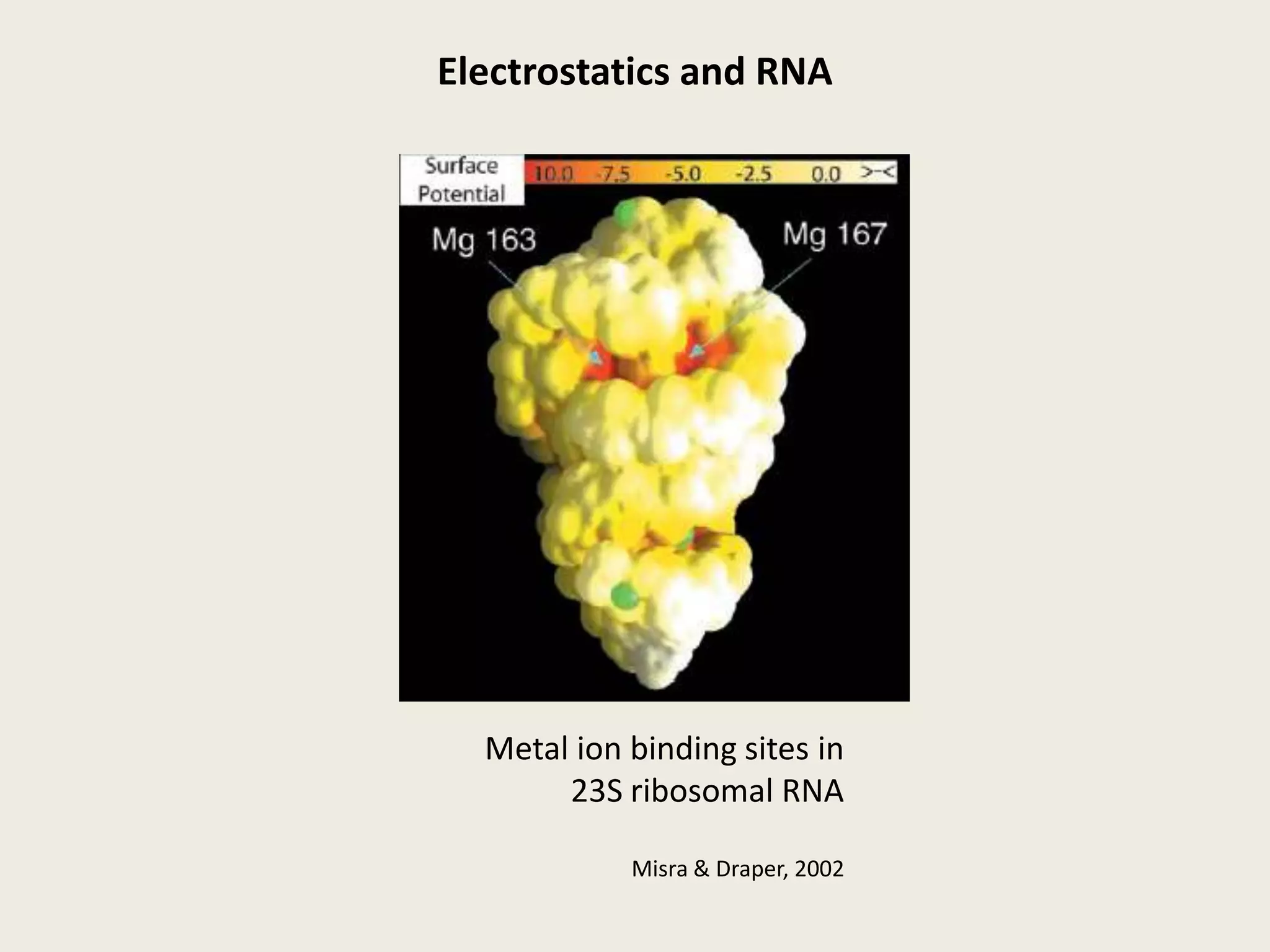
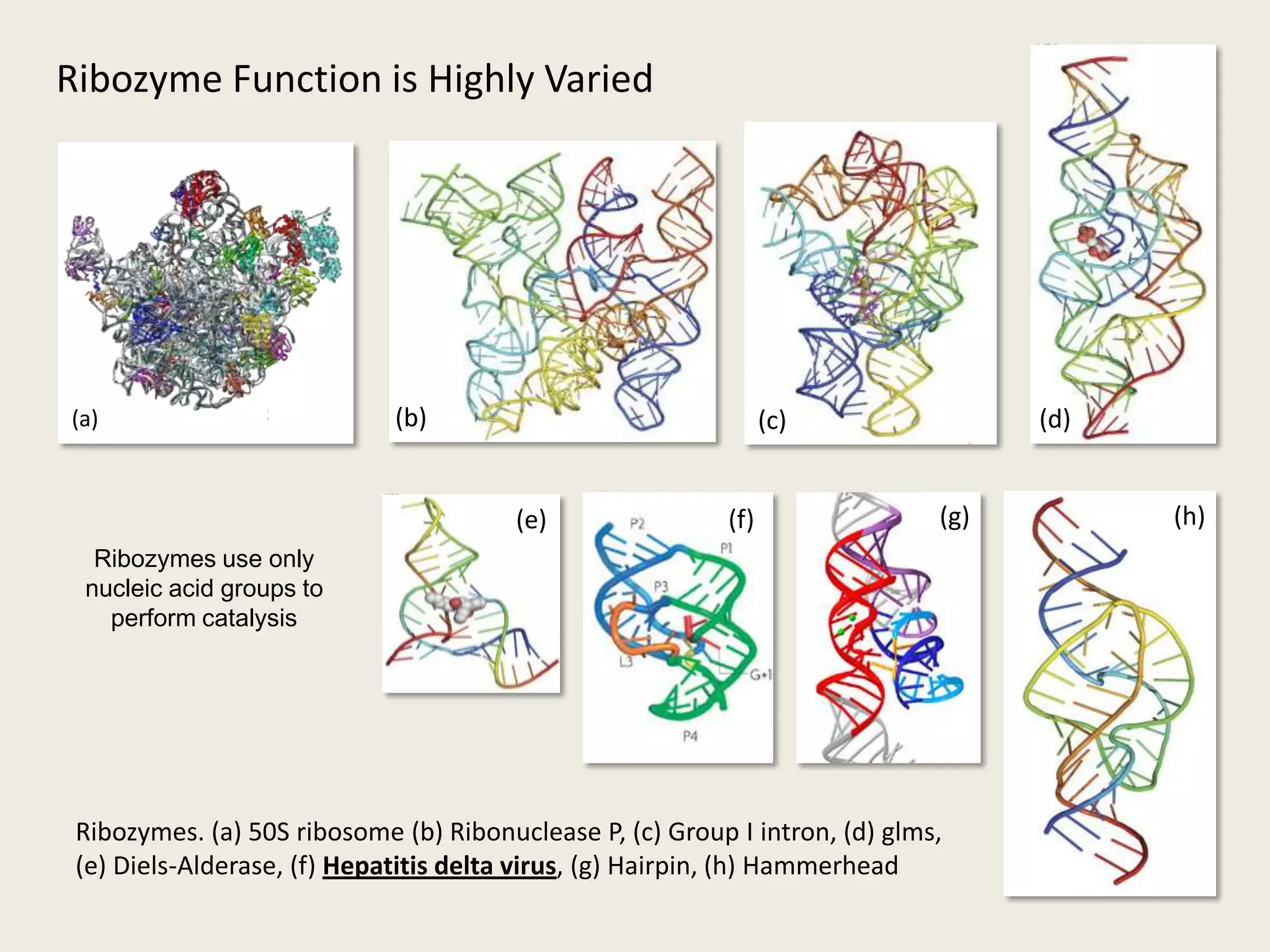
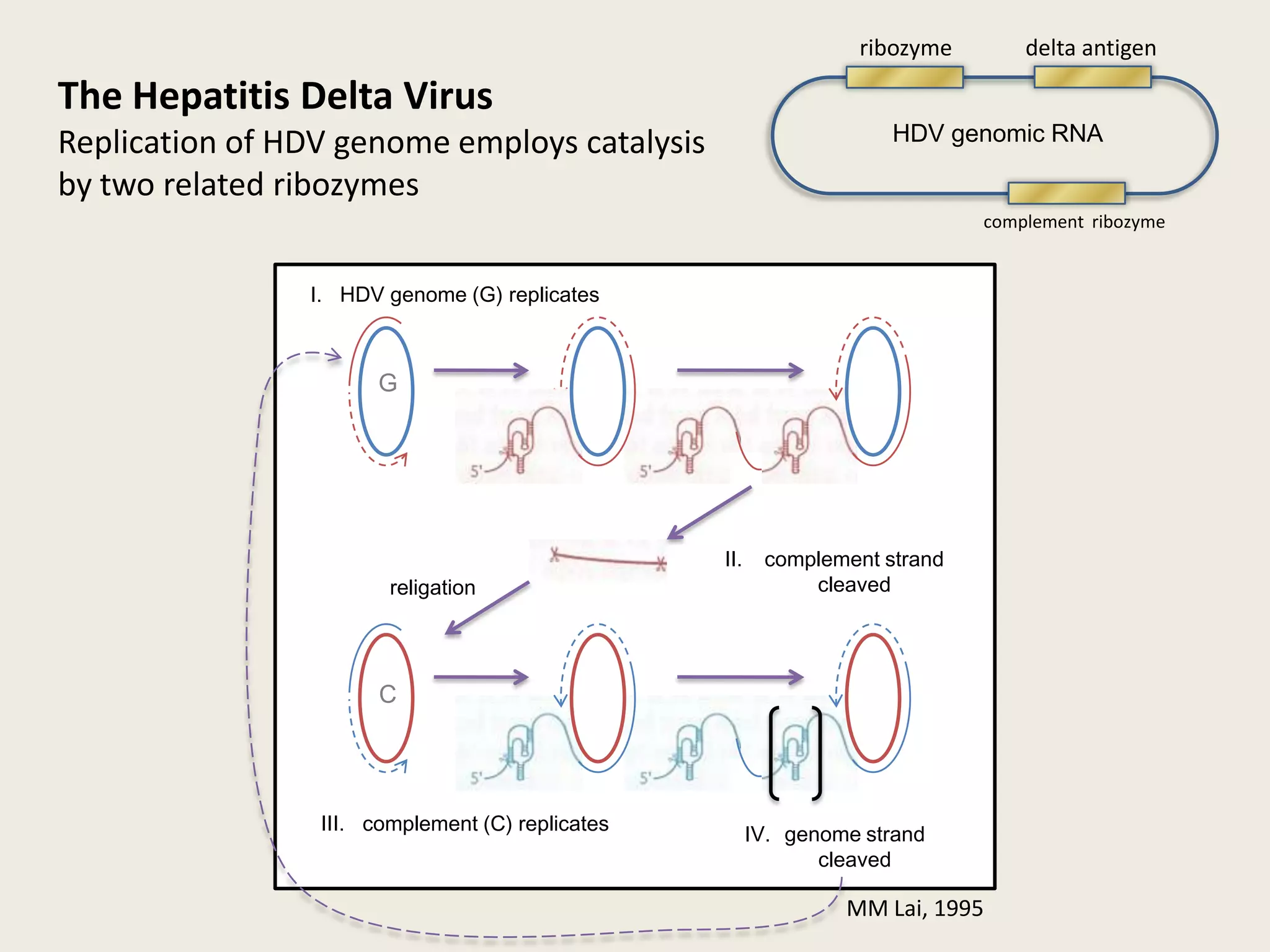
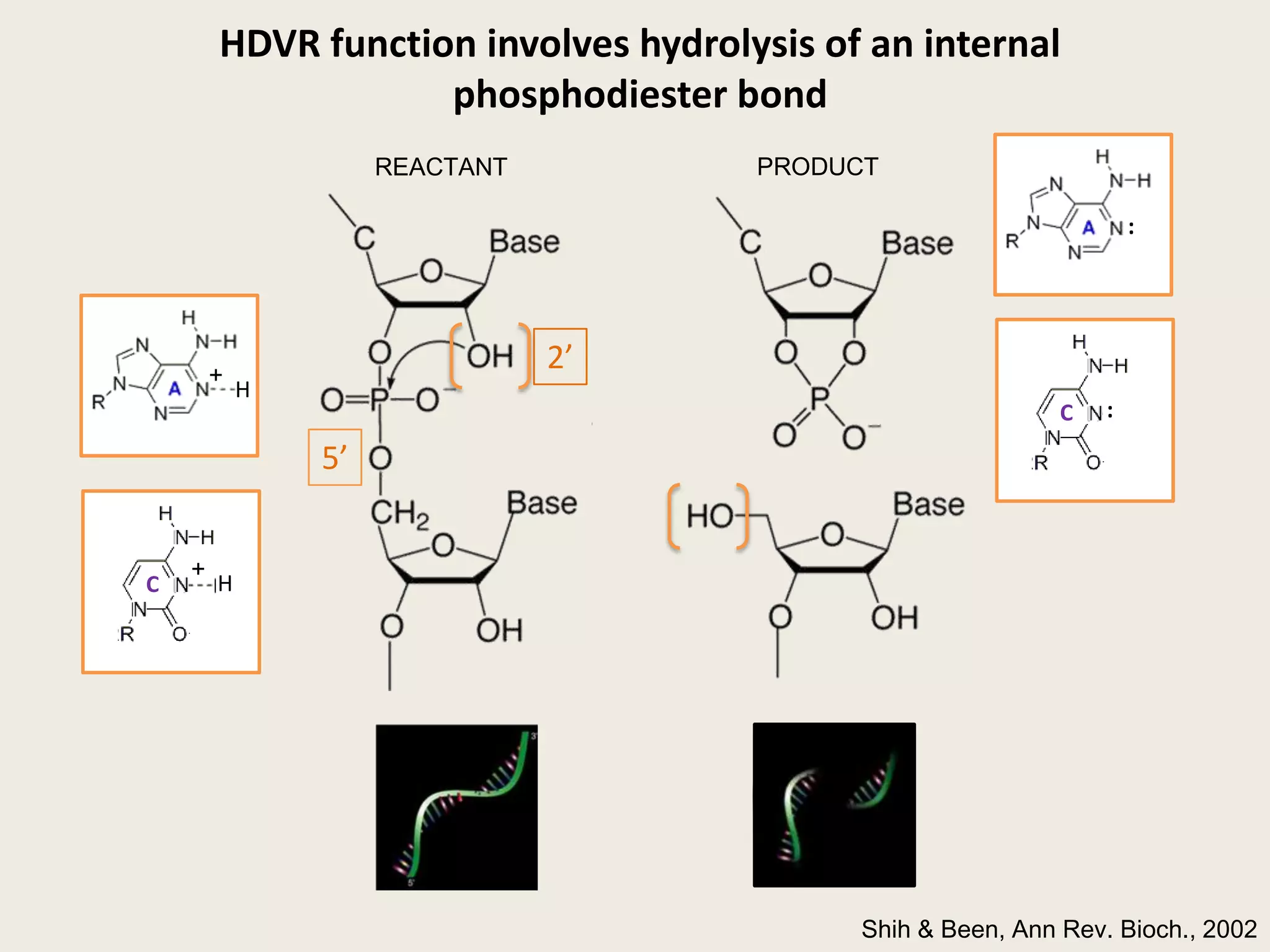
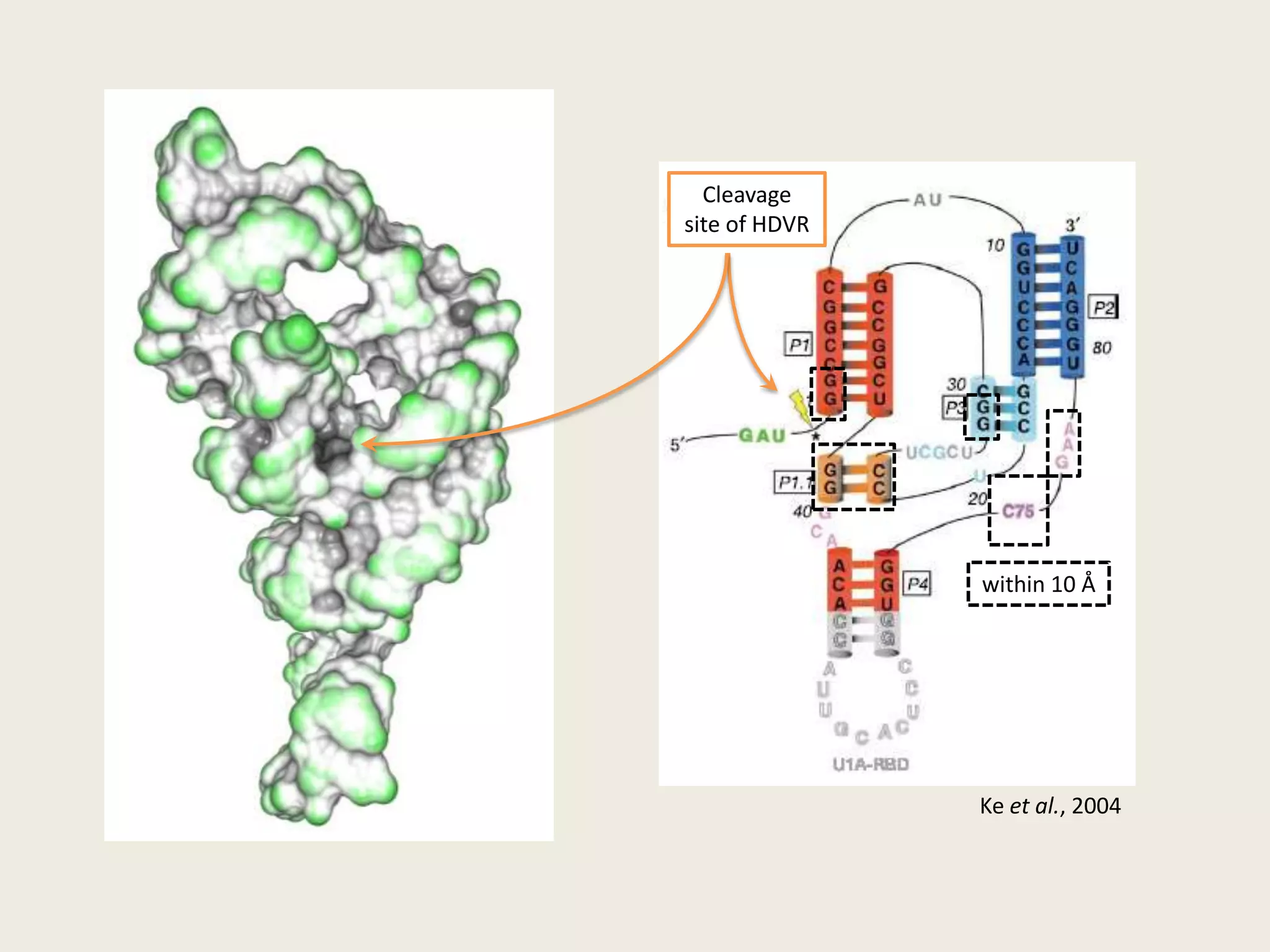
