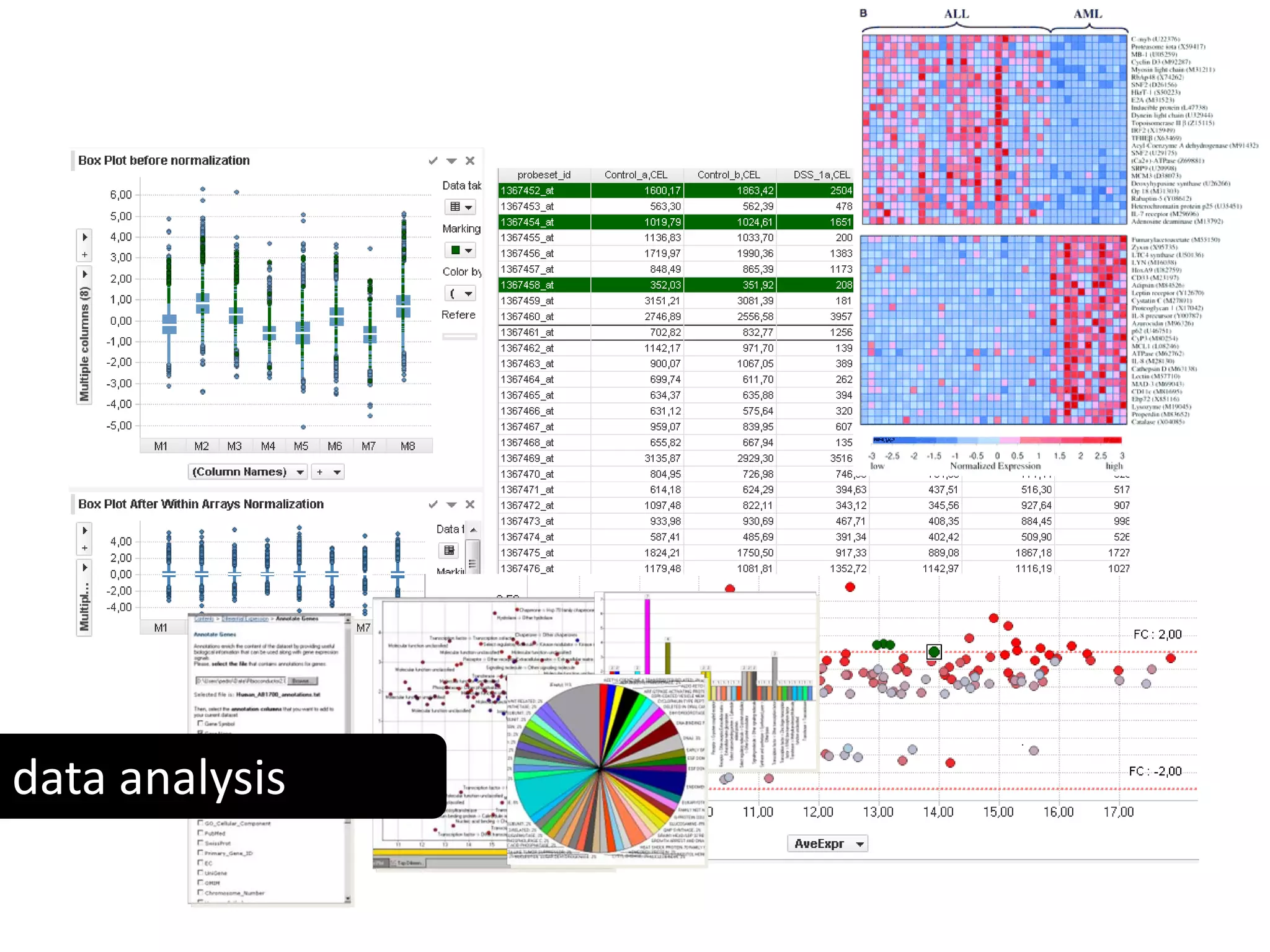
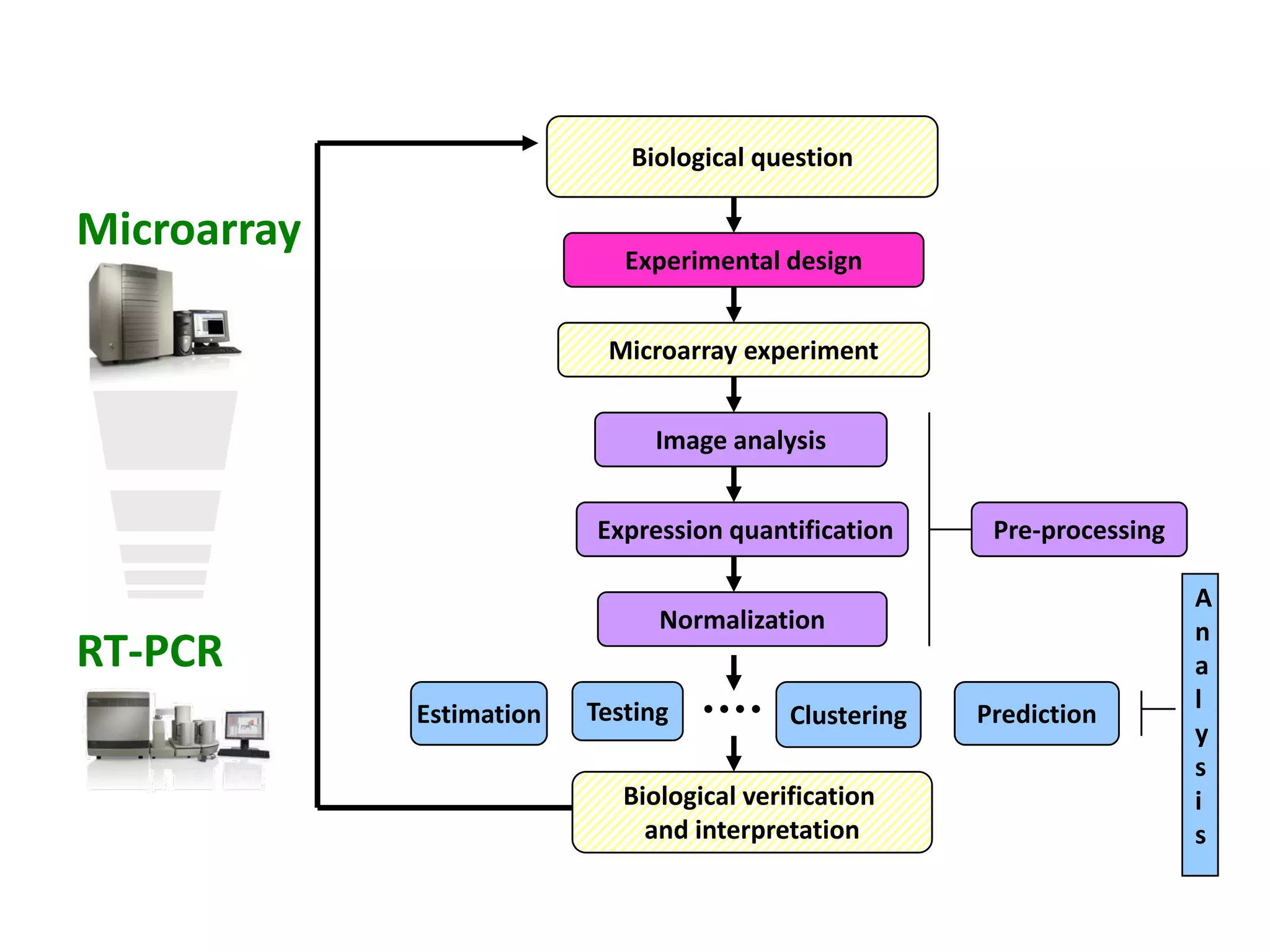


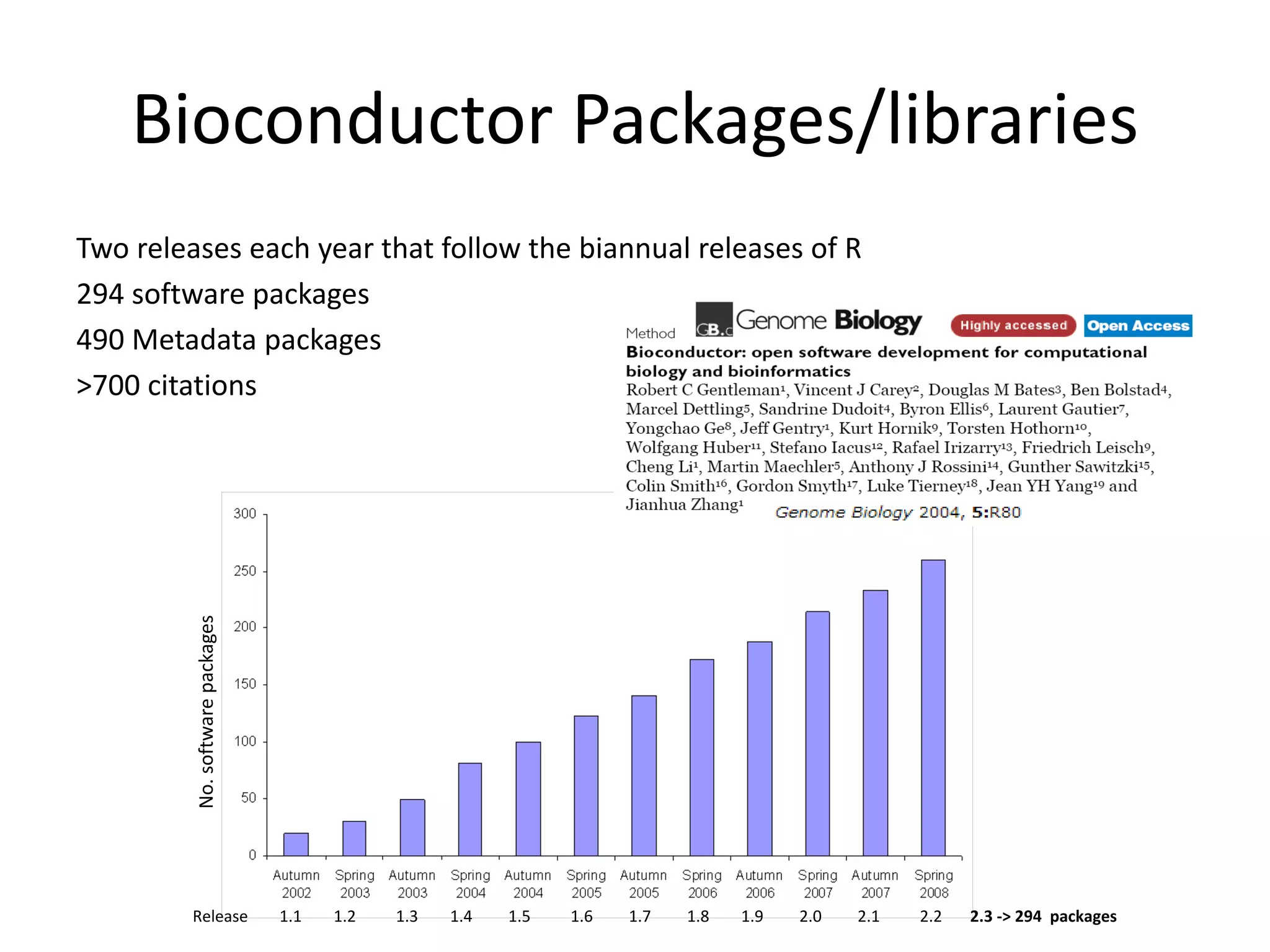

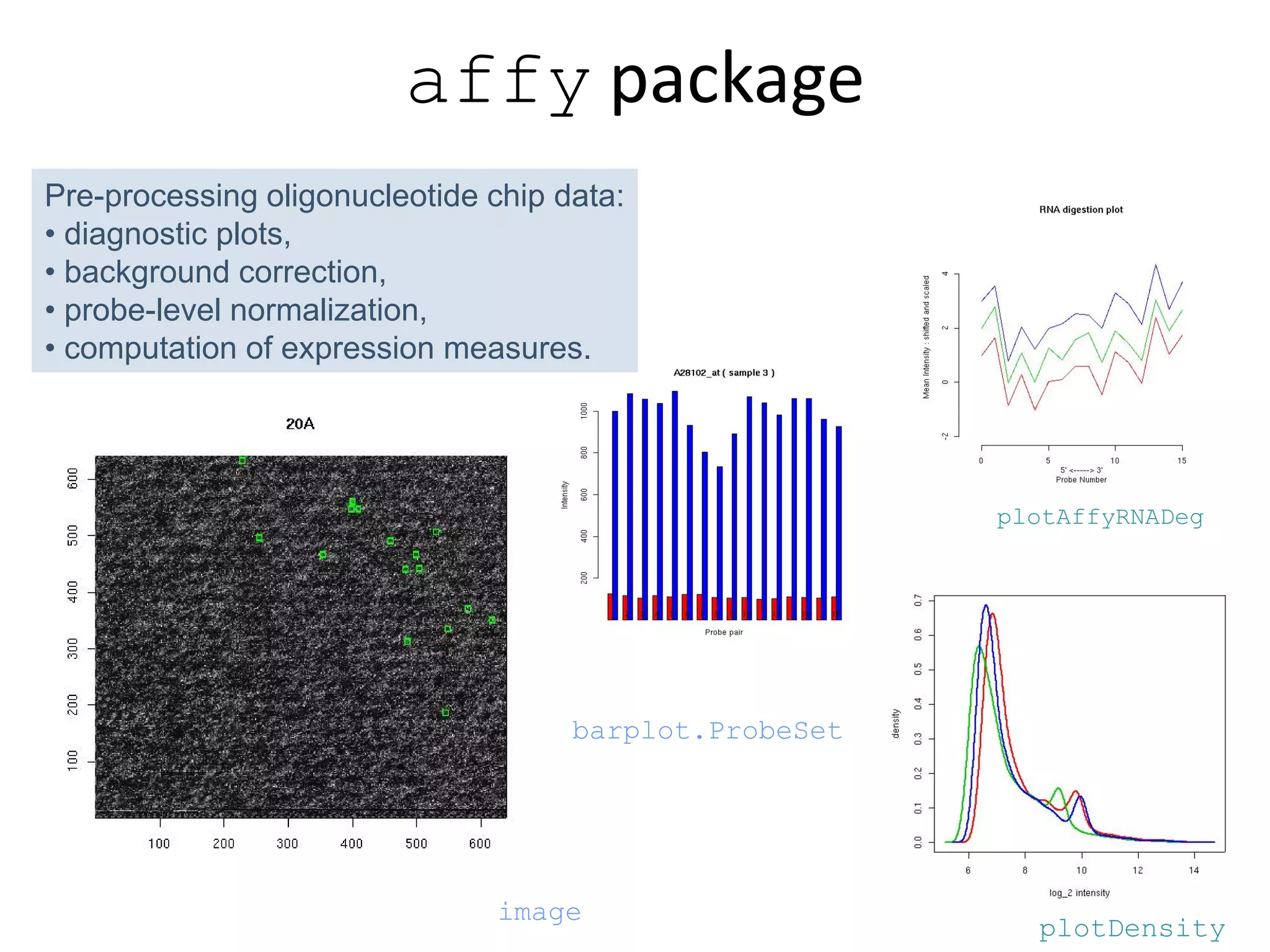

