Cool Ahsg Model
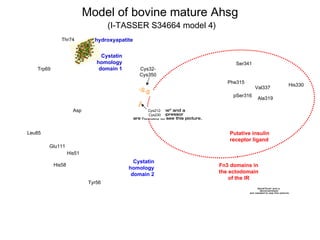
Fig. 14. A cartoon representation of the two structural faces of secreted Ahsg. There is no X-ray crystallographic or NMR-derived structure on which to base the 3-dimensional folding of Ahsg from any species. However, bioinformatics tools can be used to simulate a model which suggests recognition of hydroxyapatite by the C-terminal cystatin homology domains (residues 18-248; dotted red ovals), and the Fn3 homology domains of the insulin receptor by Ahsg residues 249-367 (dotted blue oval). The 350-residue single chain of human Ahsg (residues 18-367) was submitted to the I-TASSER website (Zhang, 2008), which uses amino acid homology of submitted protein sequences with protein sequences in the PDB database, and returns five models. I stipulated that I-TASSER take into consideration the known 5 disulfide pairs in Ahsg using the “Assign contact/distance restraints” feature, and chose Model 5 from Run S34302. PyMol for Mac (http://www.pymol.org/) to visualize the resulting pdb-formatted file generated by I-TASSER. Some of the –S-S- bonds are depicted in orange. Regions of strong α-helicity are colored blue; β-sheet structures yellow, and the unstructured backbone colored green.
Recommended
Recommended
More Related Content
Viewers also liked
Viewers also liked (12)
Cool Ahsg Model
- 1. Model of bovine mature Ahsg (I-TASSER S34664 model 4) Phe315 Ala319 Val337 Ser341 Cys32-Cys350 Thr74 His330 His58 Trp69 Tyr56 Leu85 Asp 115 Glu111 Putative insulin receptor ligand Cystatin homology domain 1 Cystatin homology domain 2 His51 Fn3 domains in the ectodomain of the IR hydroxyapatite pSer316 Cys212-Cys230 -S-S
- 2. Model of human mature Ahsg (I-TASSER S34302) http://zhang.bioinformatics.ku.edu/I-TASSER/ Thr158 Cystatin homology domain 2 Thr37 Ile48 Gln174 Phe229 Lys237 Cys358 Cys32 -S-S- Gln308 Arg311 Gly240 Thr248 Cys96 His90 Pro254 Cys132 Cystatin homology domain 1 Cys142-Cys145 Gly348 Val347 Lys338 Arg337 Putative insulin receptor ligand Hydroxyapatite crystal Fibronectin-III domains In the insulin receptor