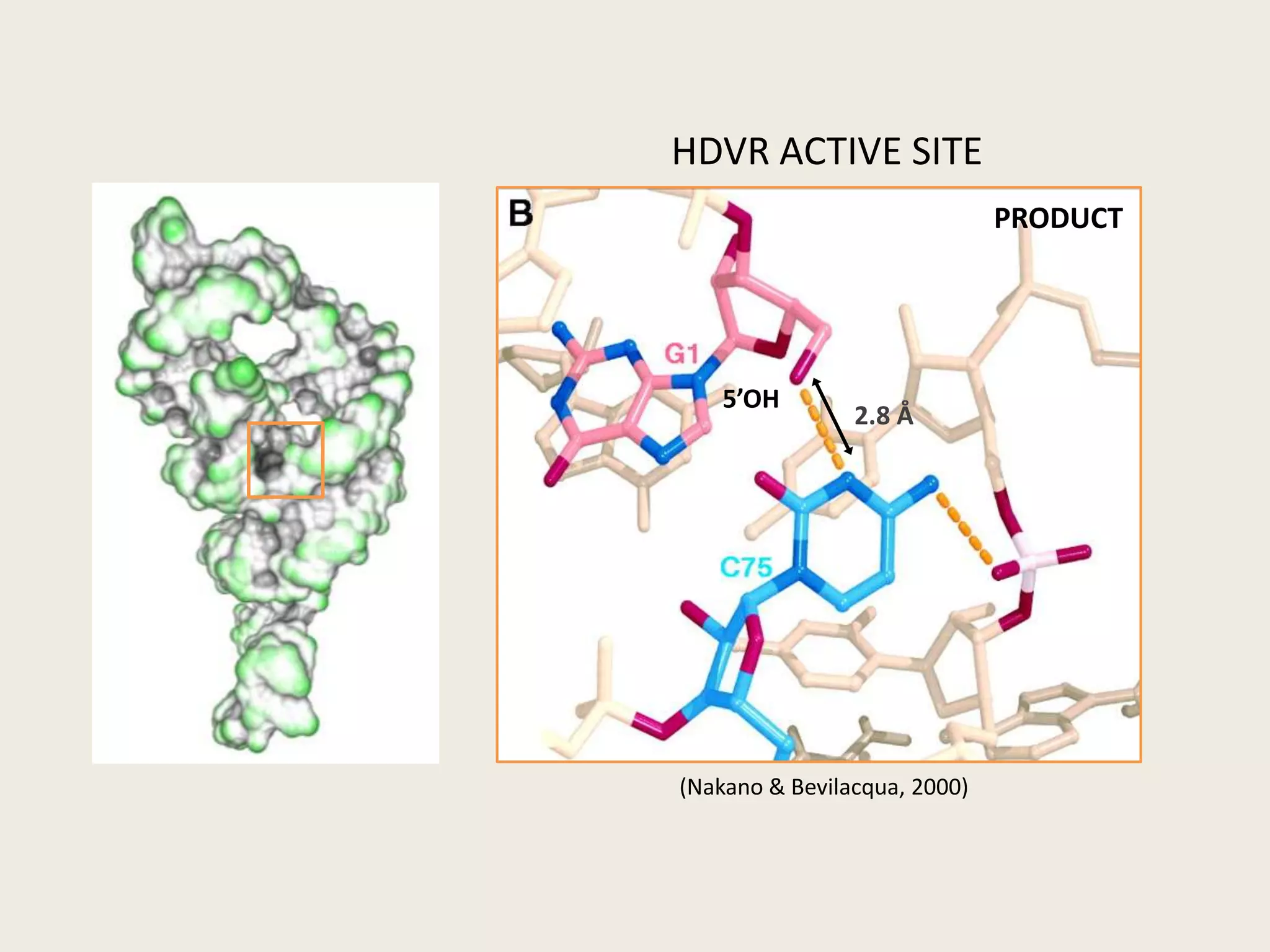
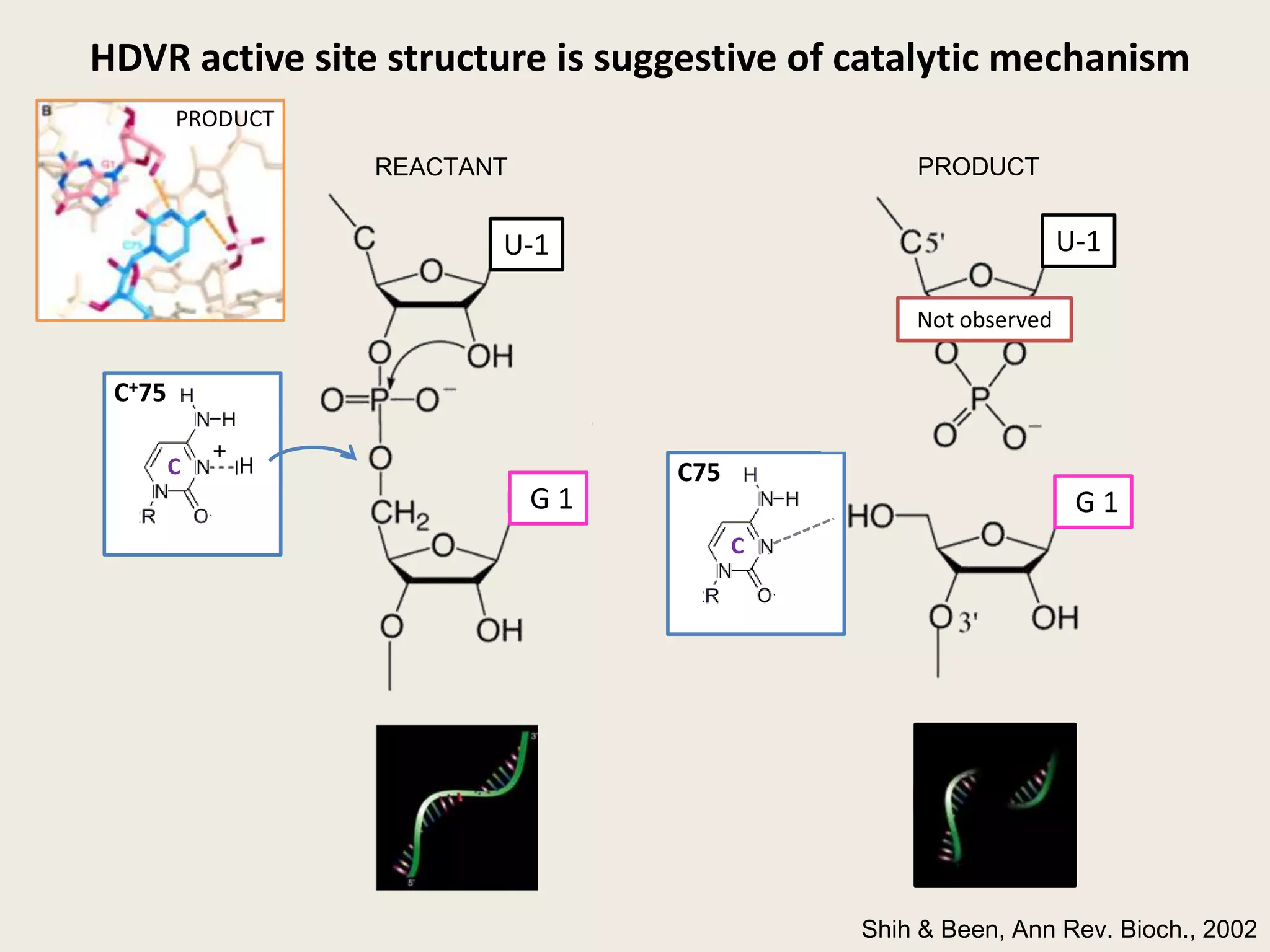
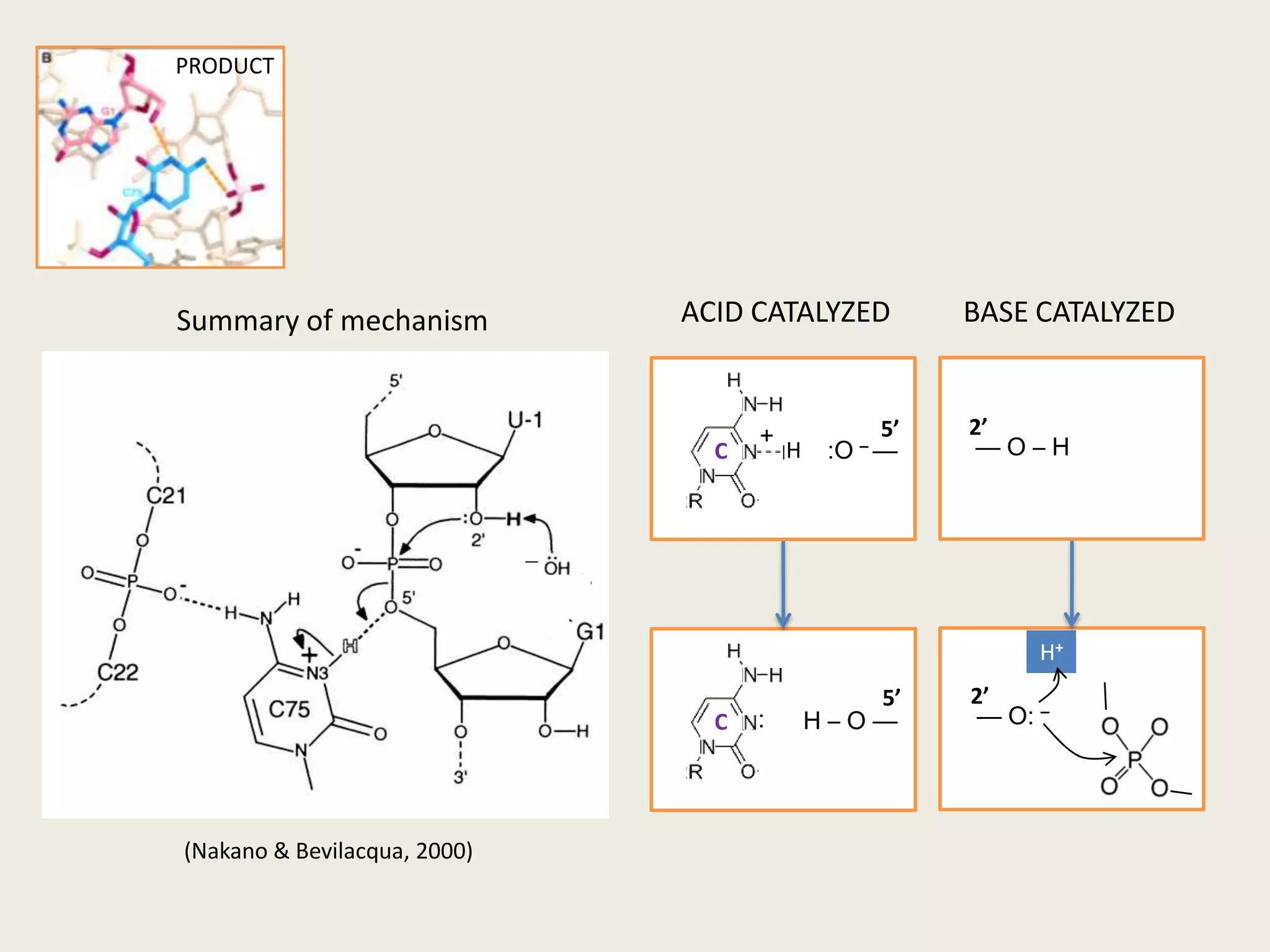
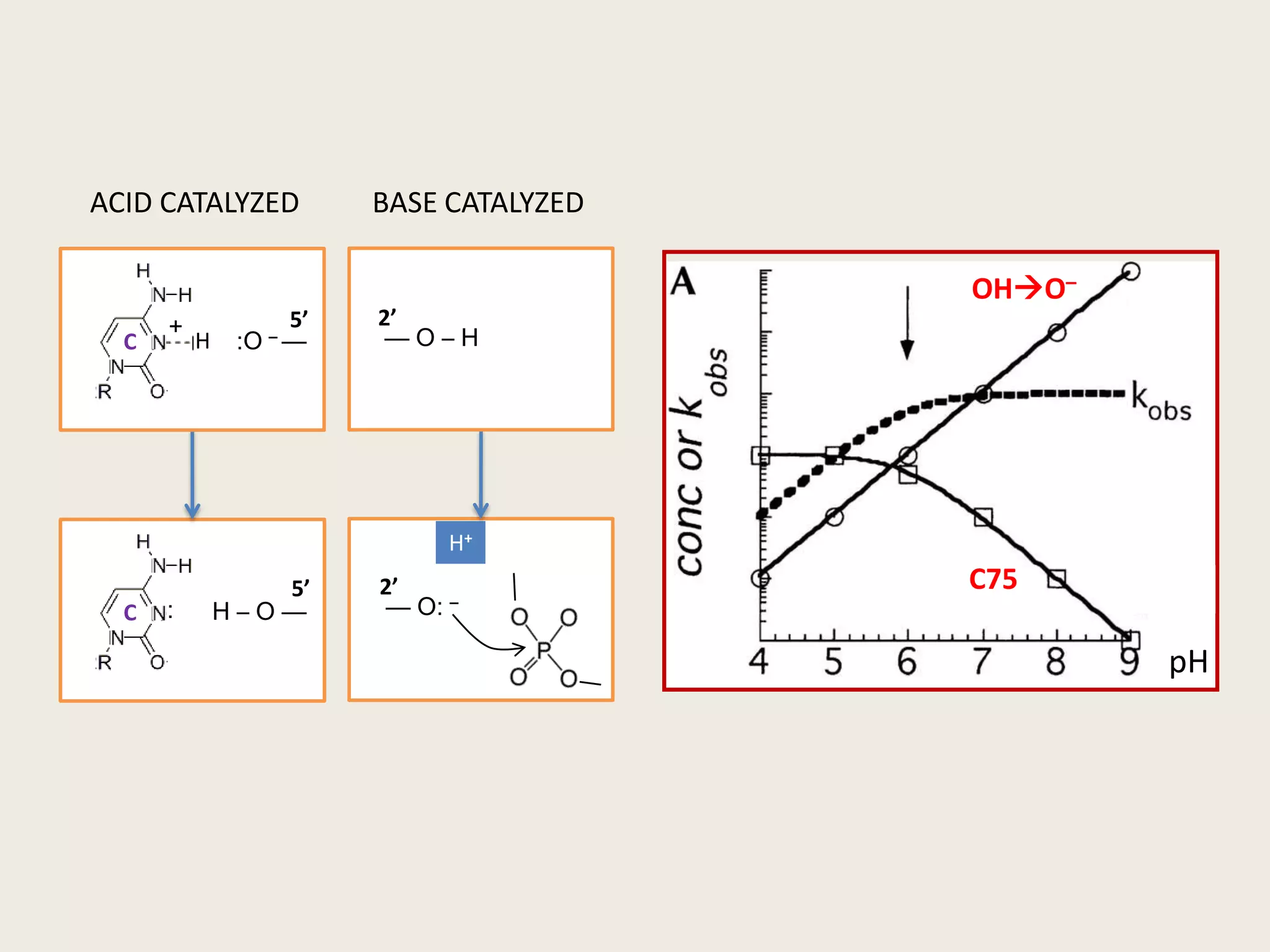
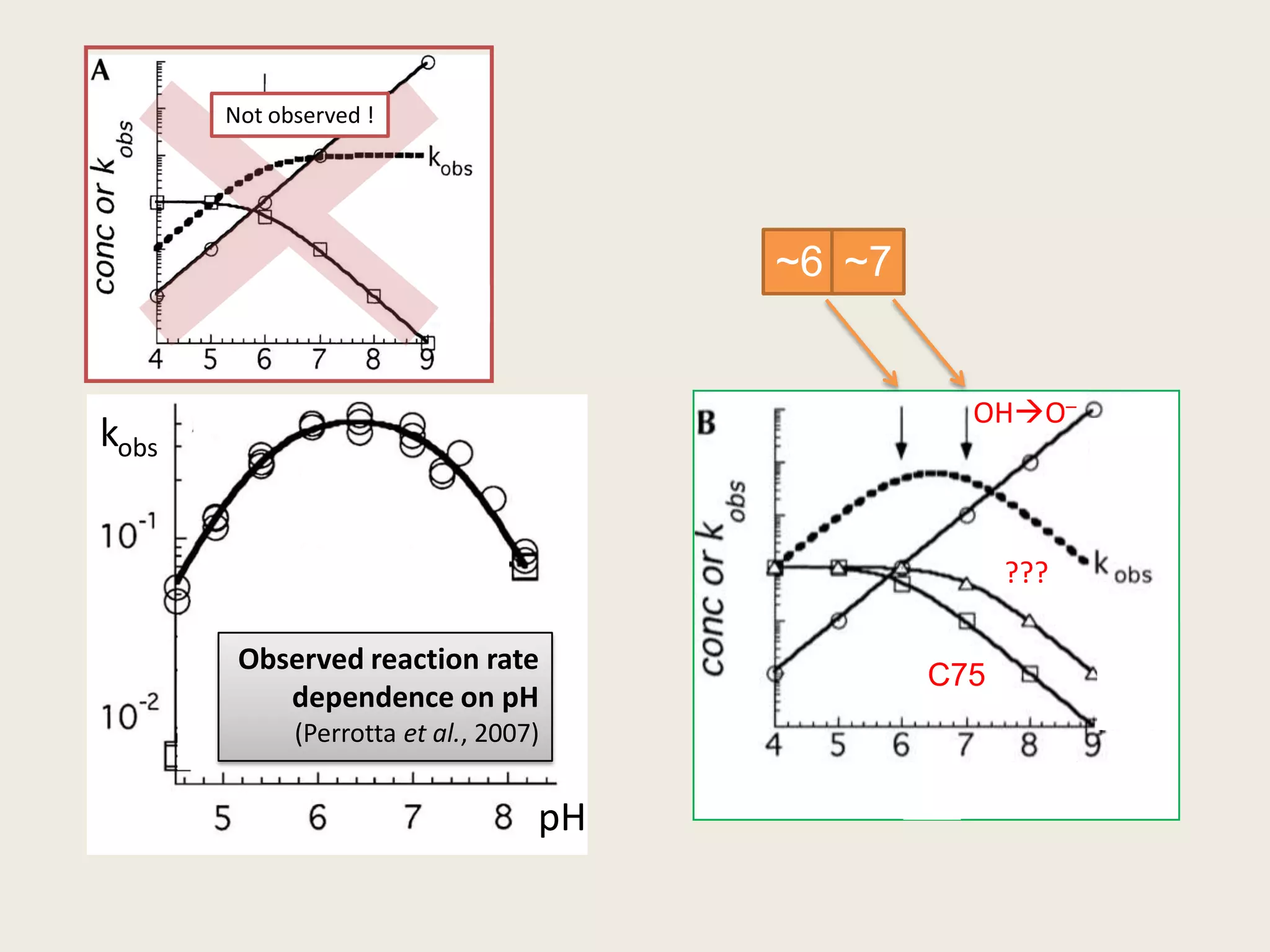