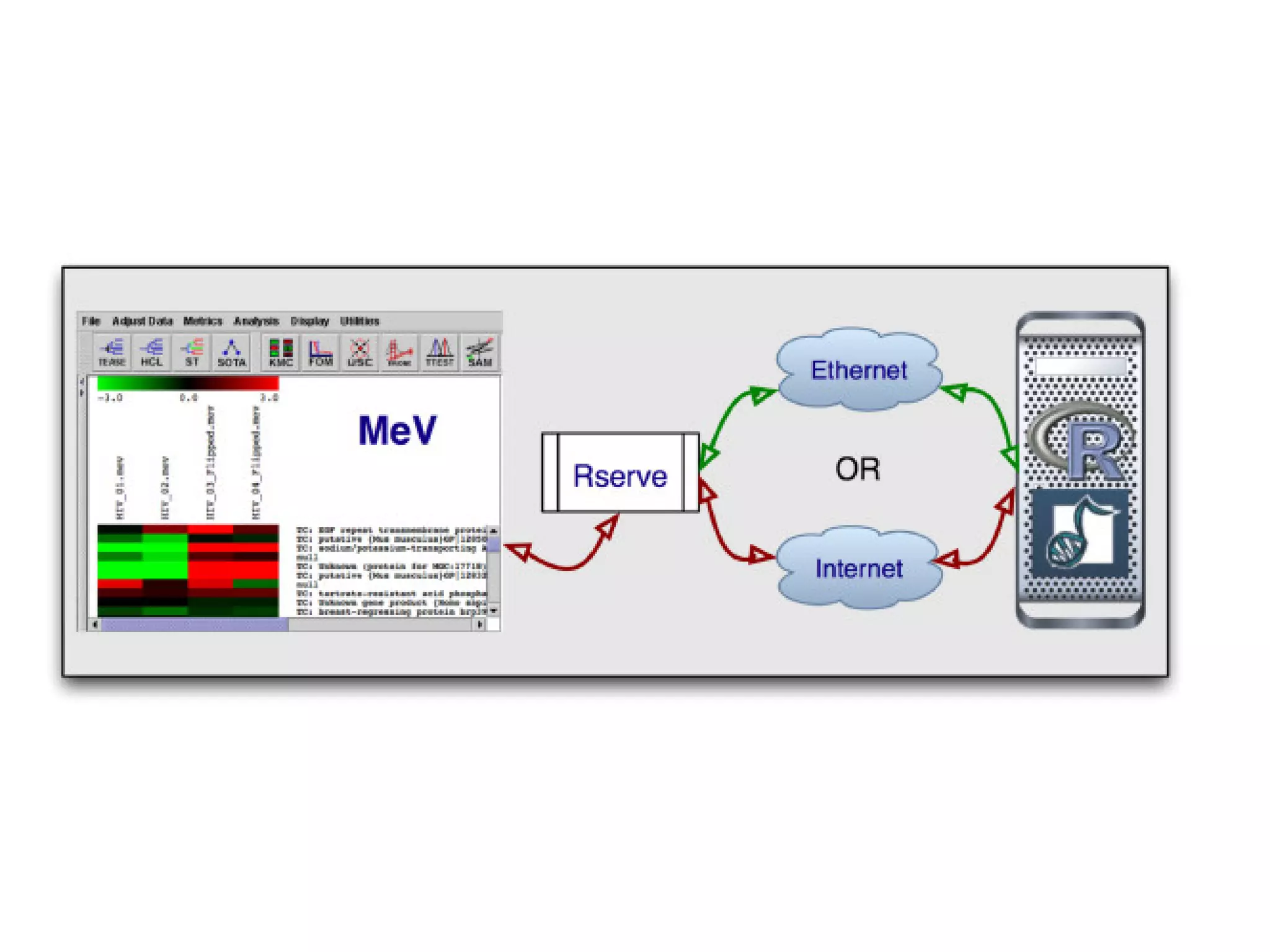

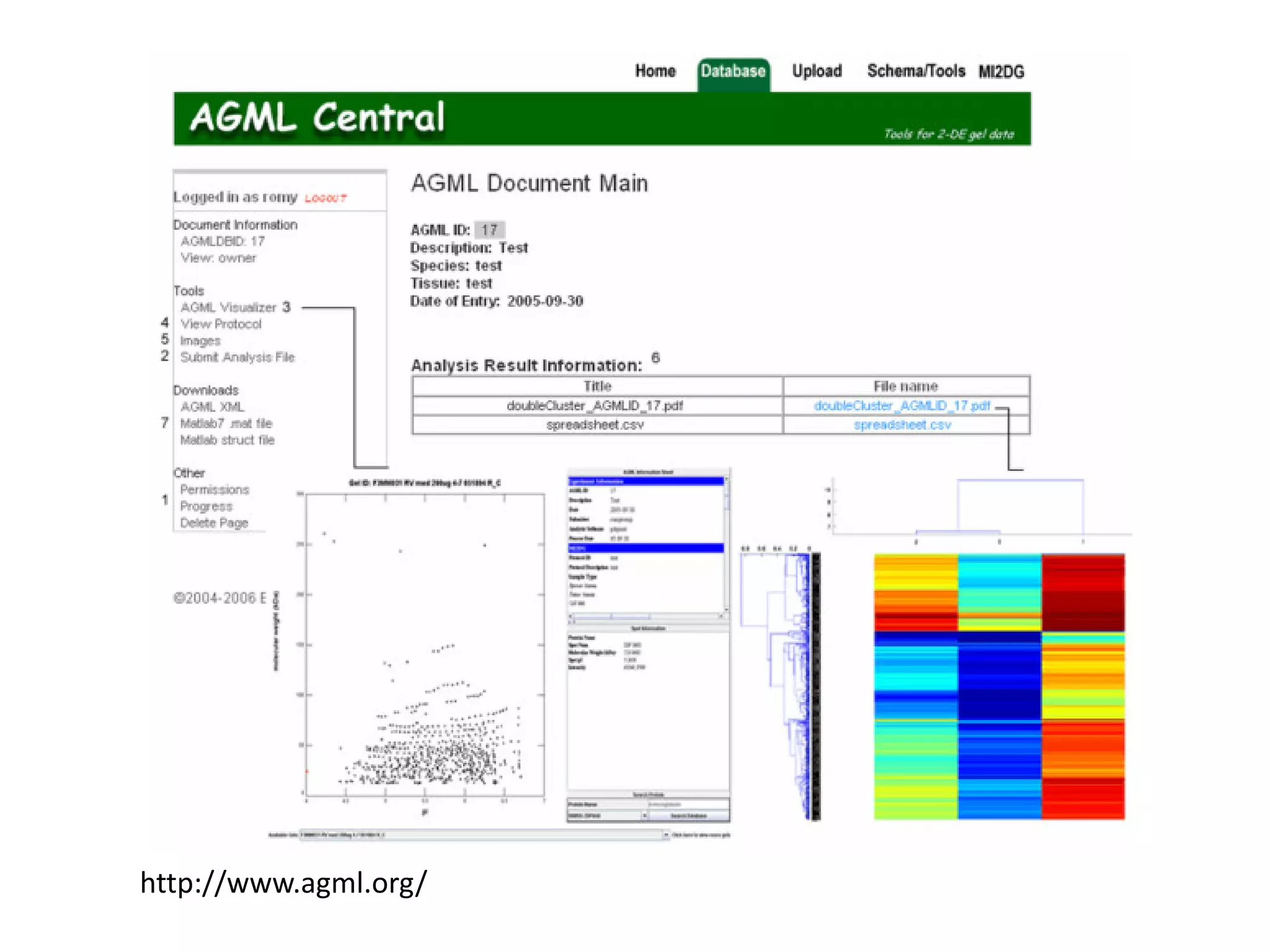
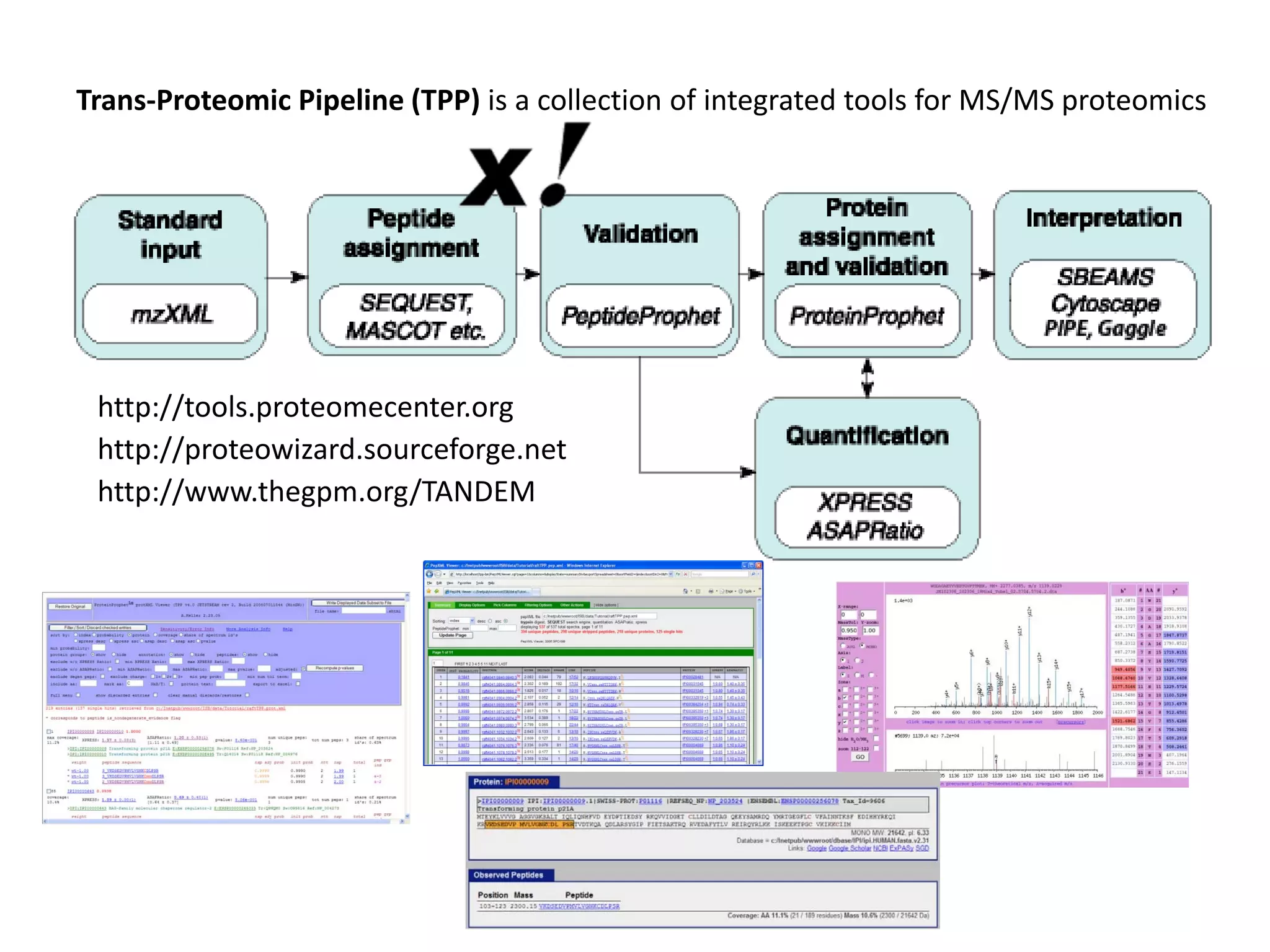