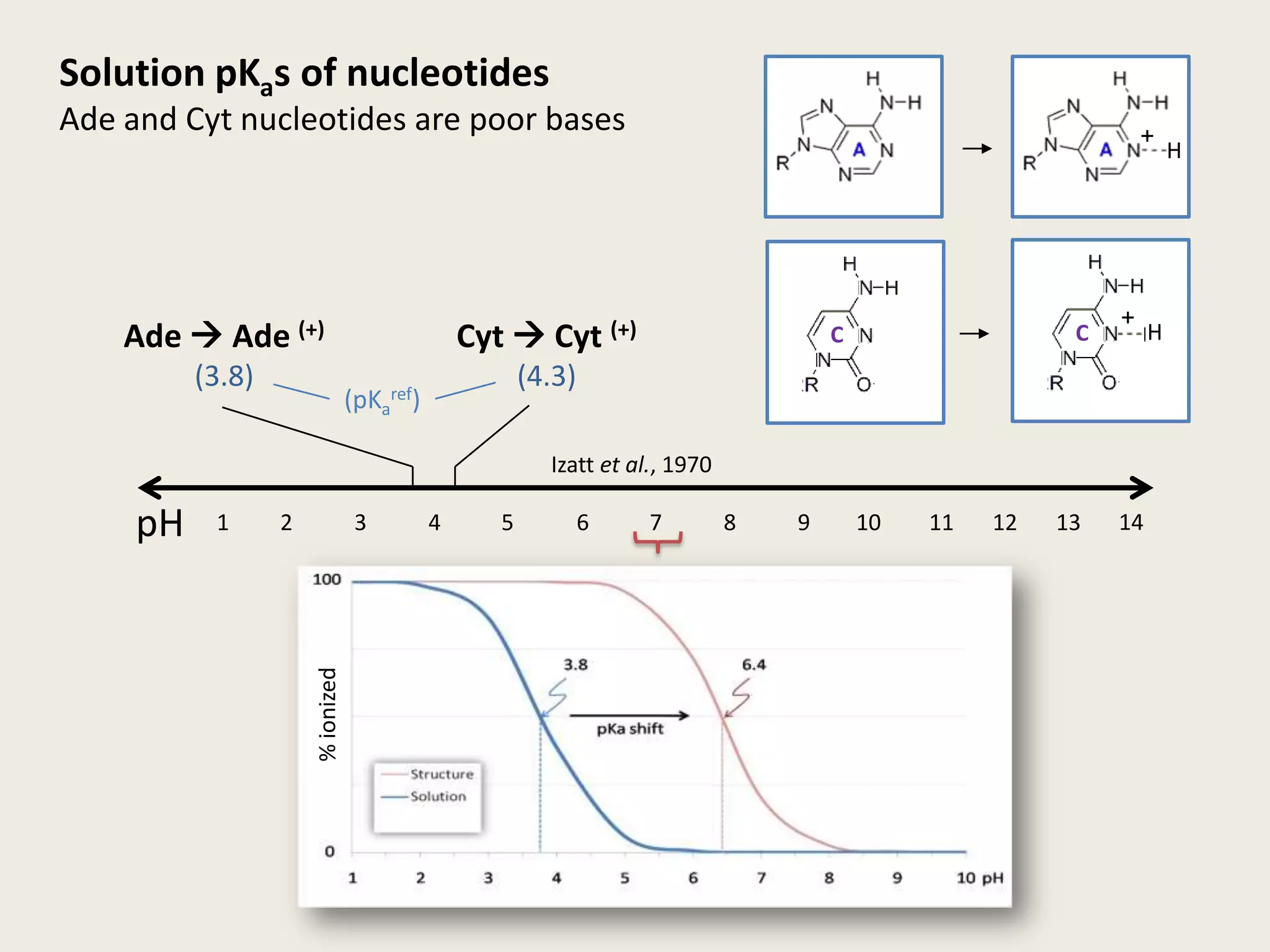
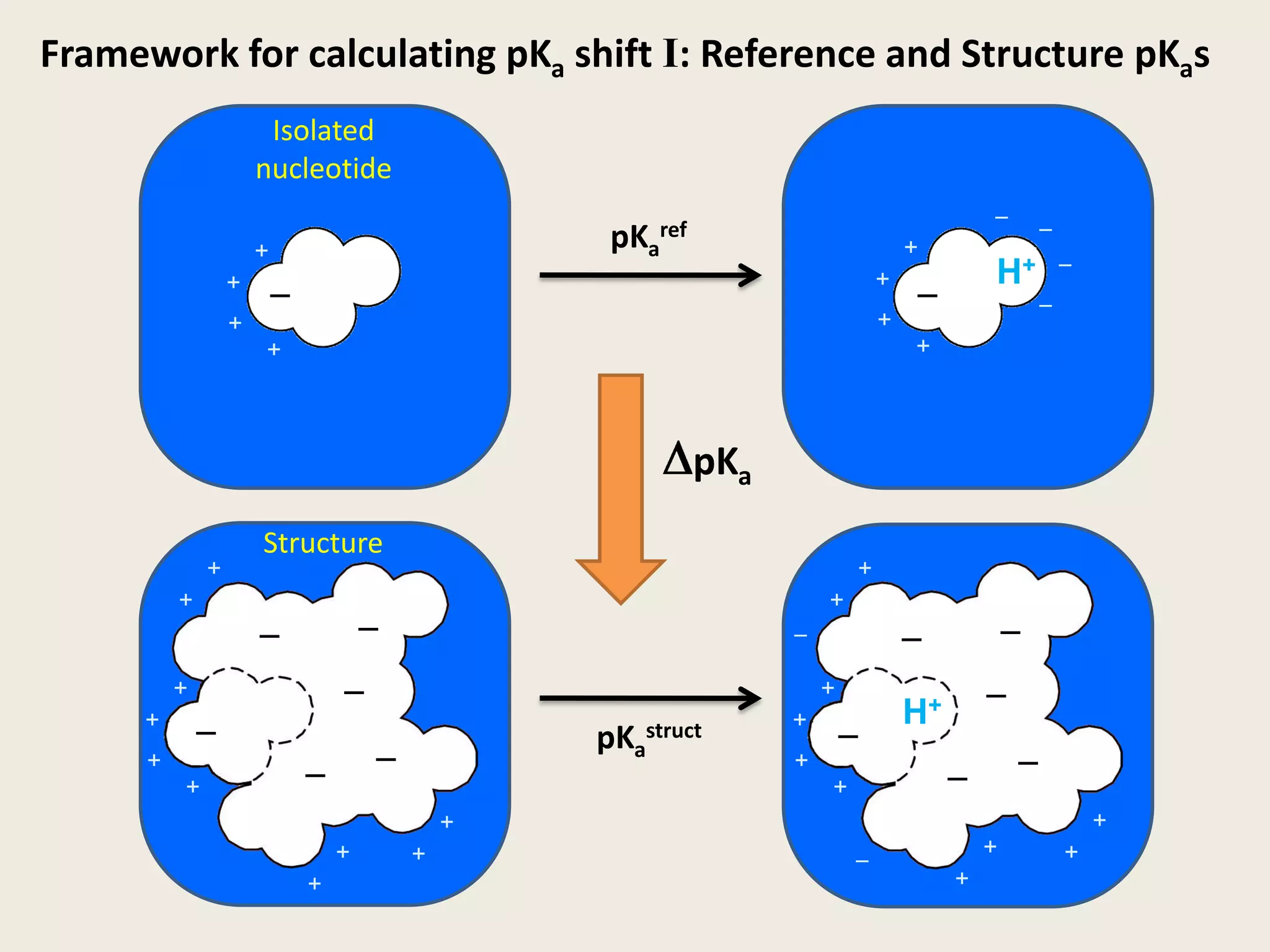
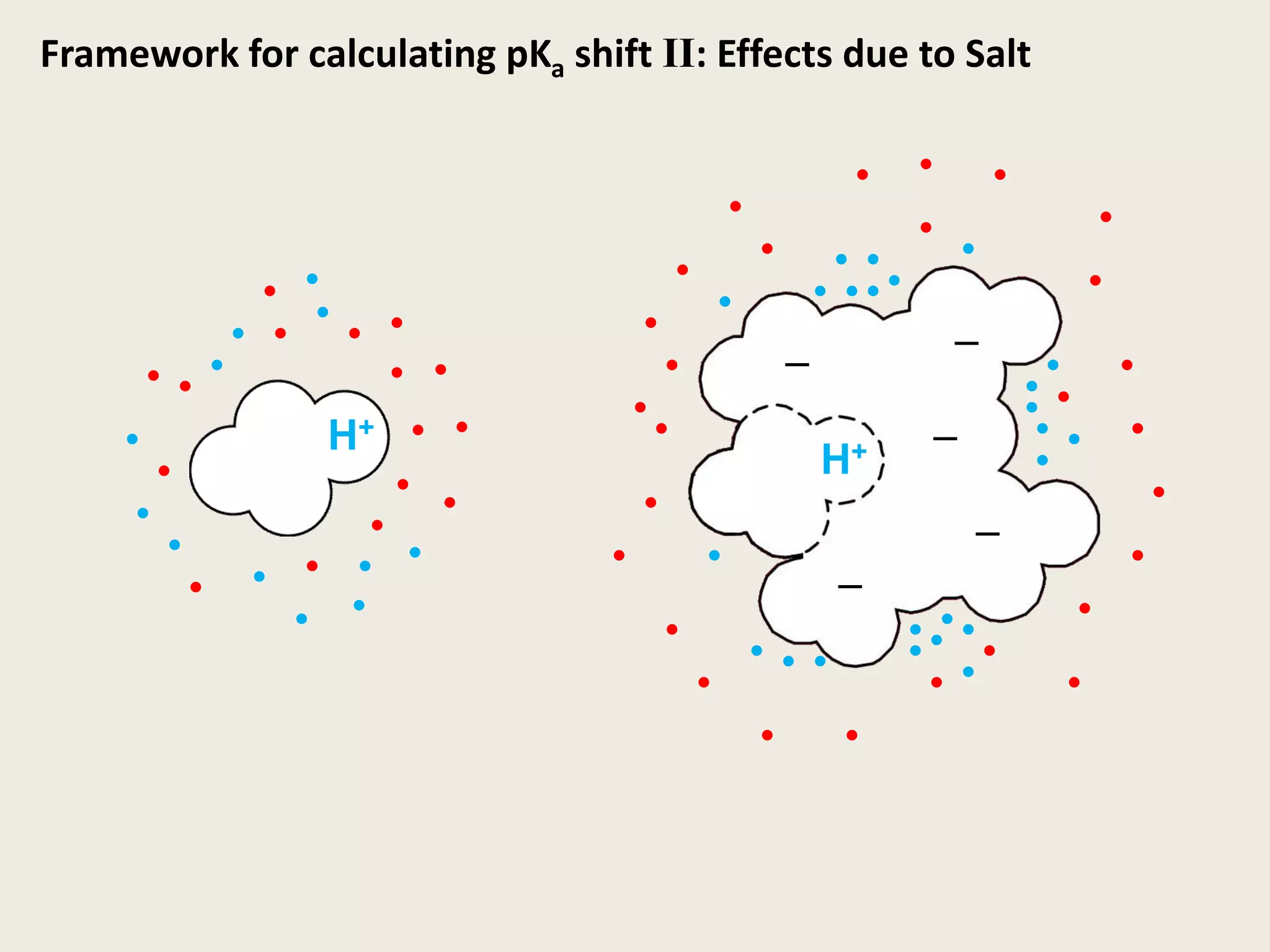
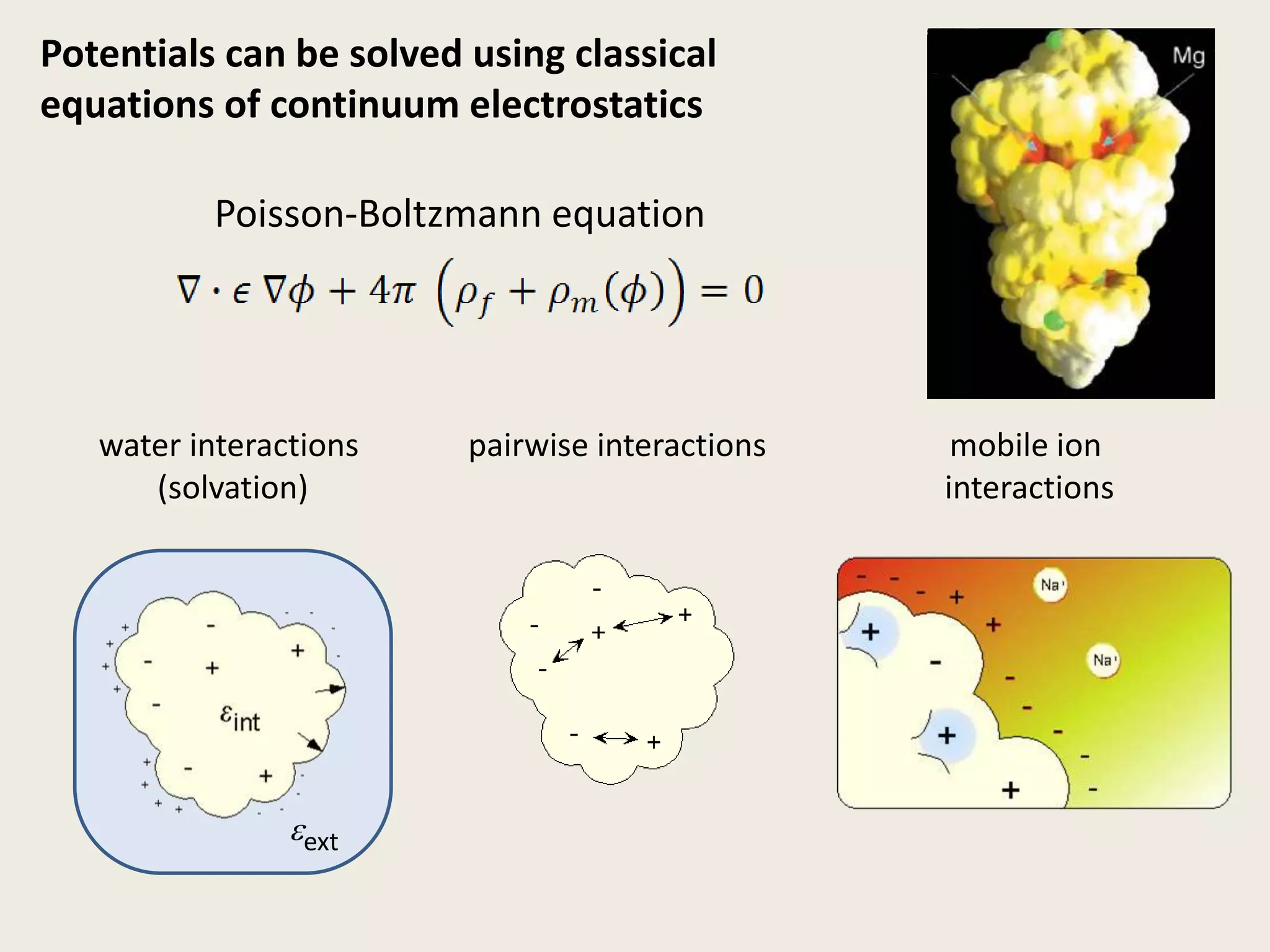
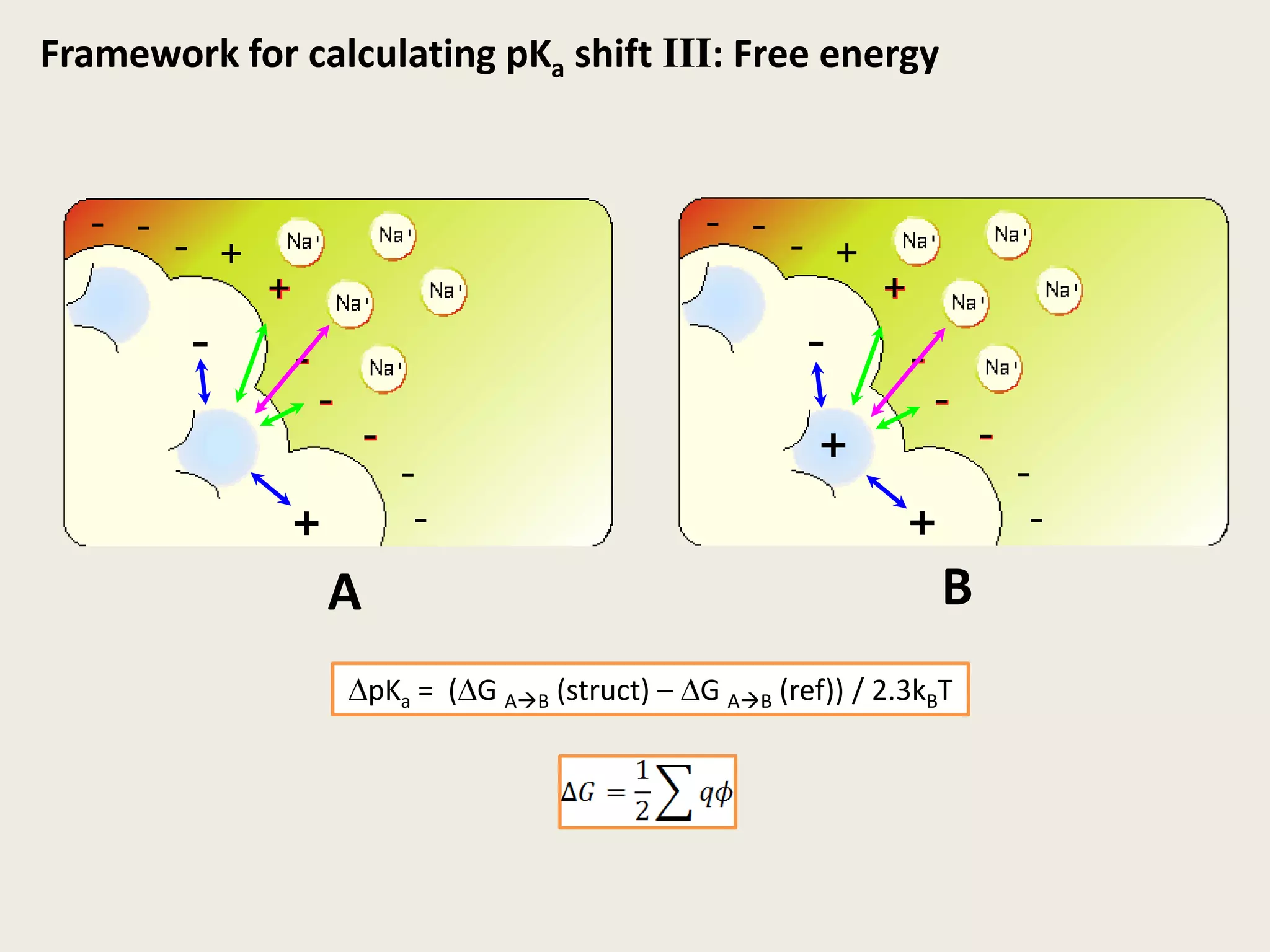
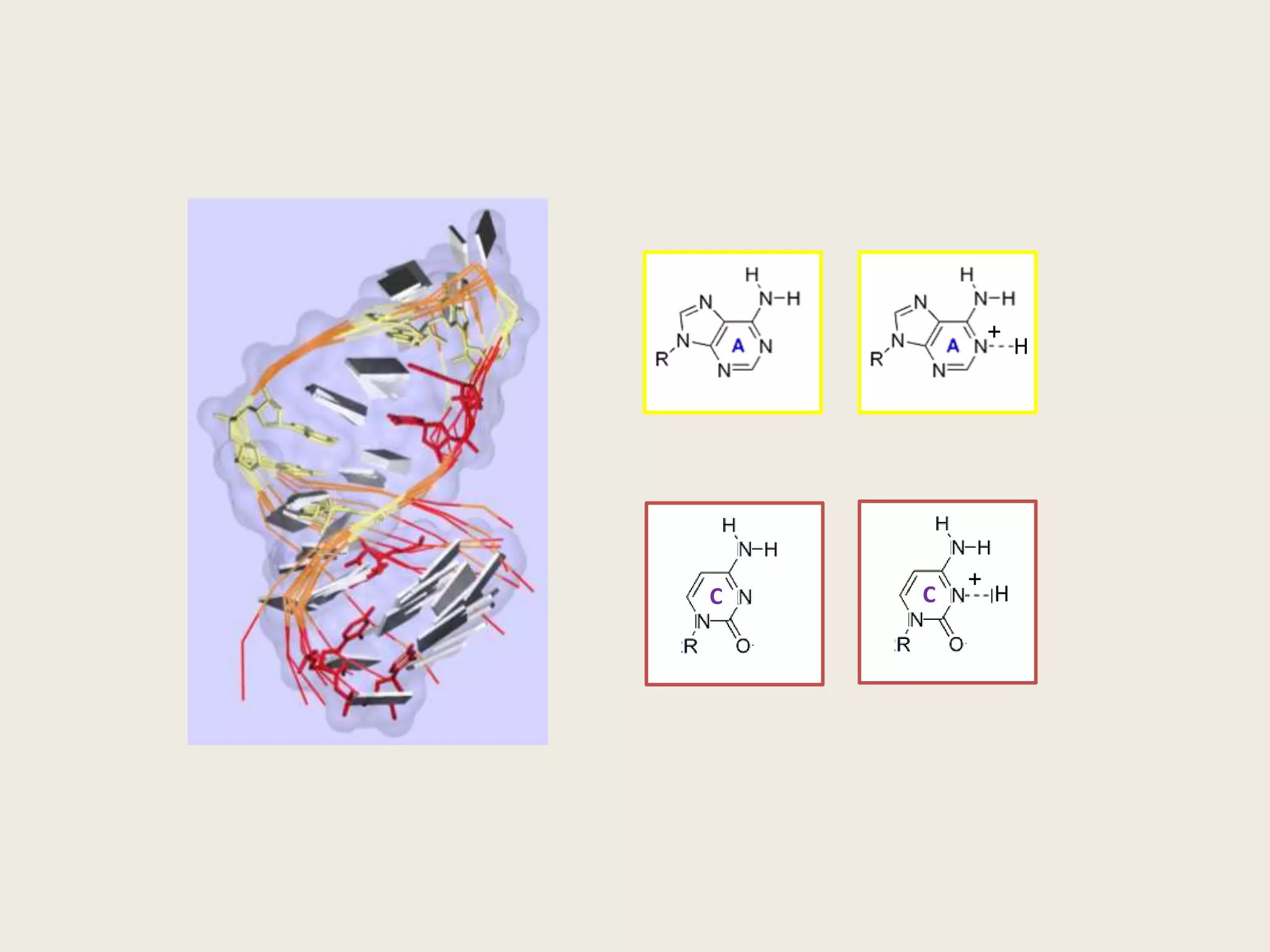
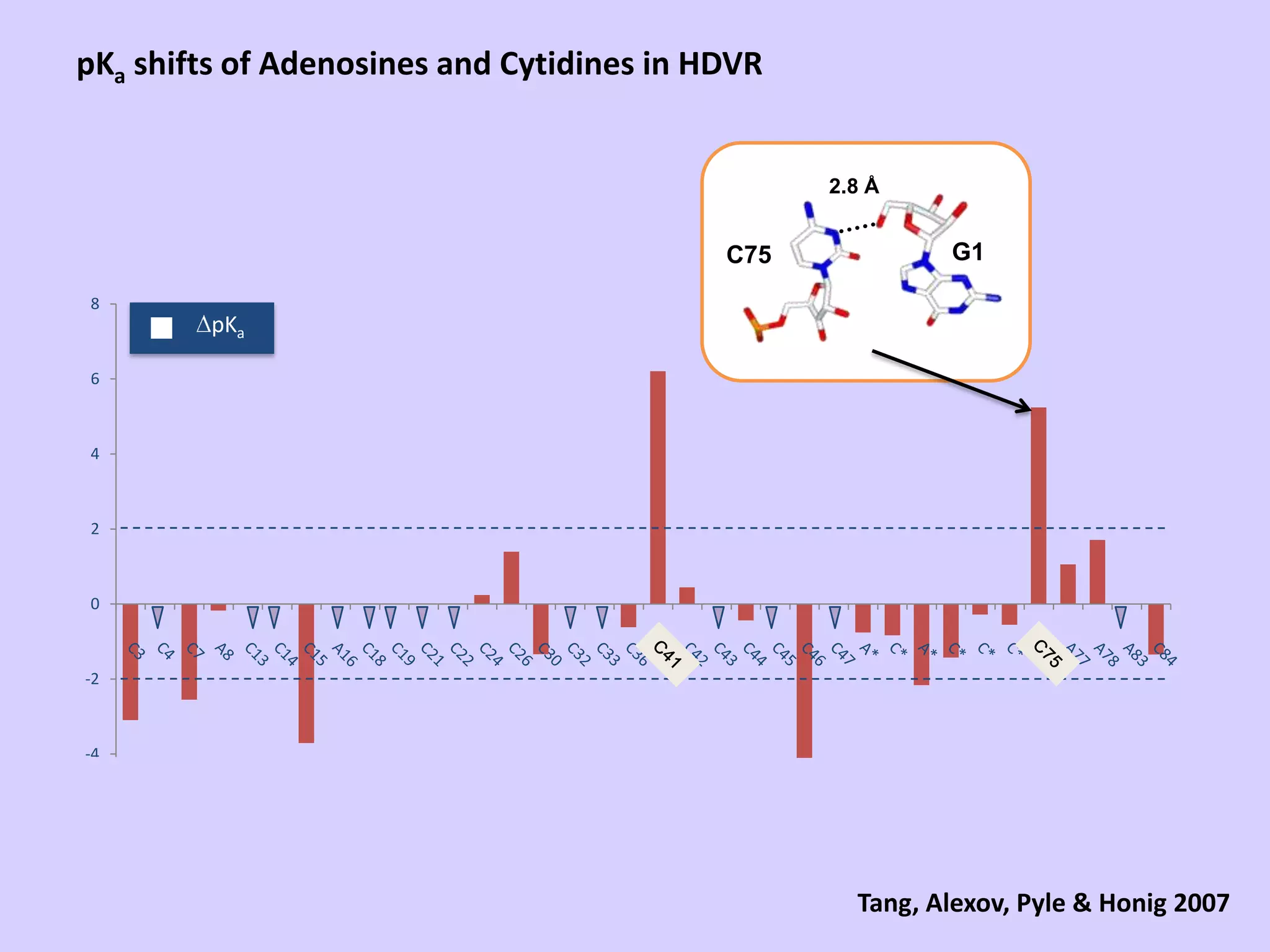
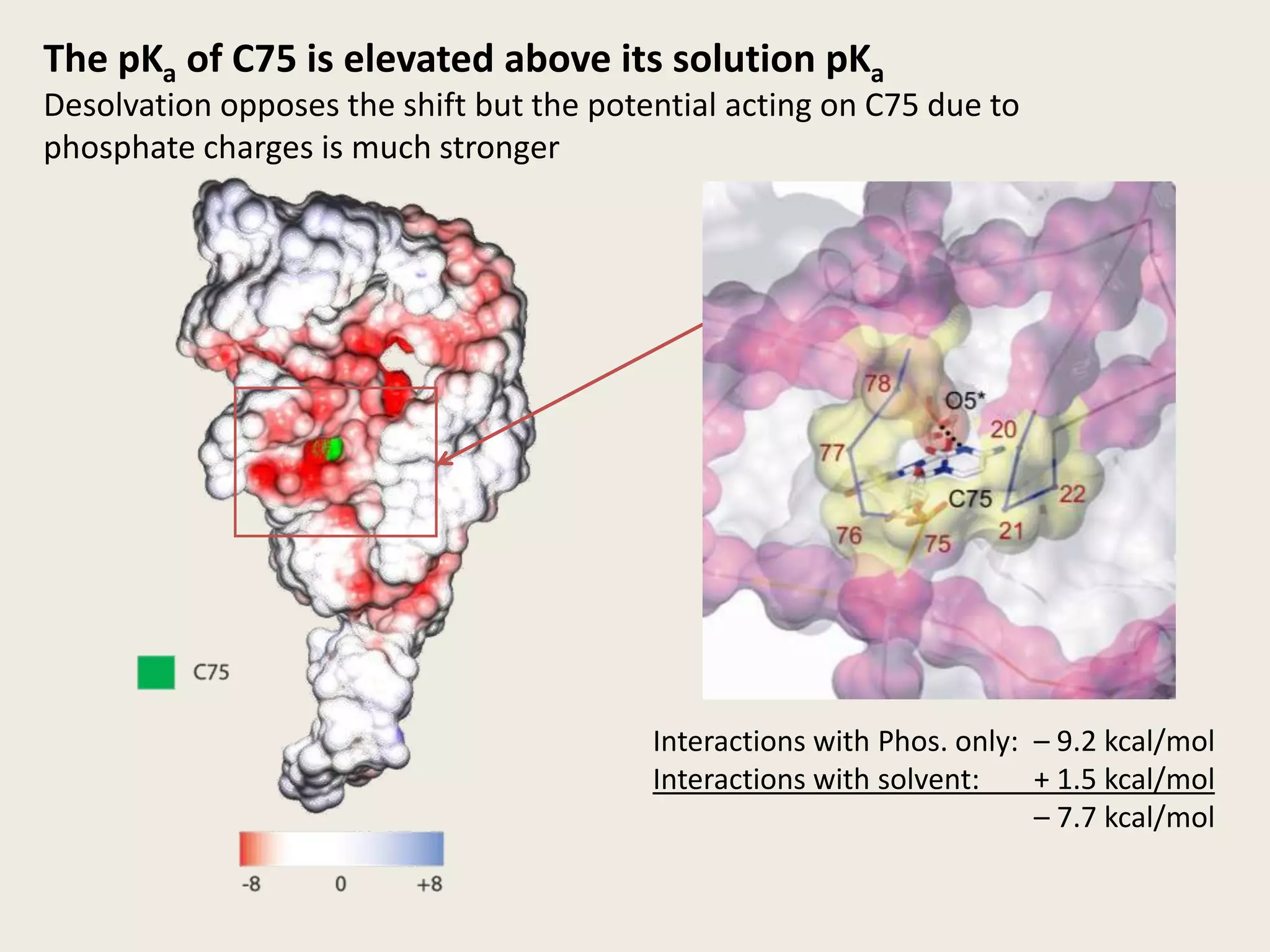
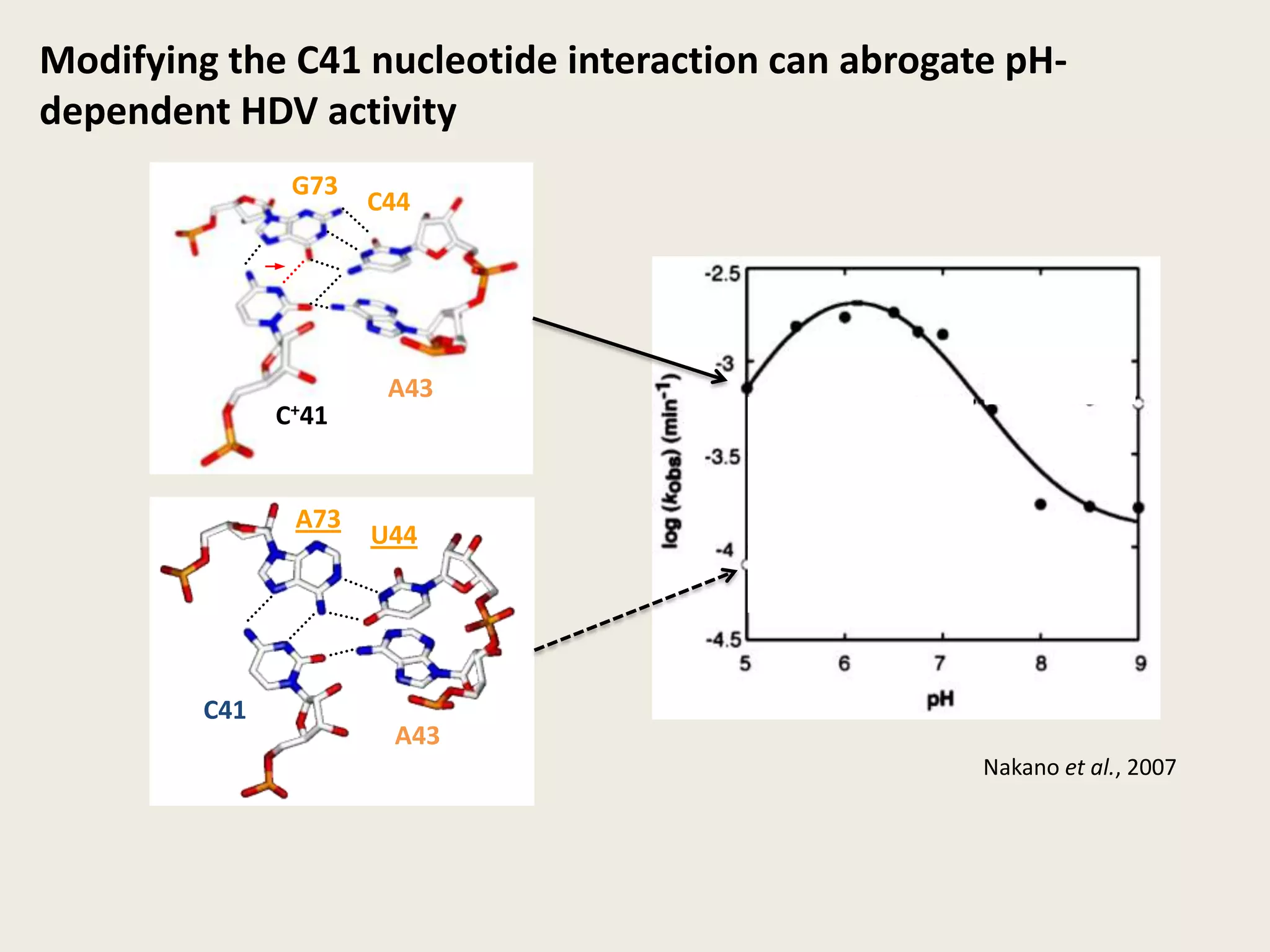
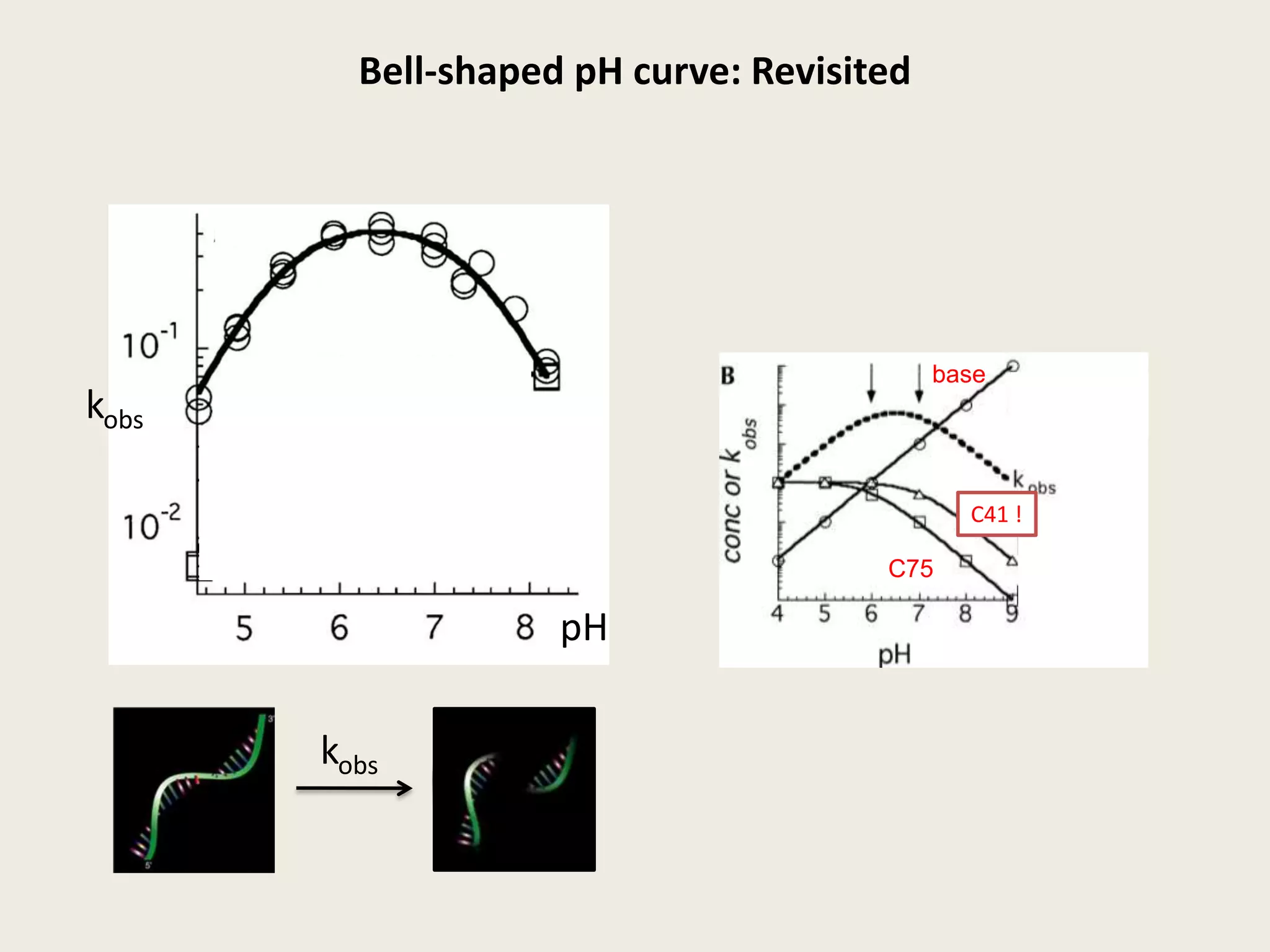