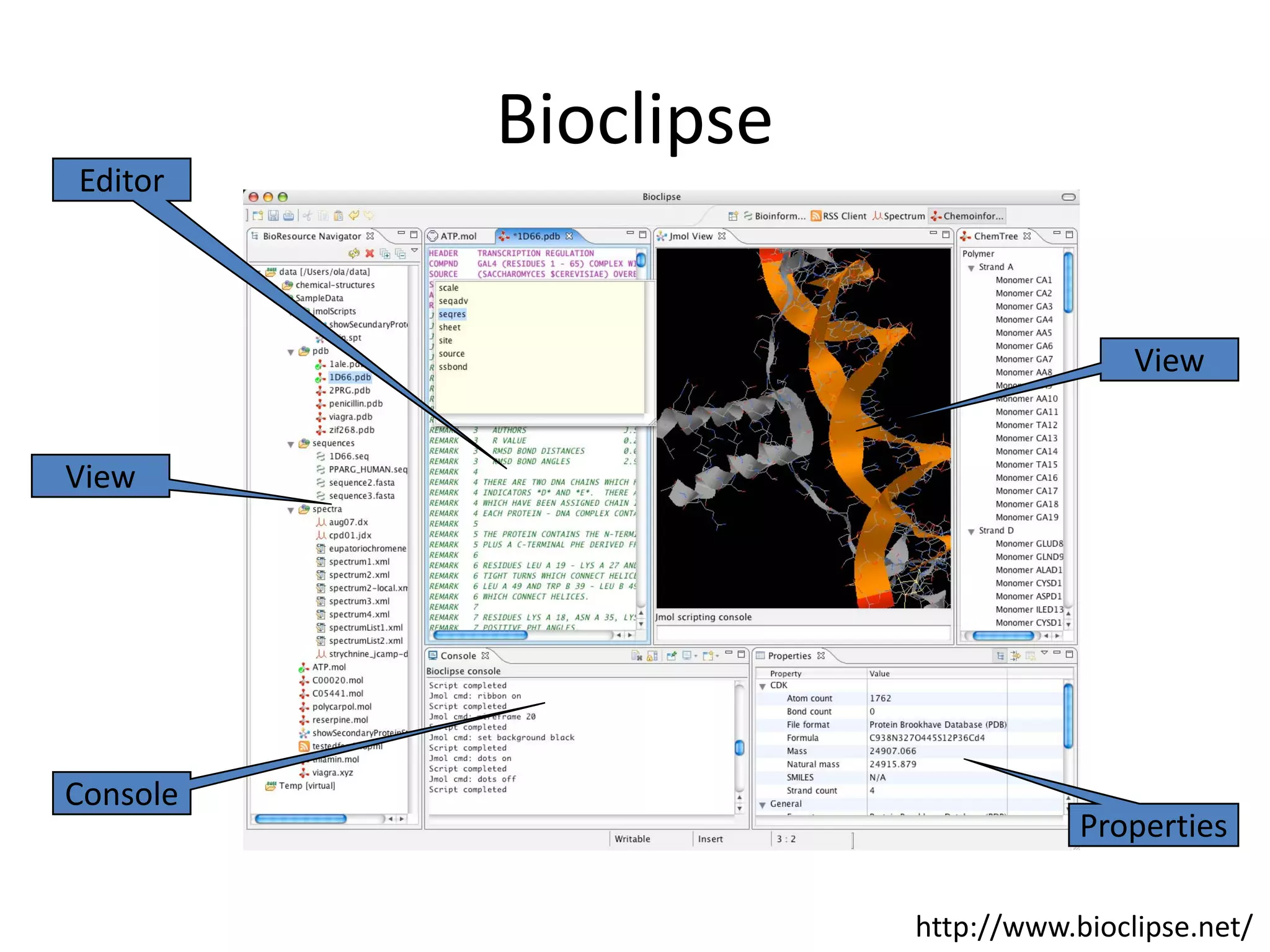
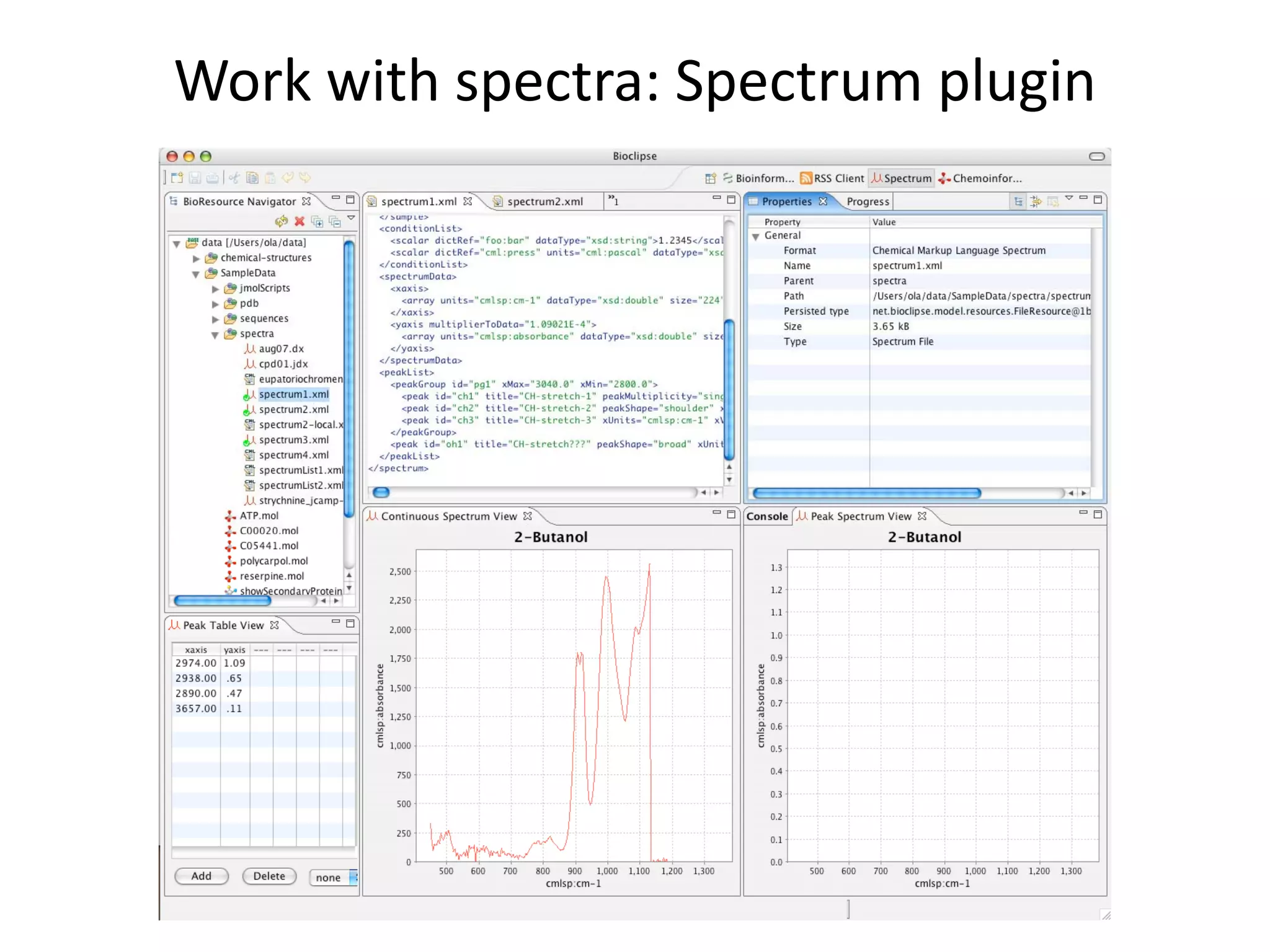
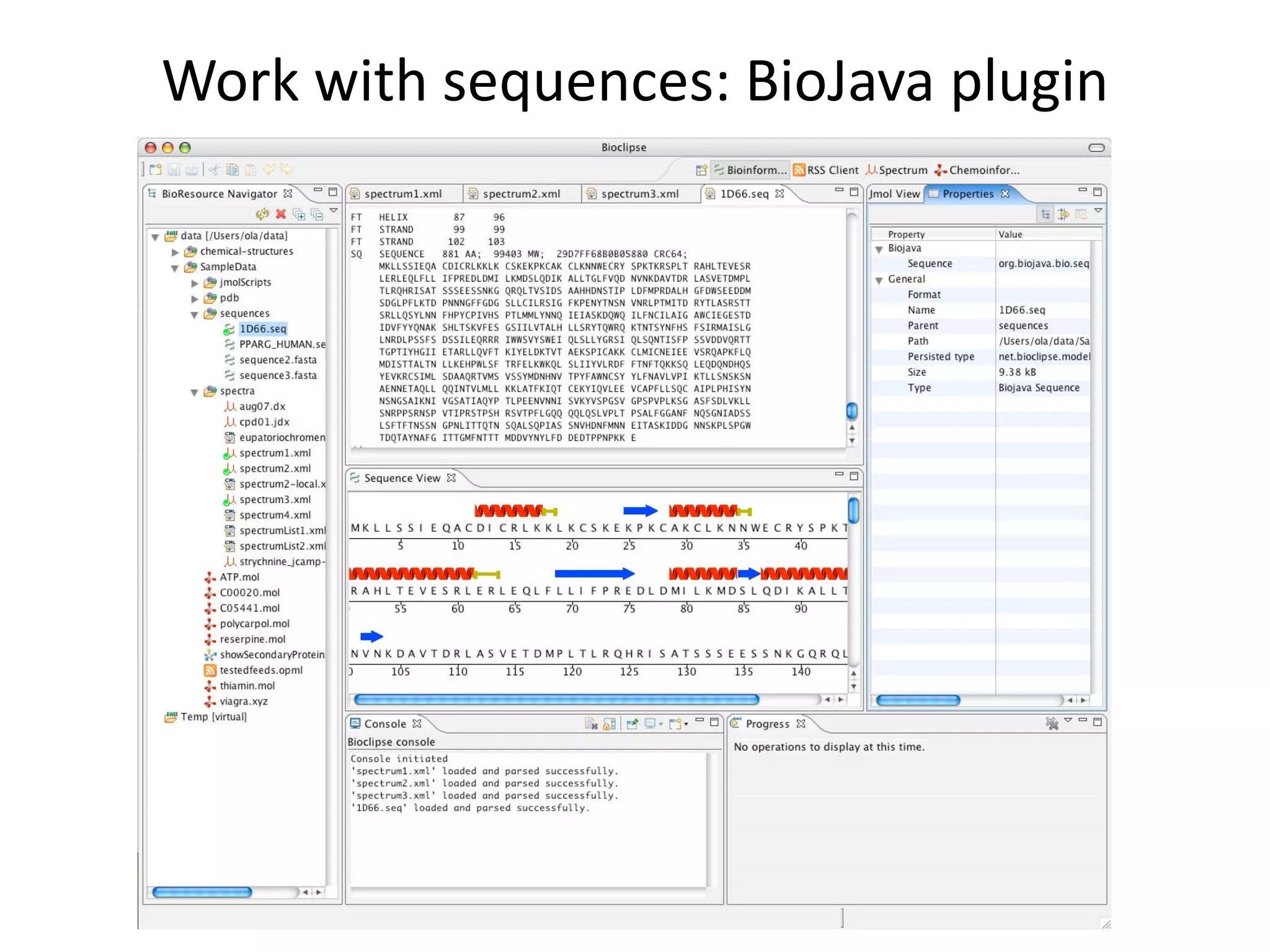
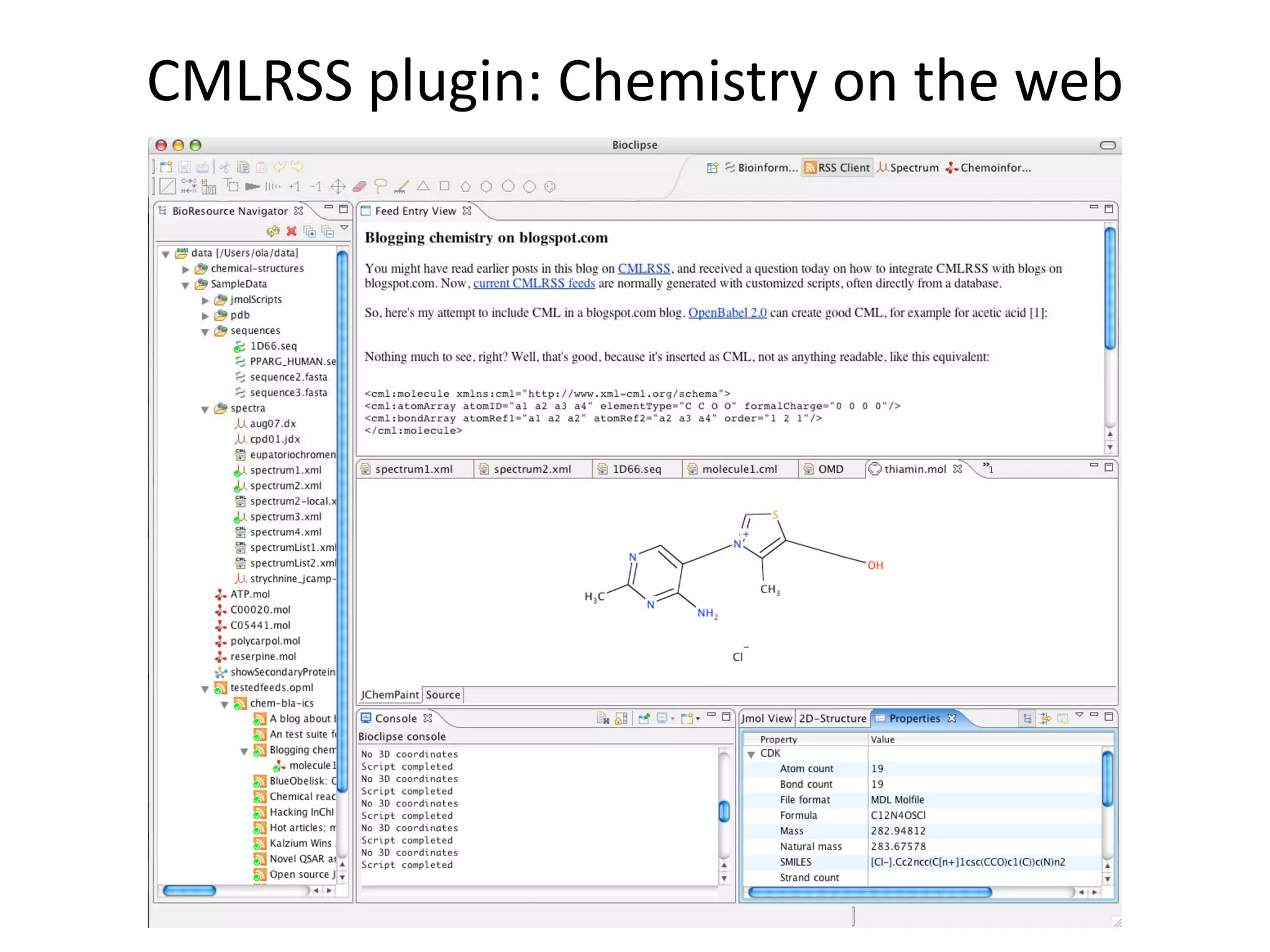