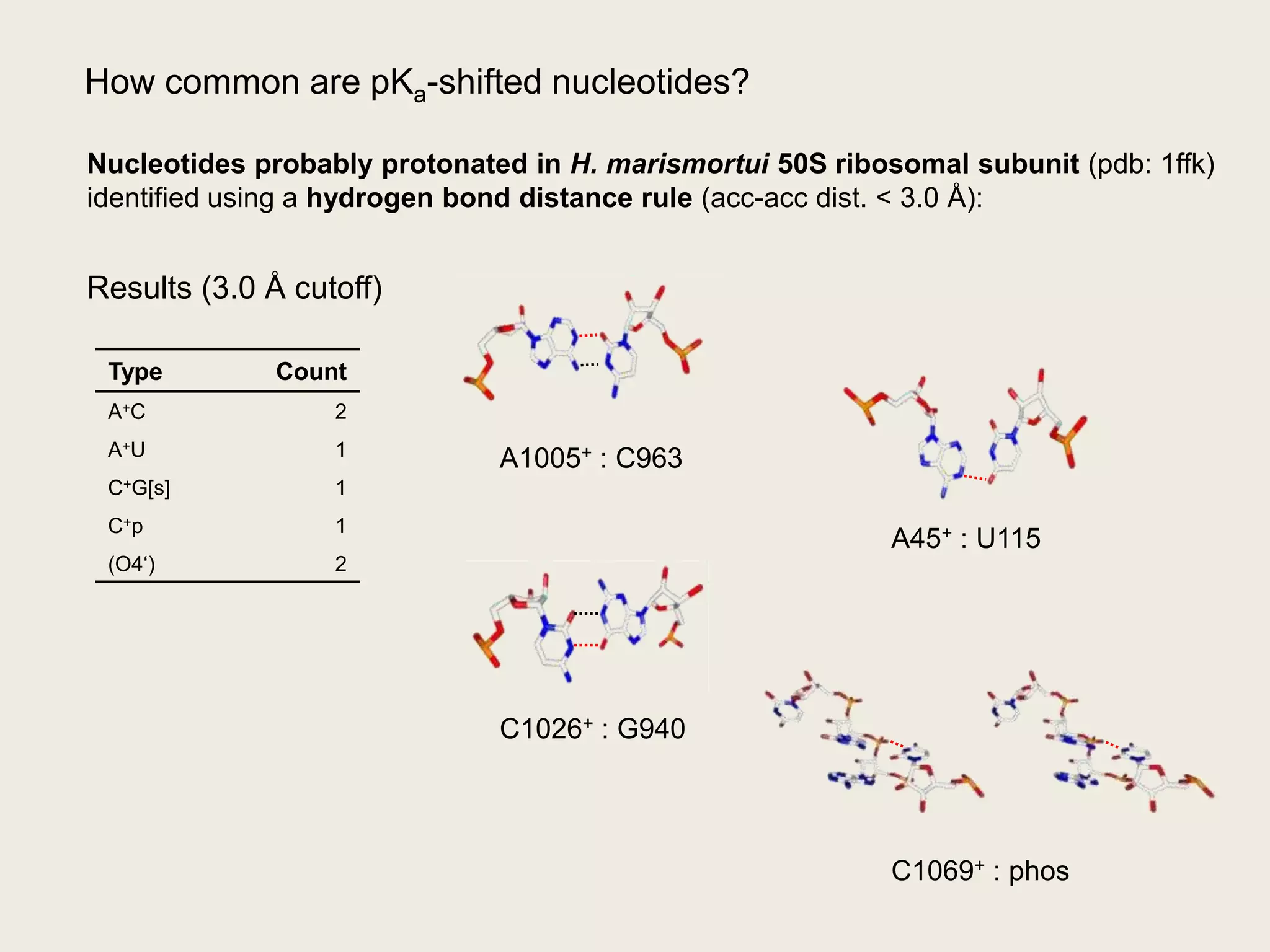
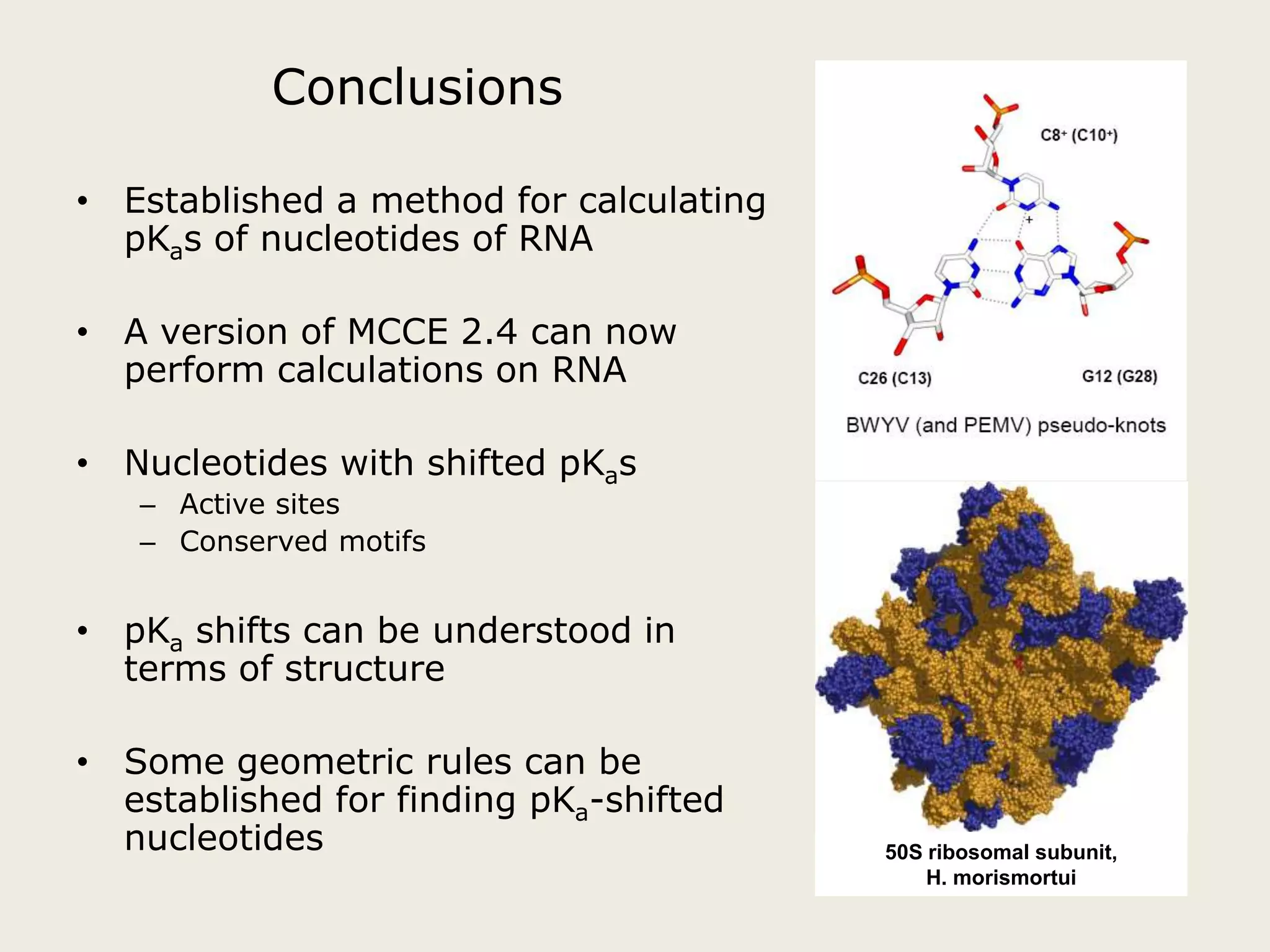
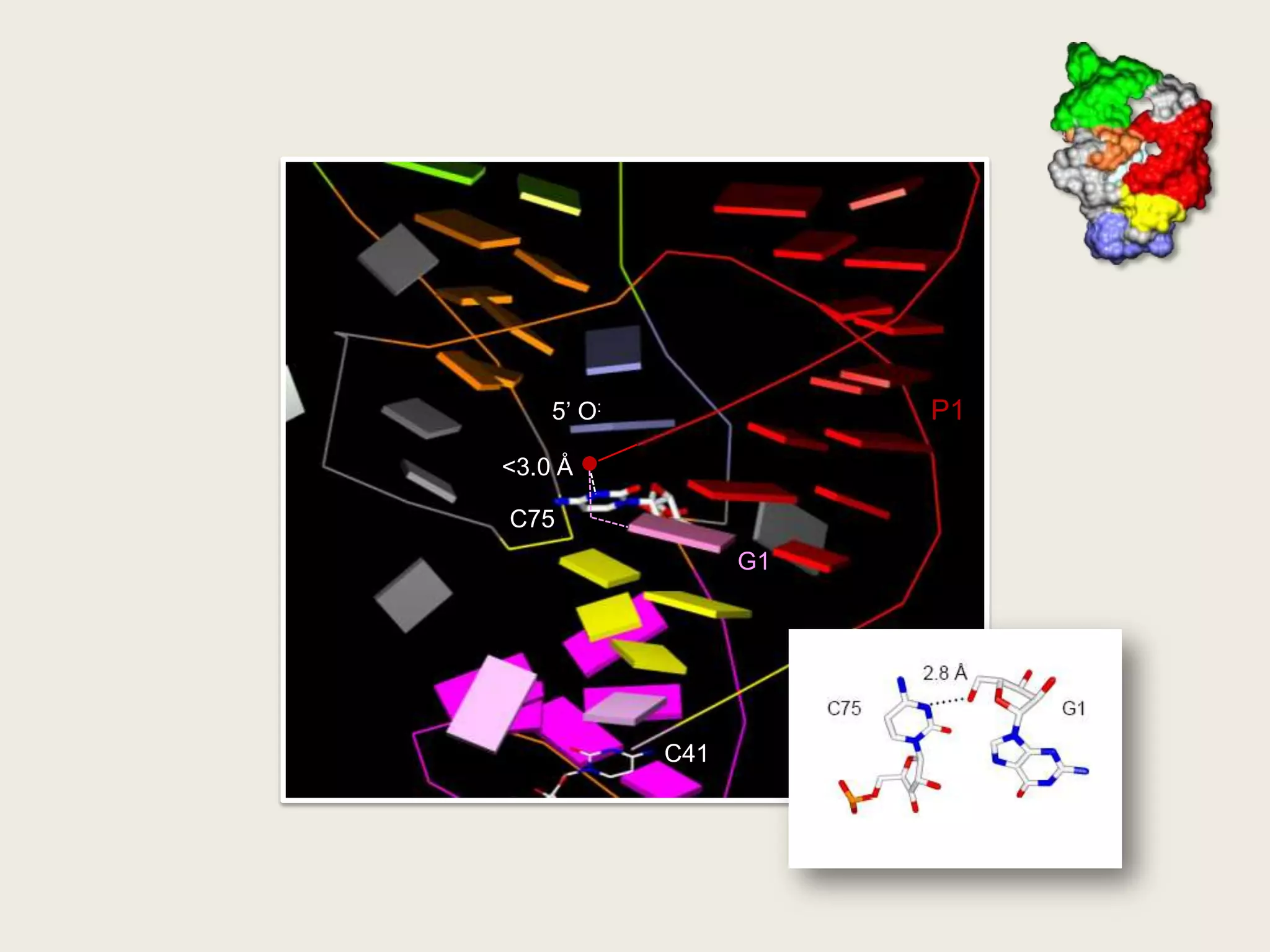
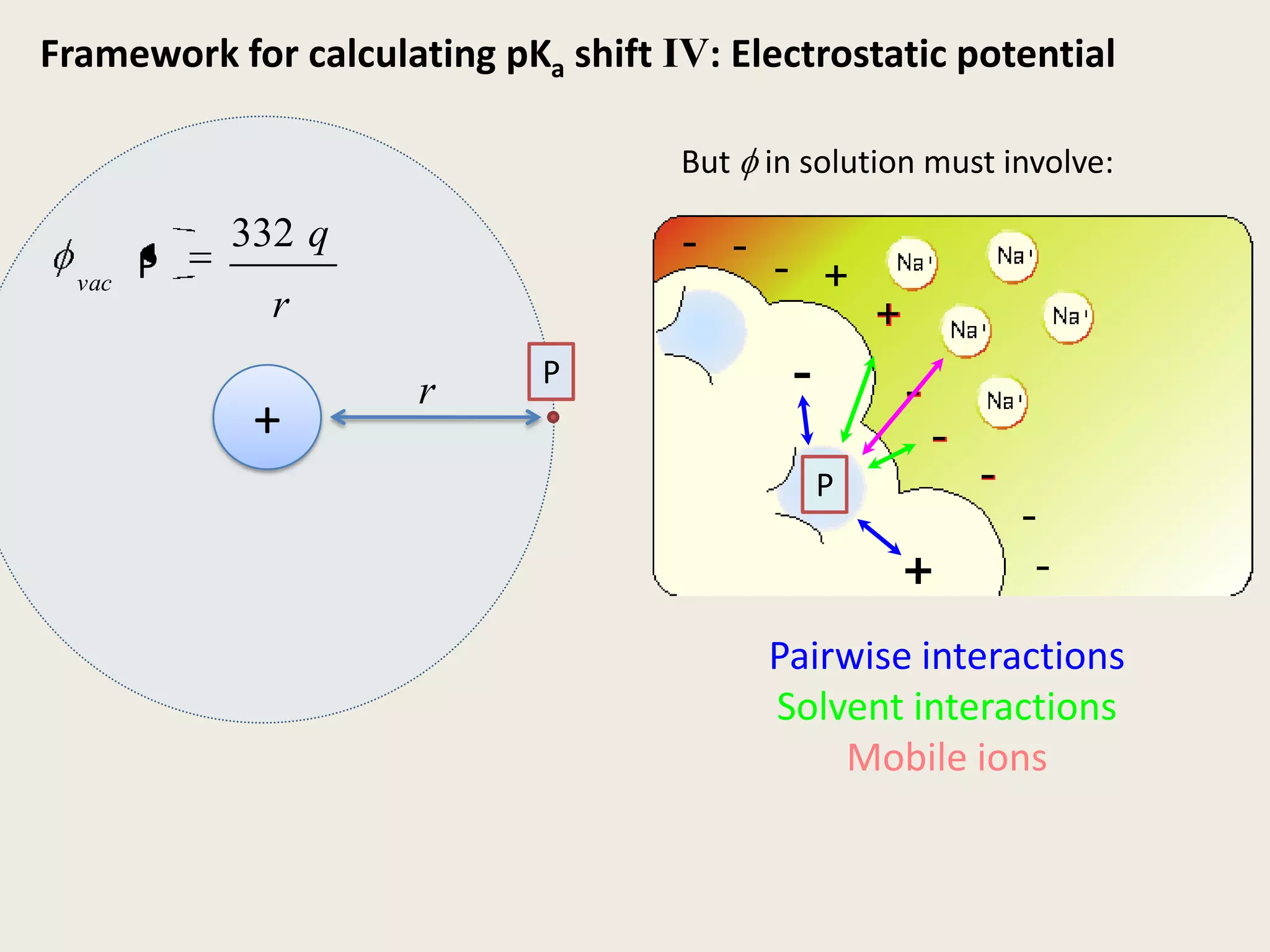
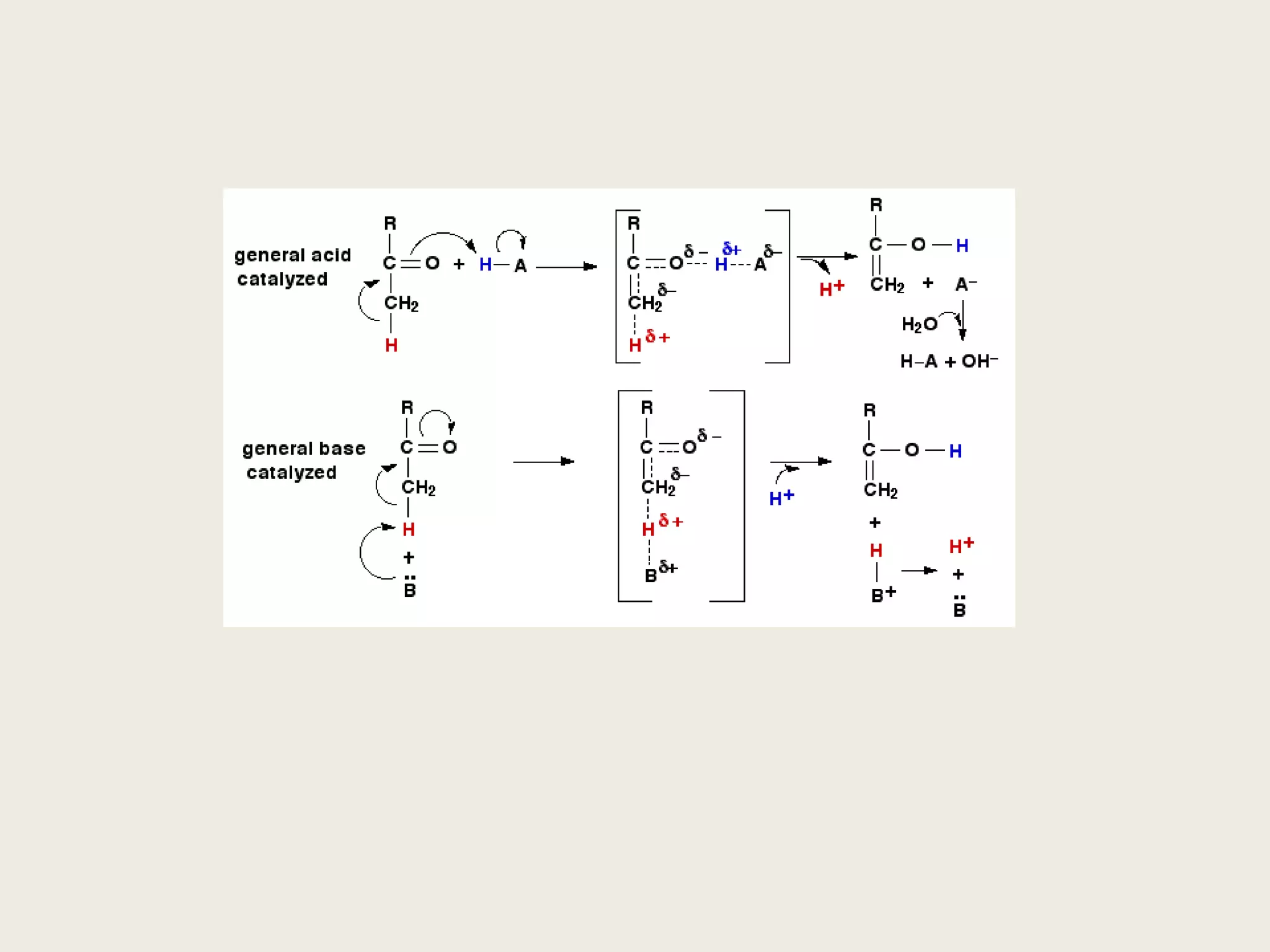
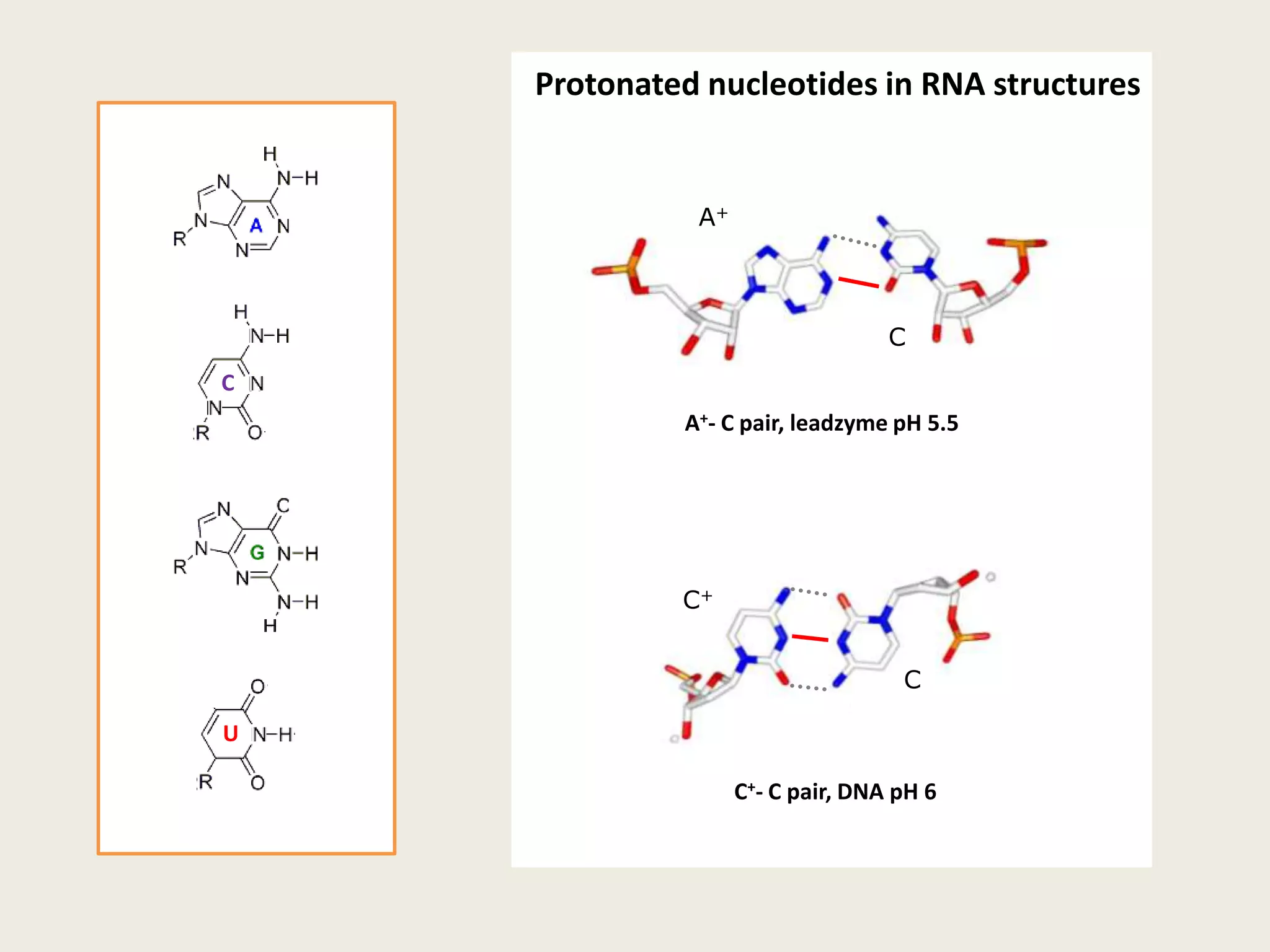
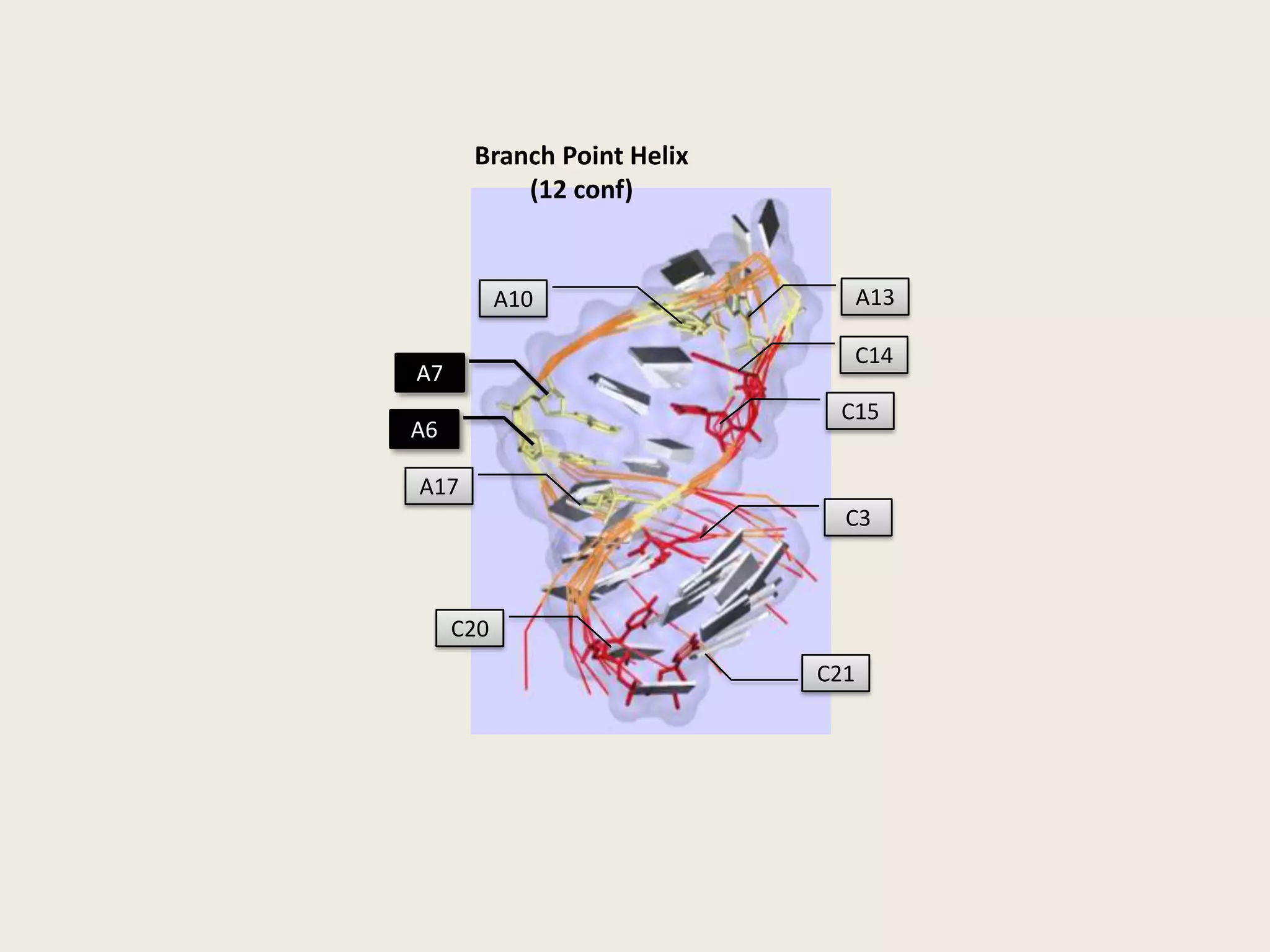
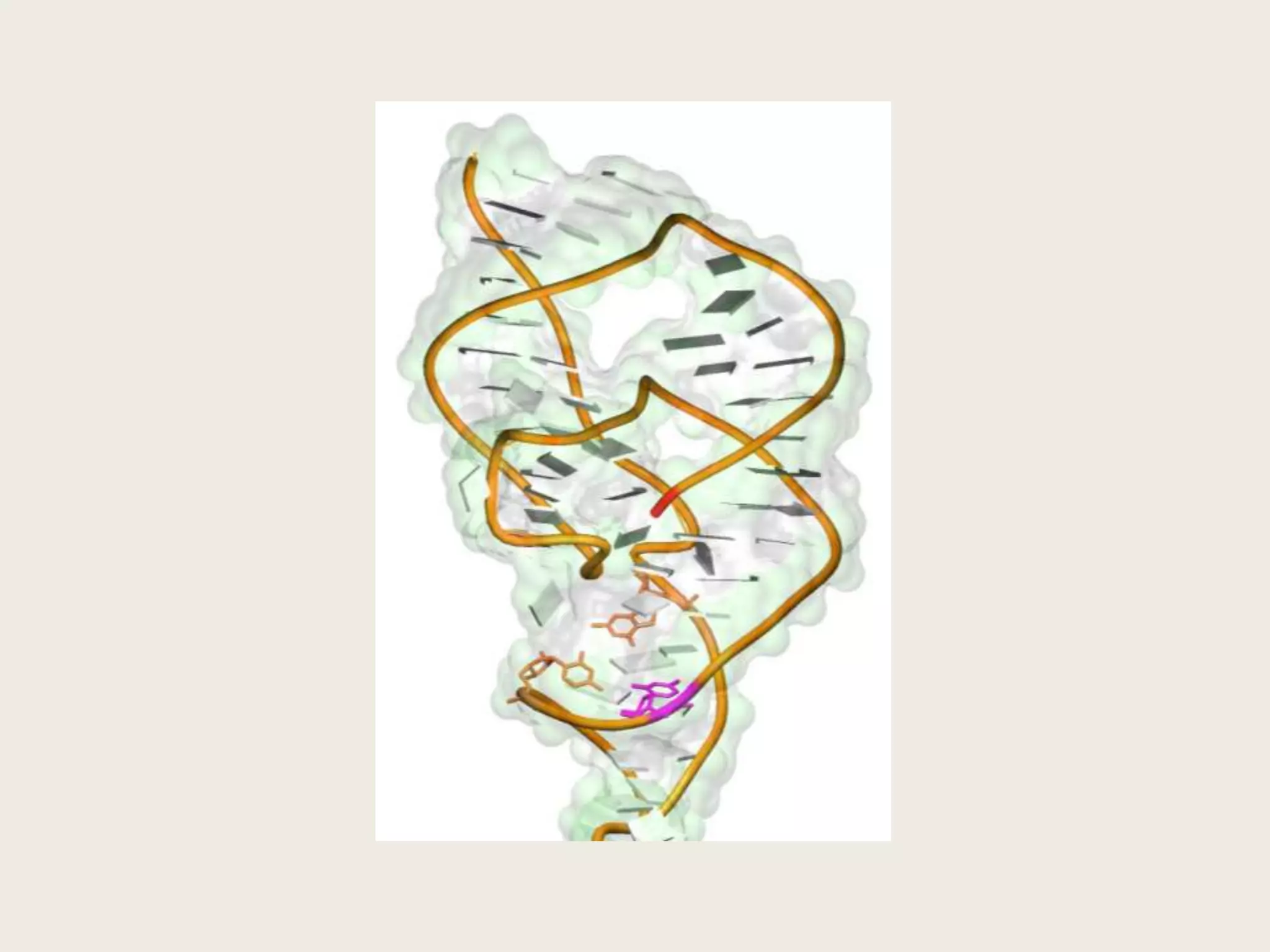
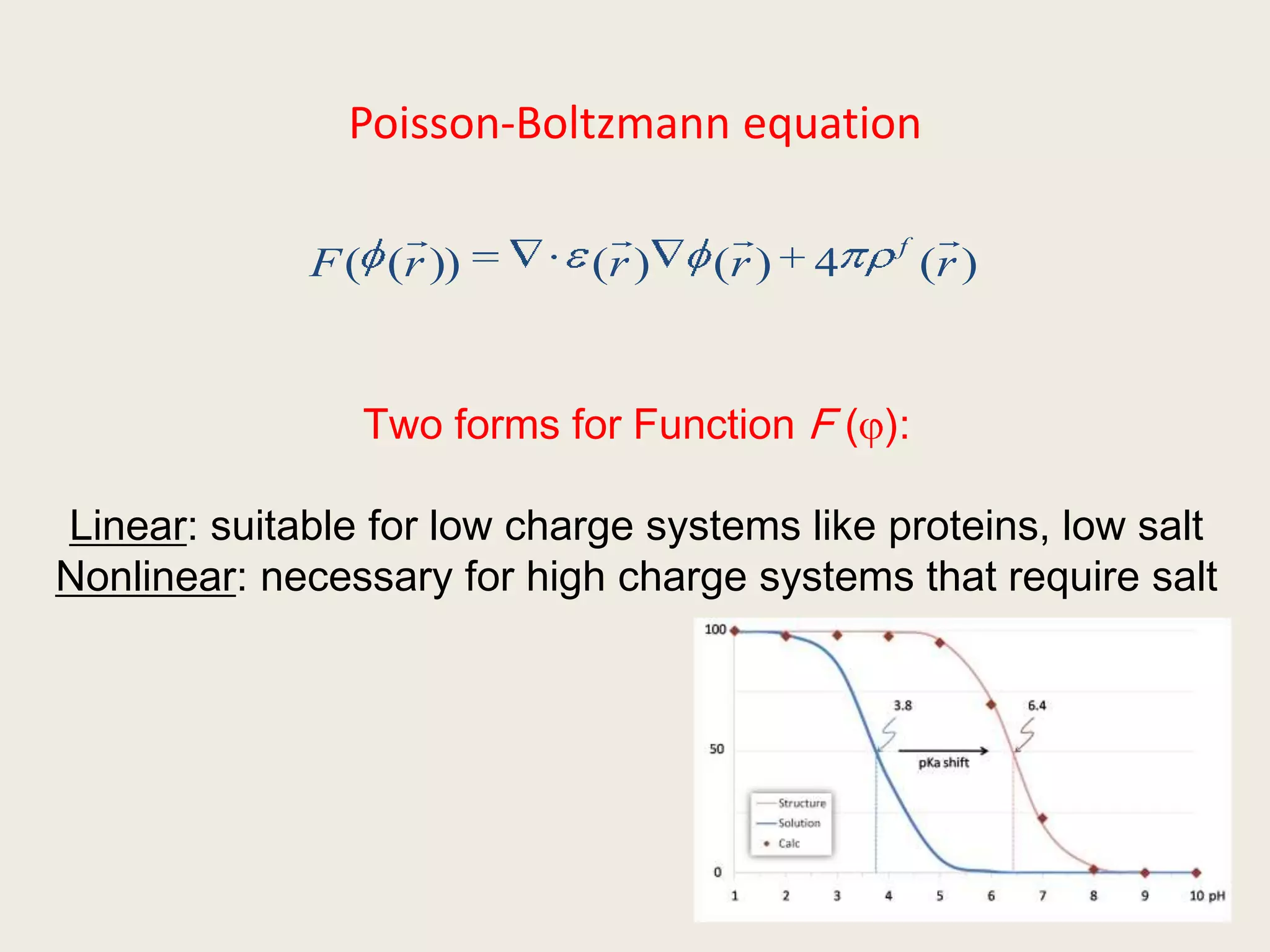
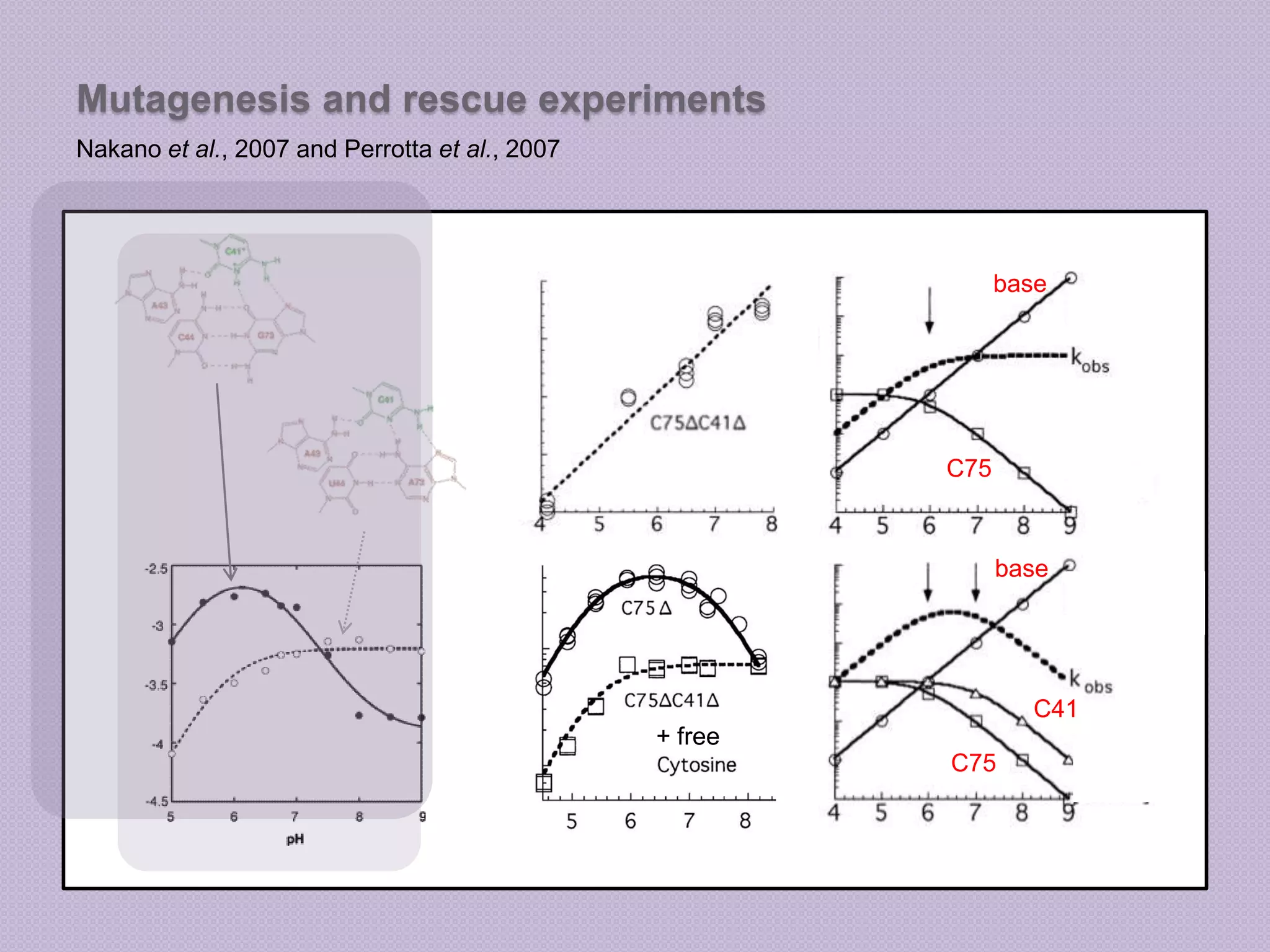
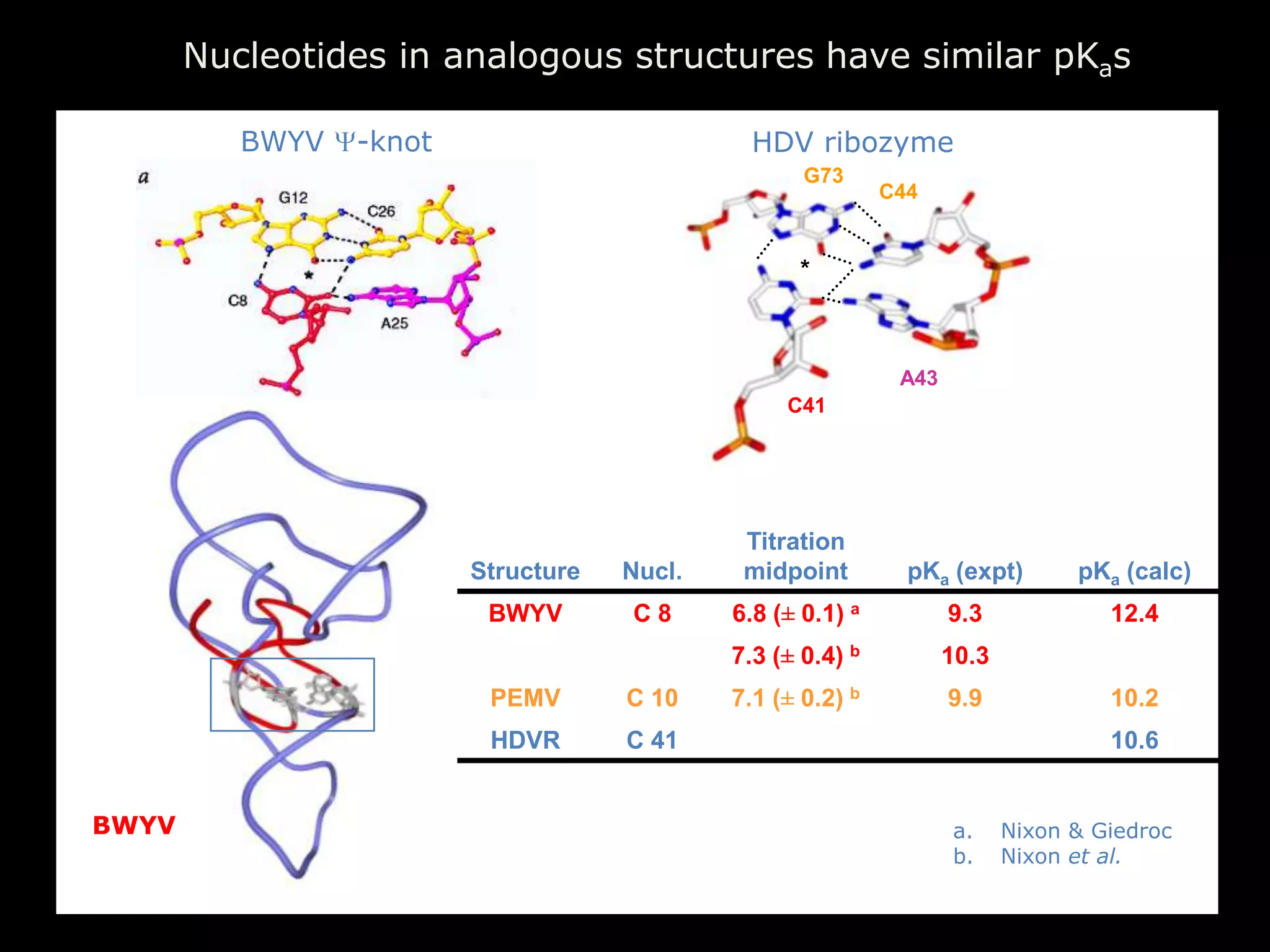