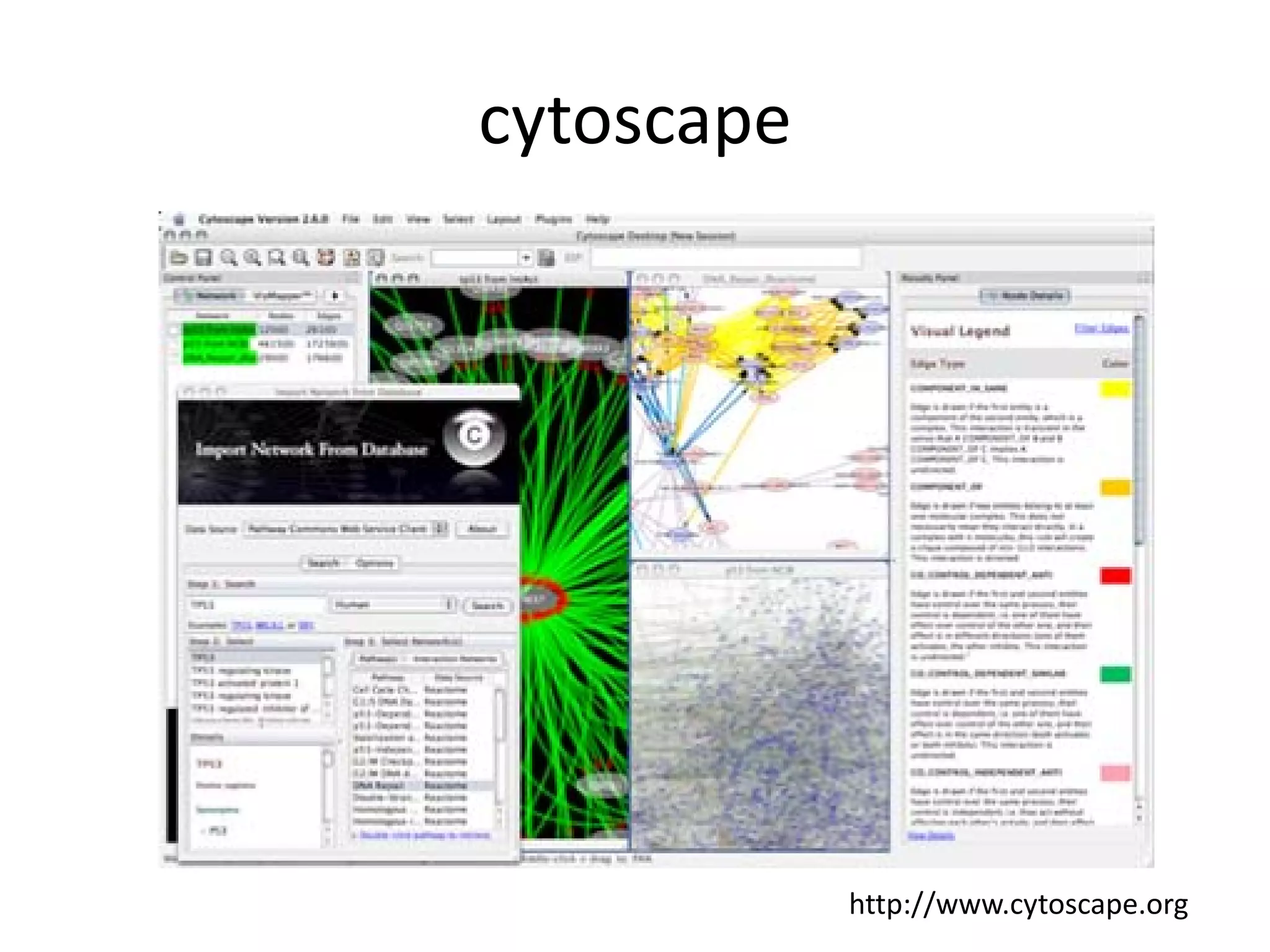
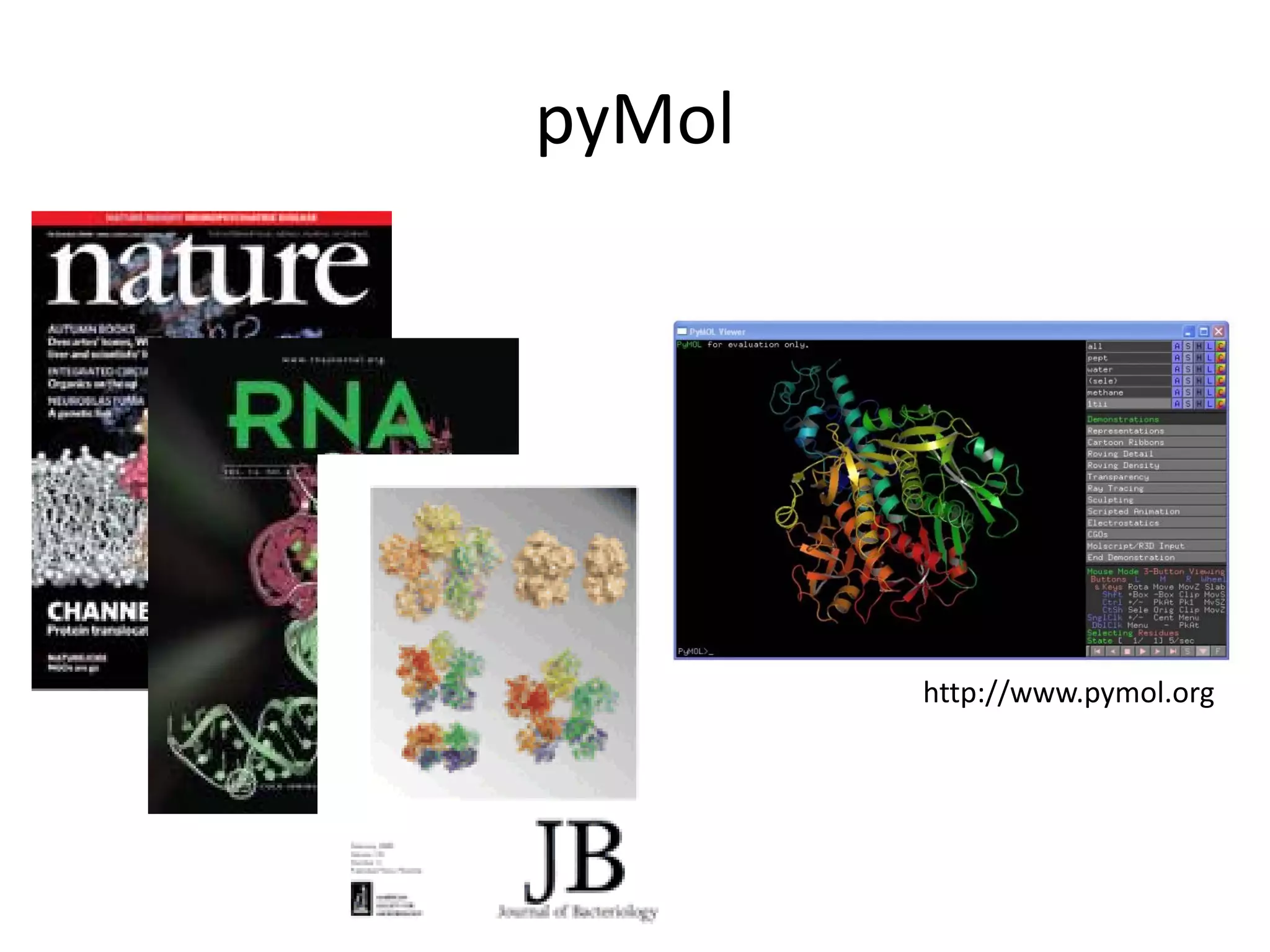


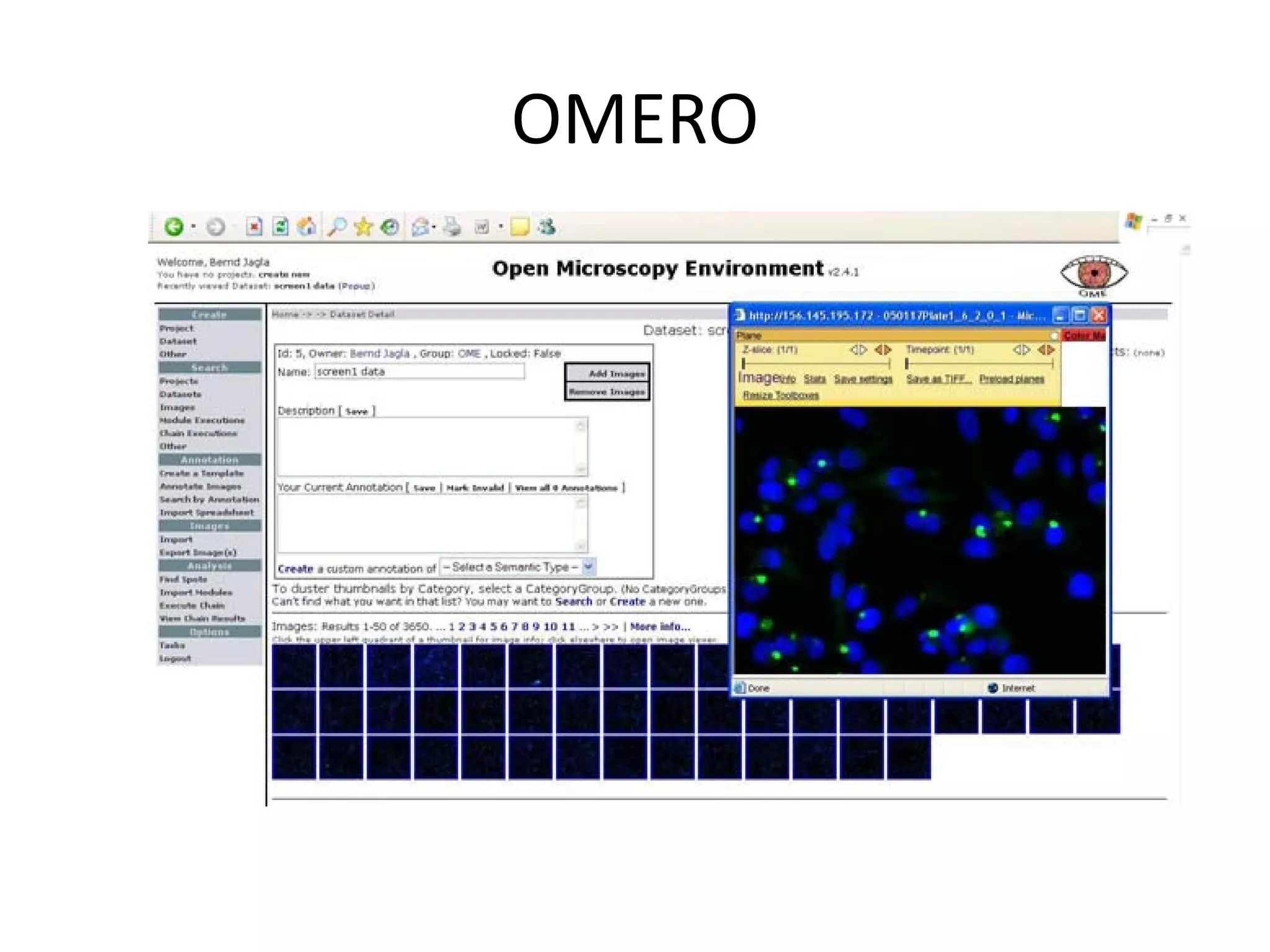
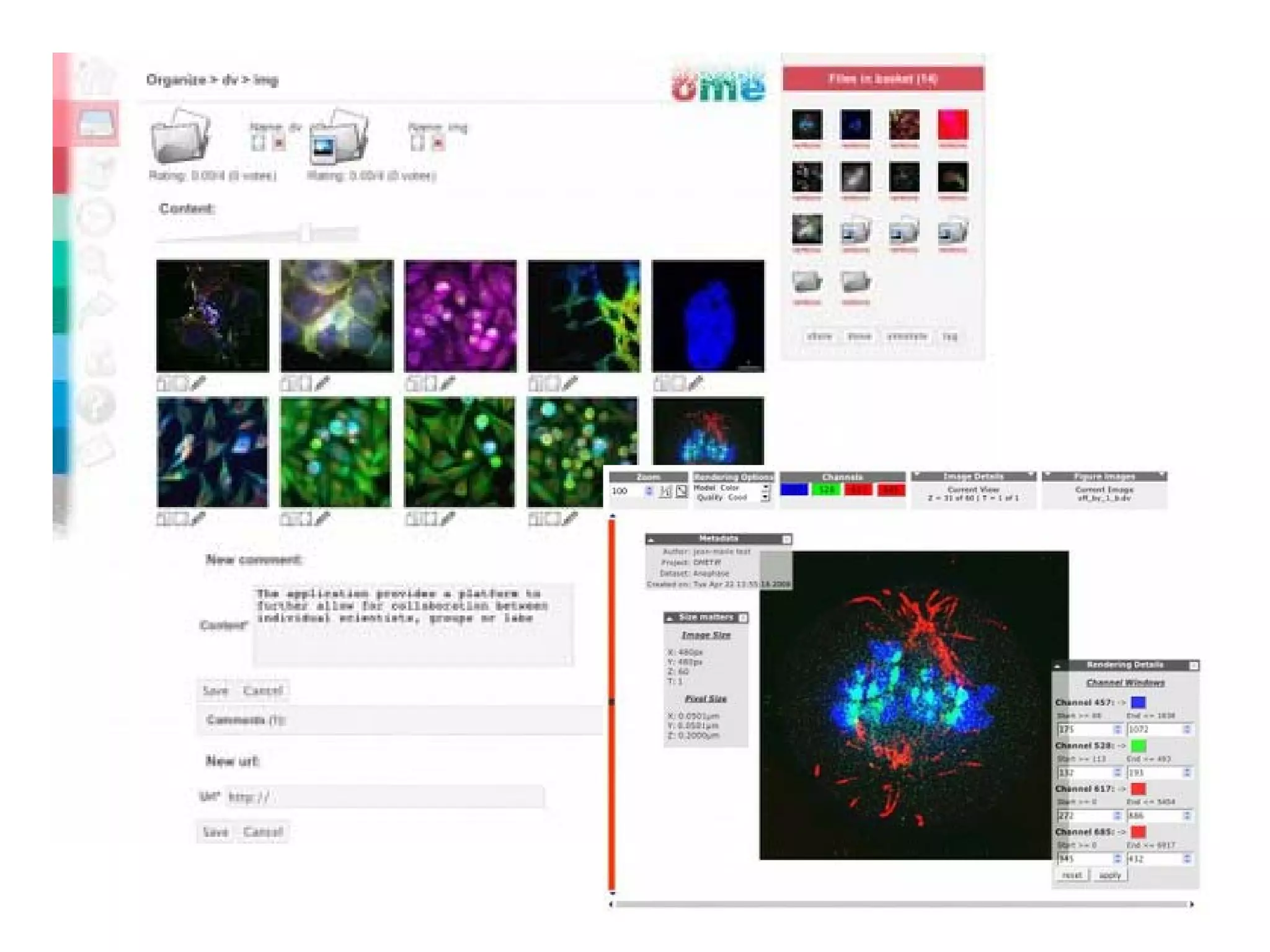
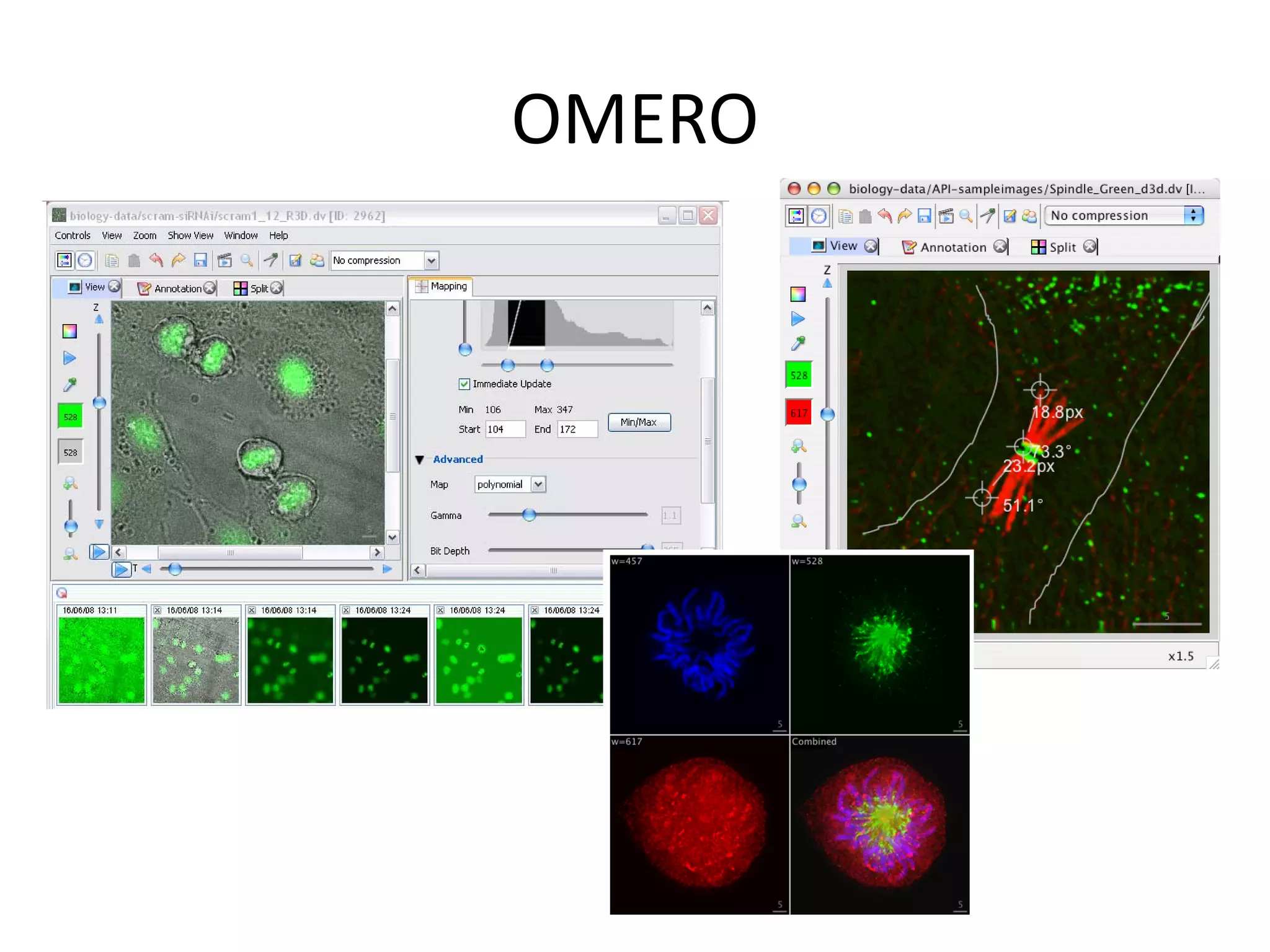










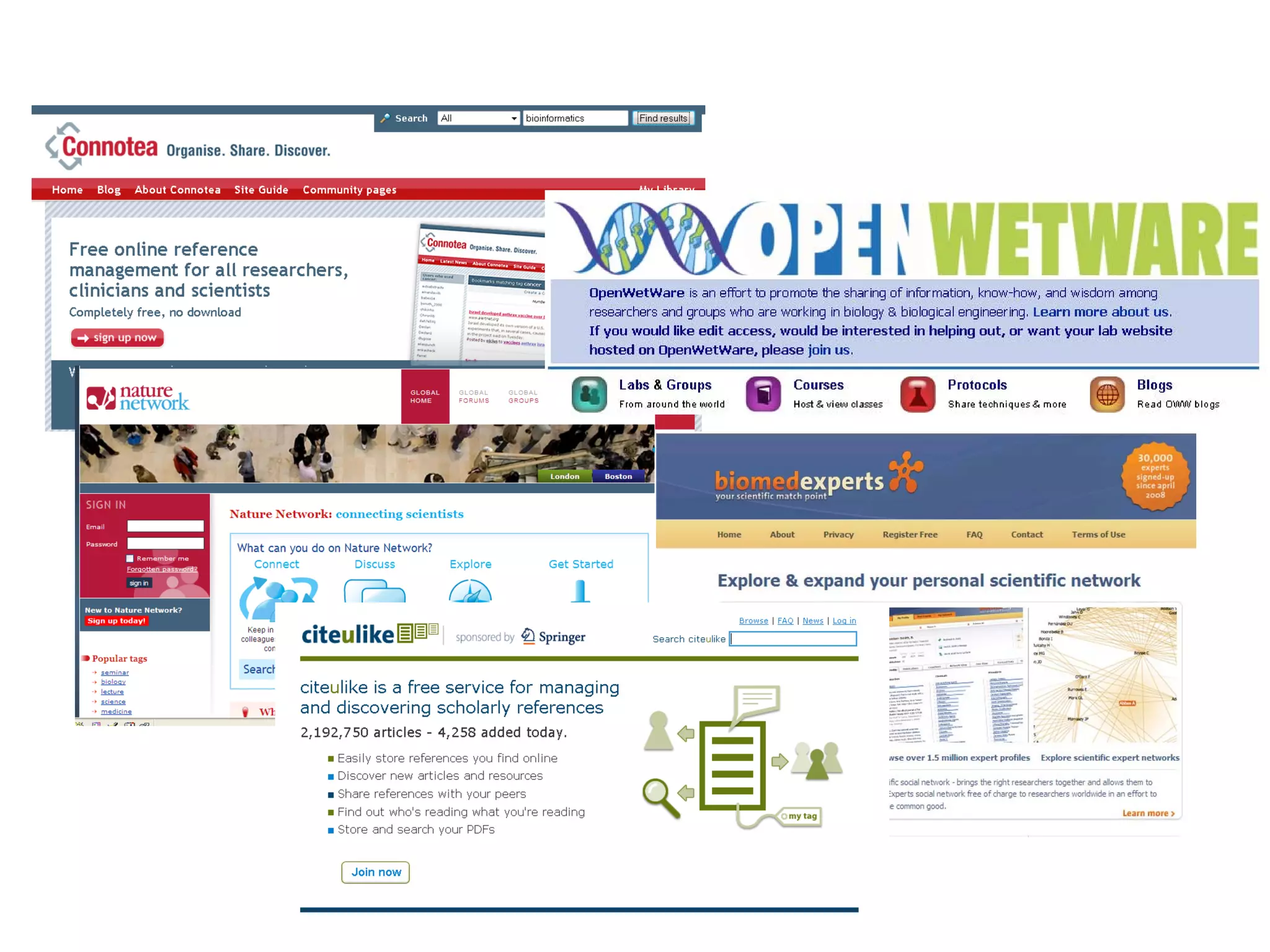



This document discusses the importance of open source software and open data in biomedical research. It notes that biological data is growing exponentially and highlights several open source bioinformatics tools like EMBOSS and web services provided by the EBI that enable researchers to access and analyze data. The document advocates for open standards to facilitate data integration and management across different omics domains.




































































