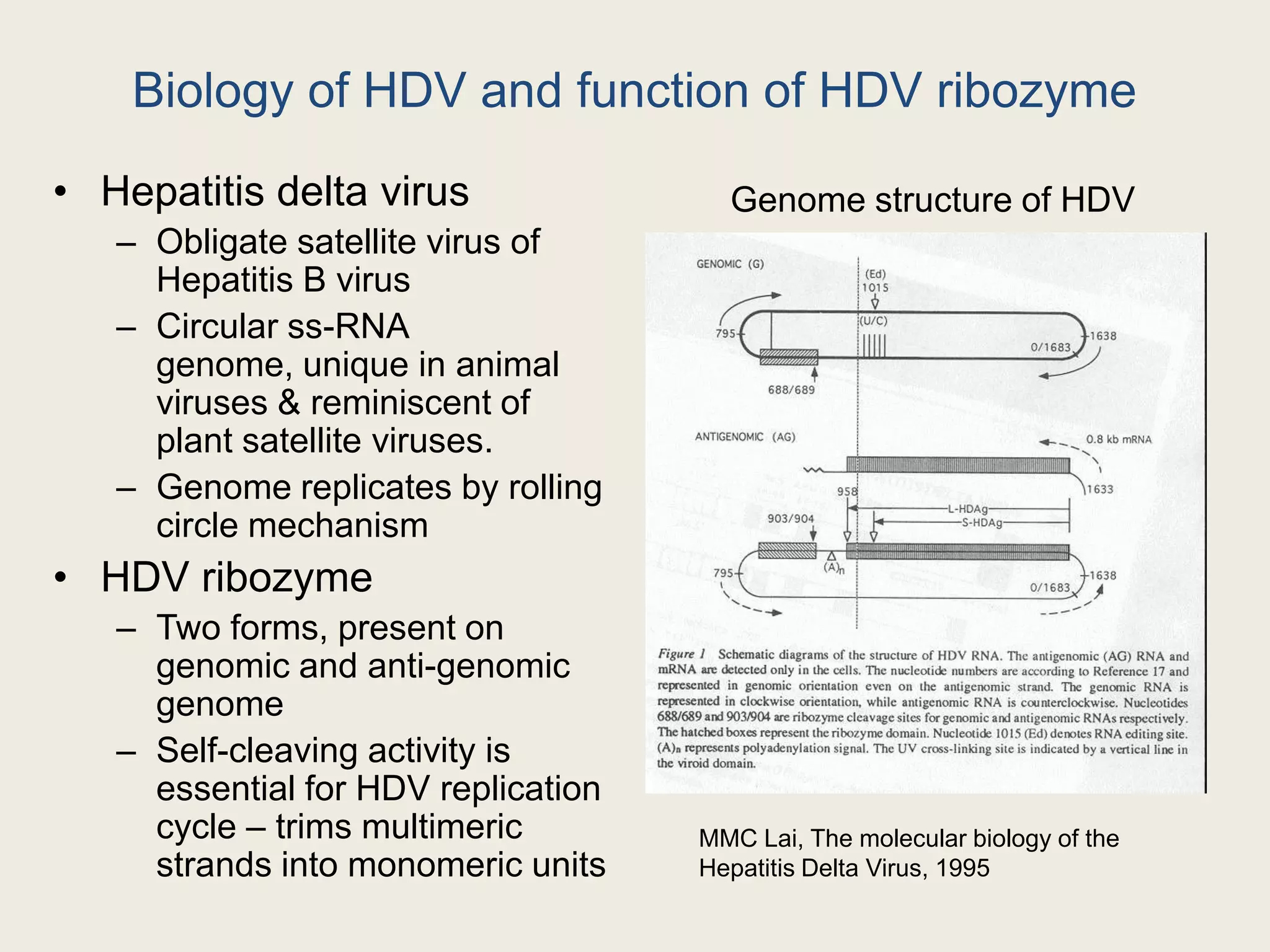
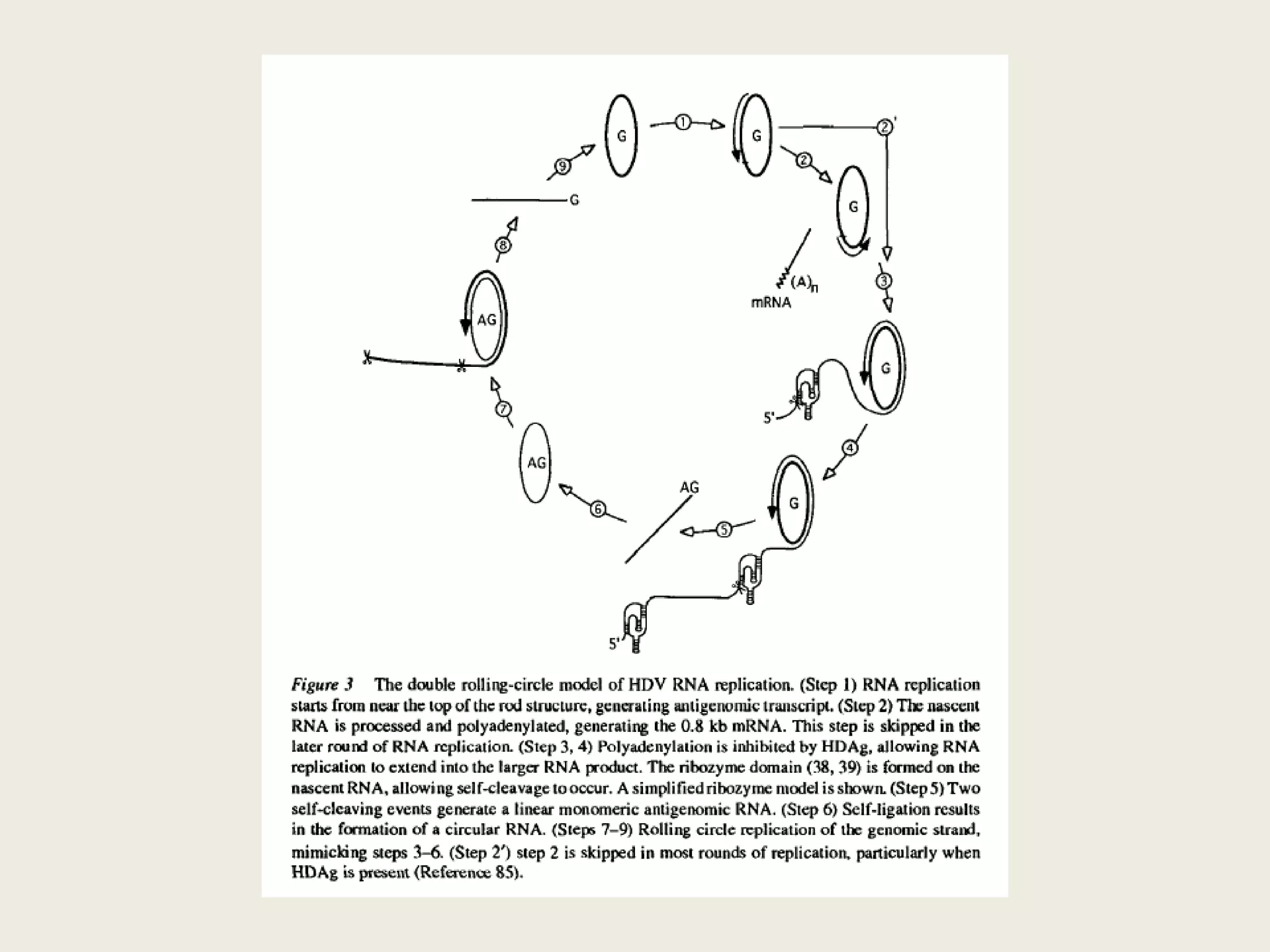
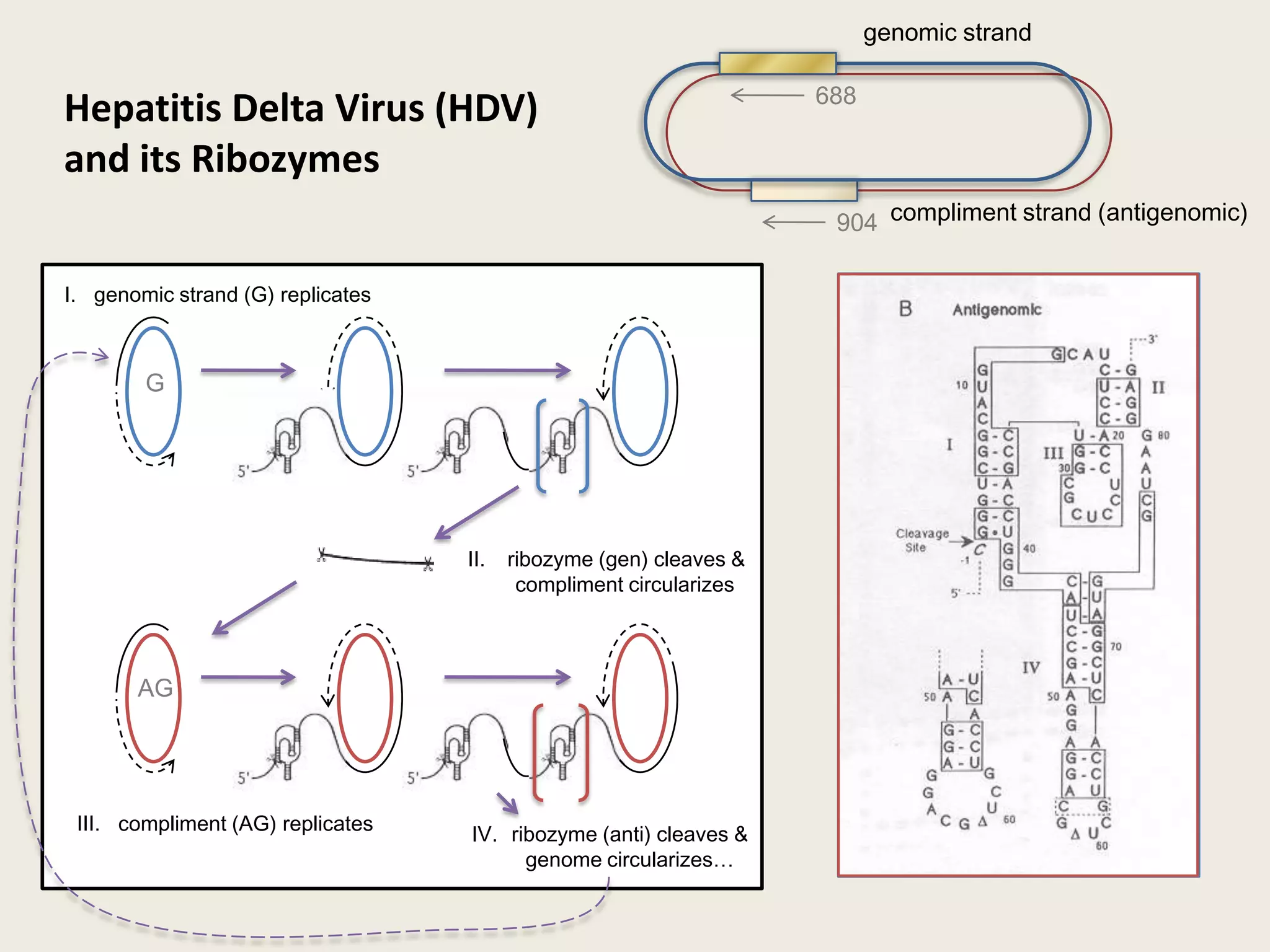
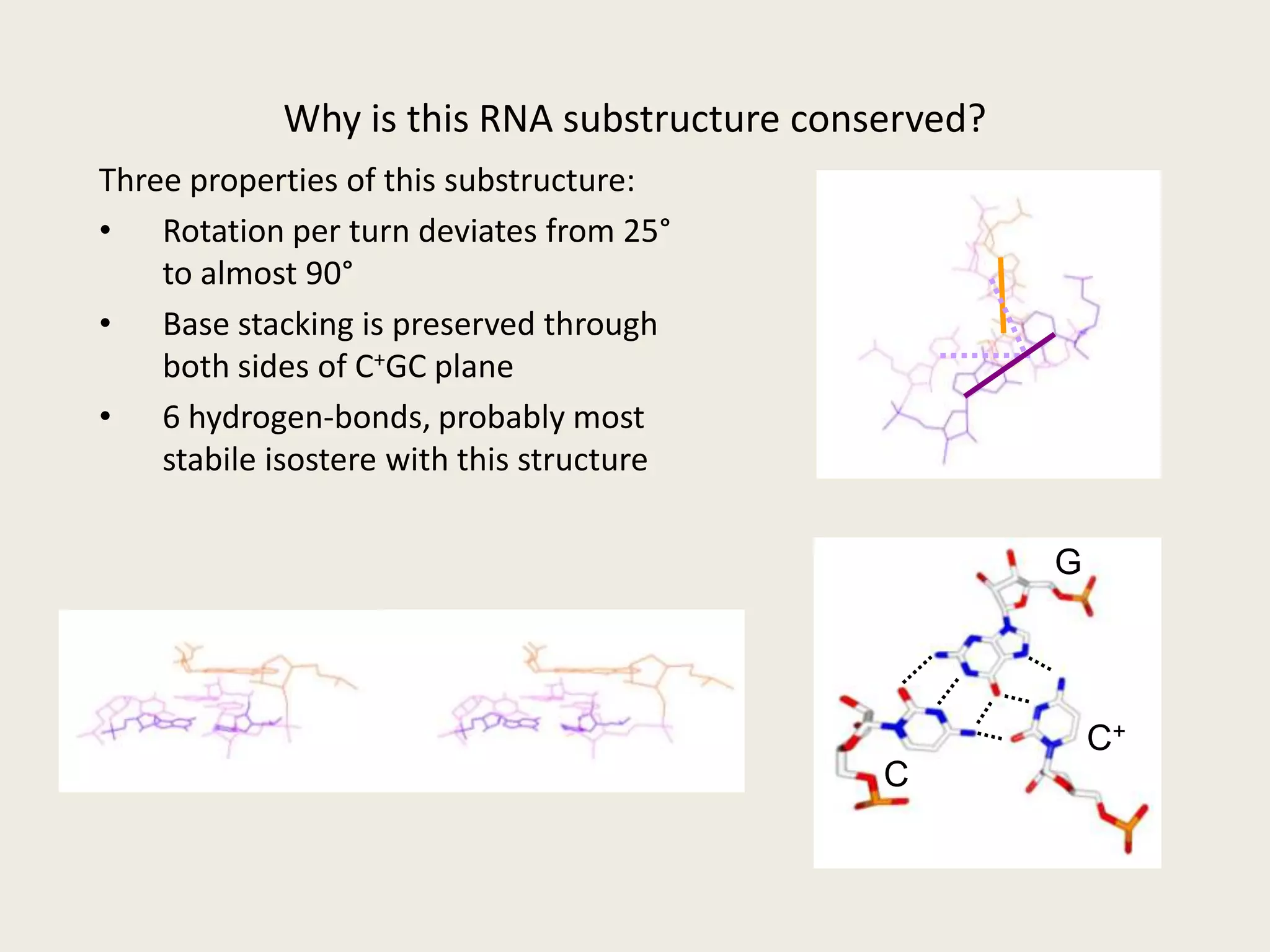
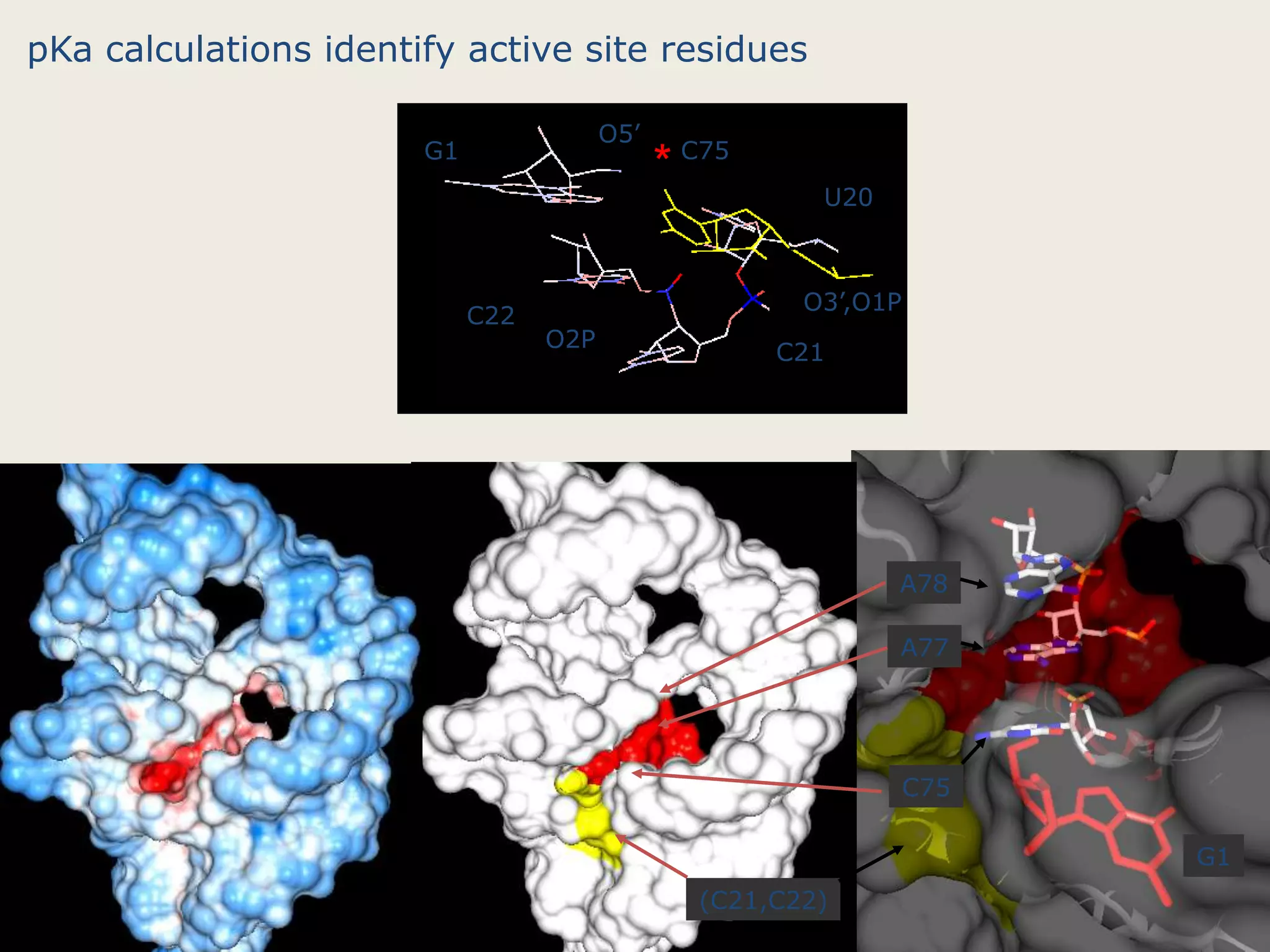
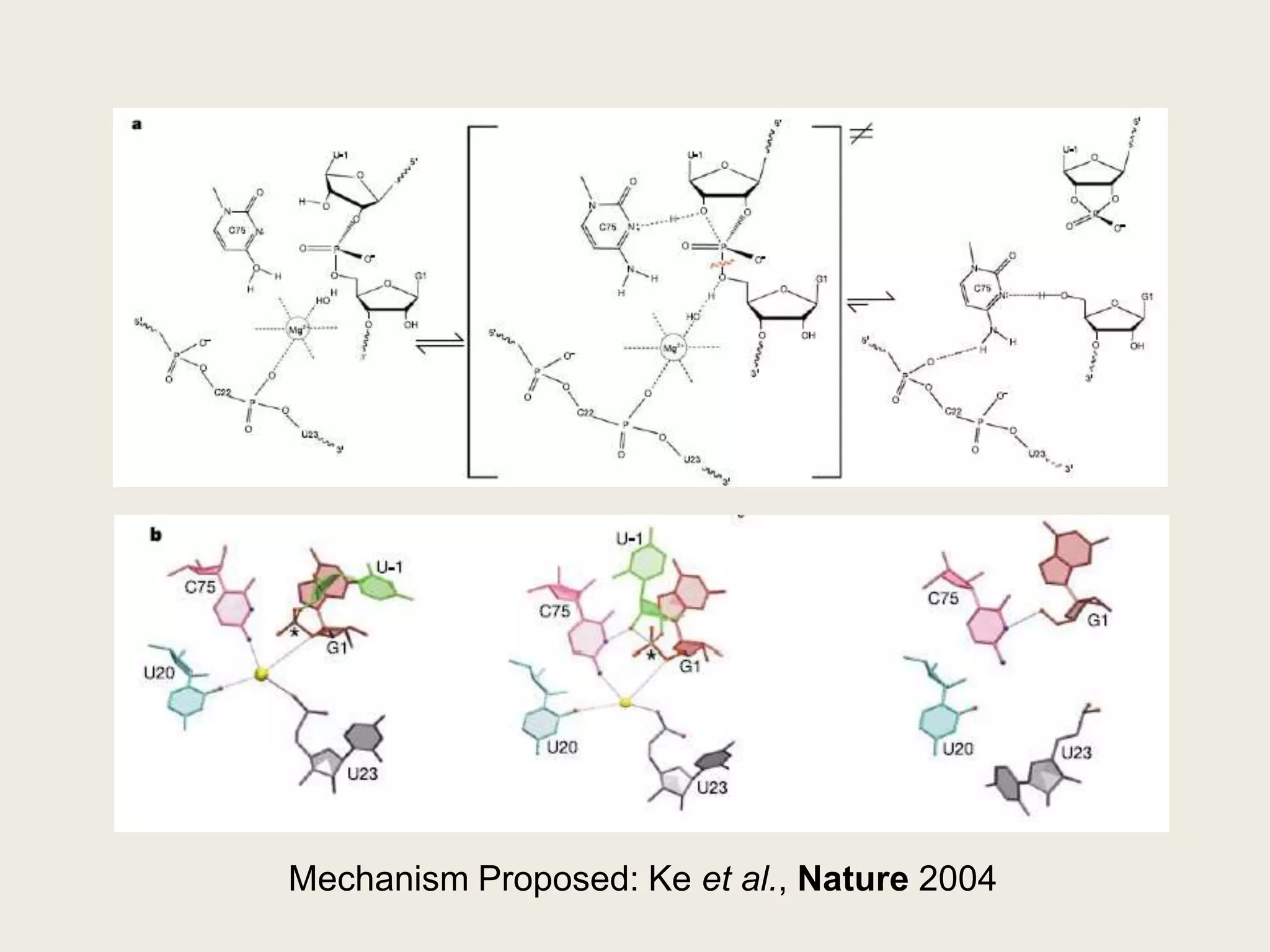
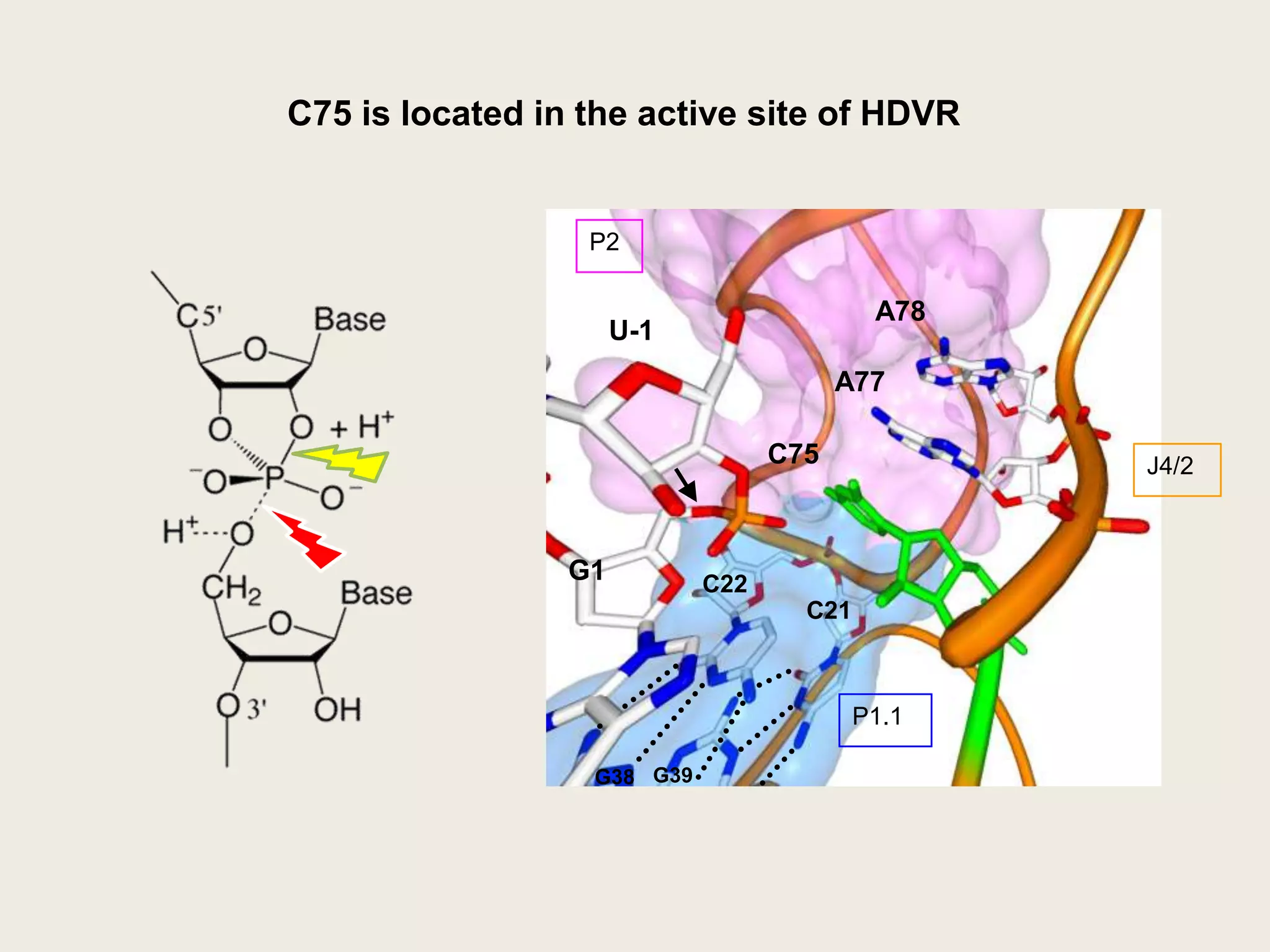
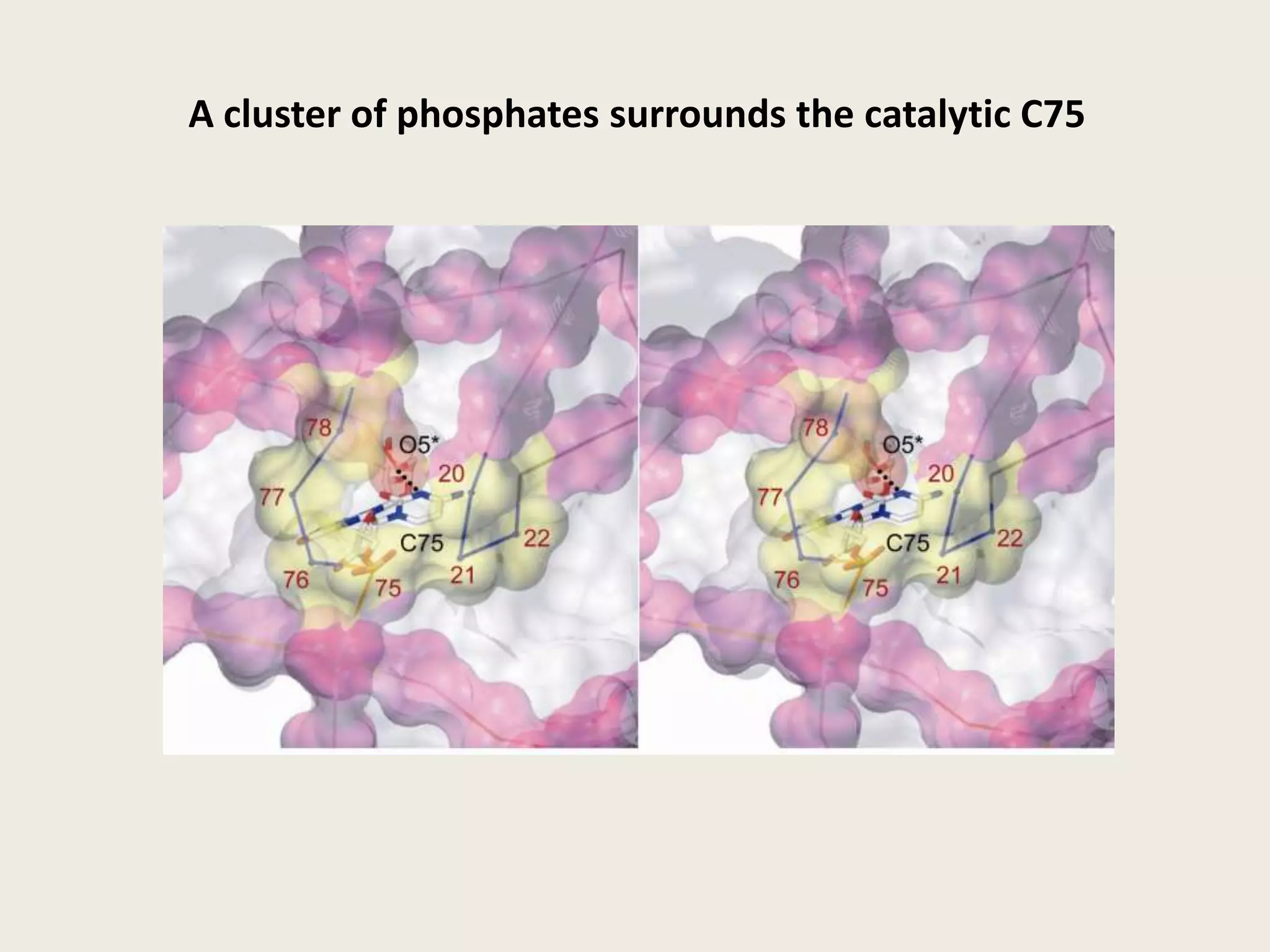
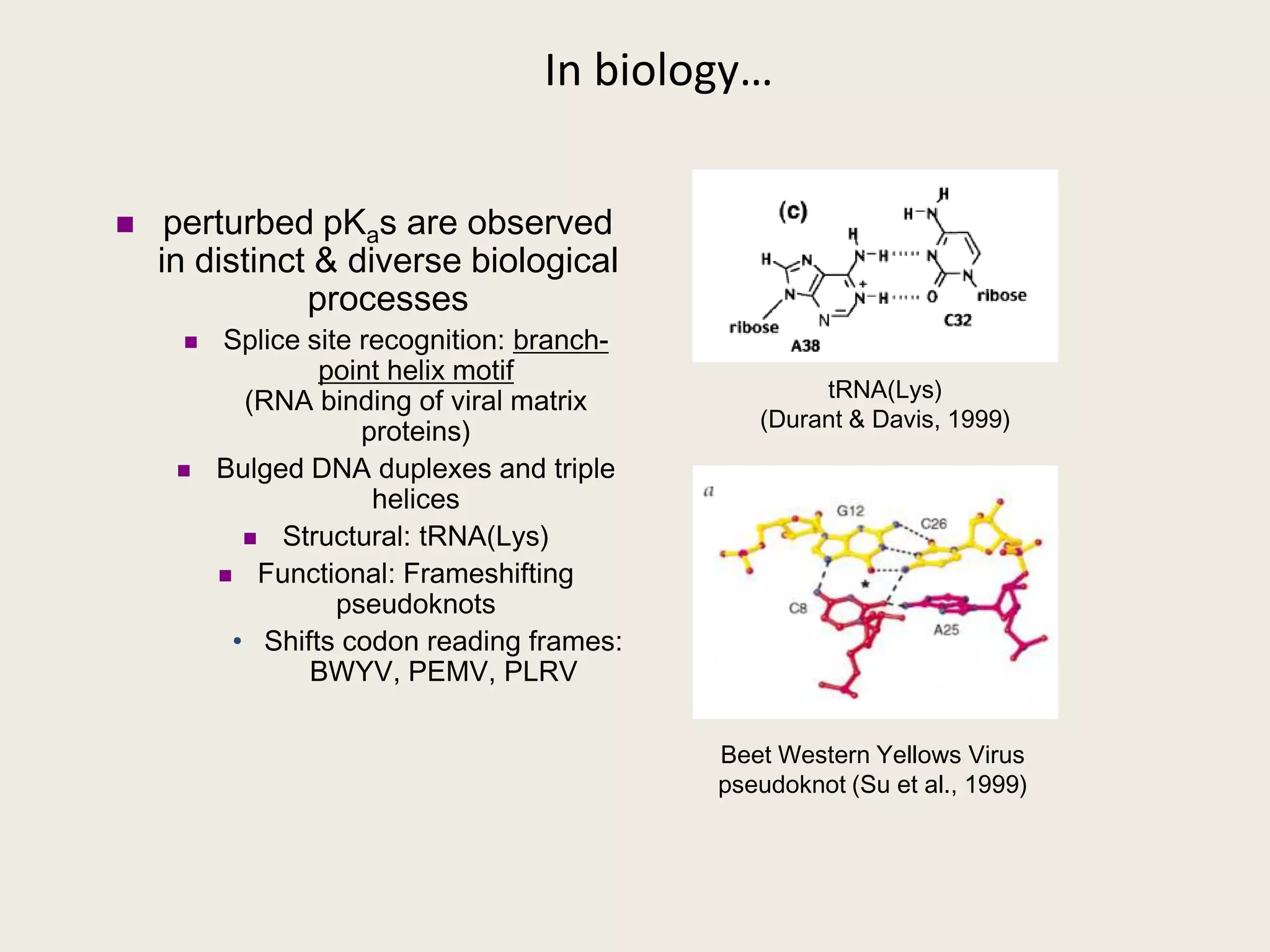
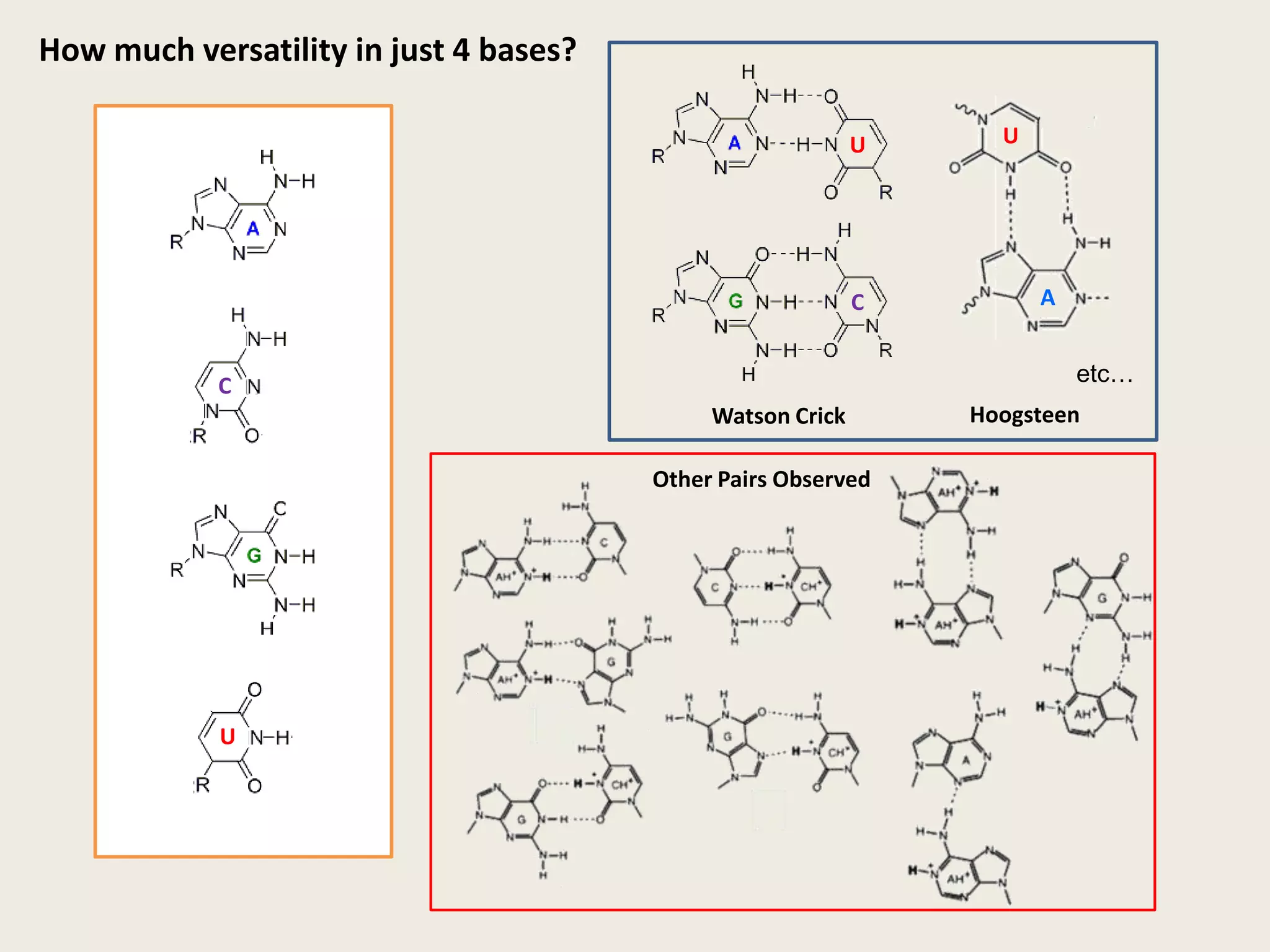
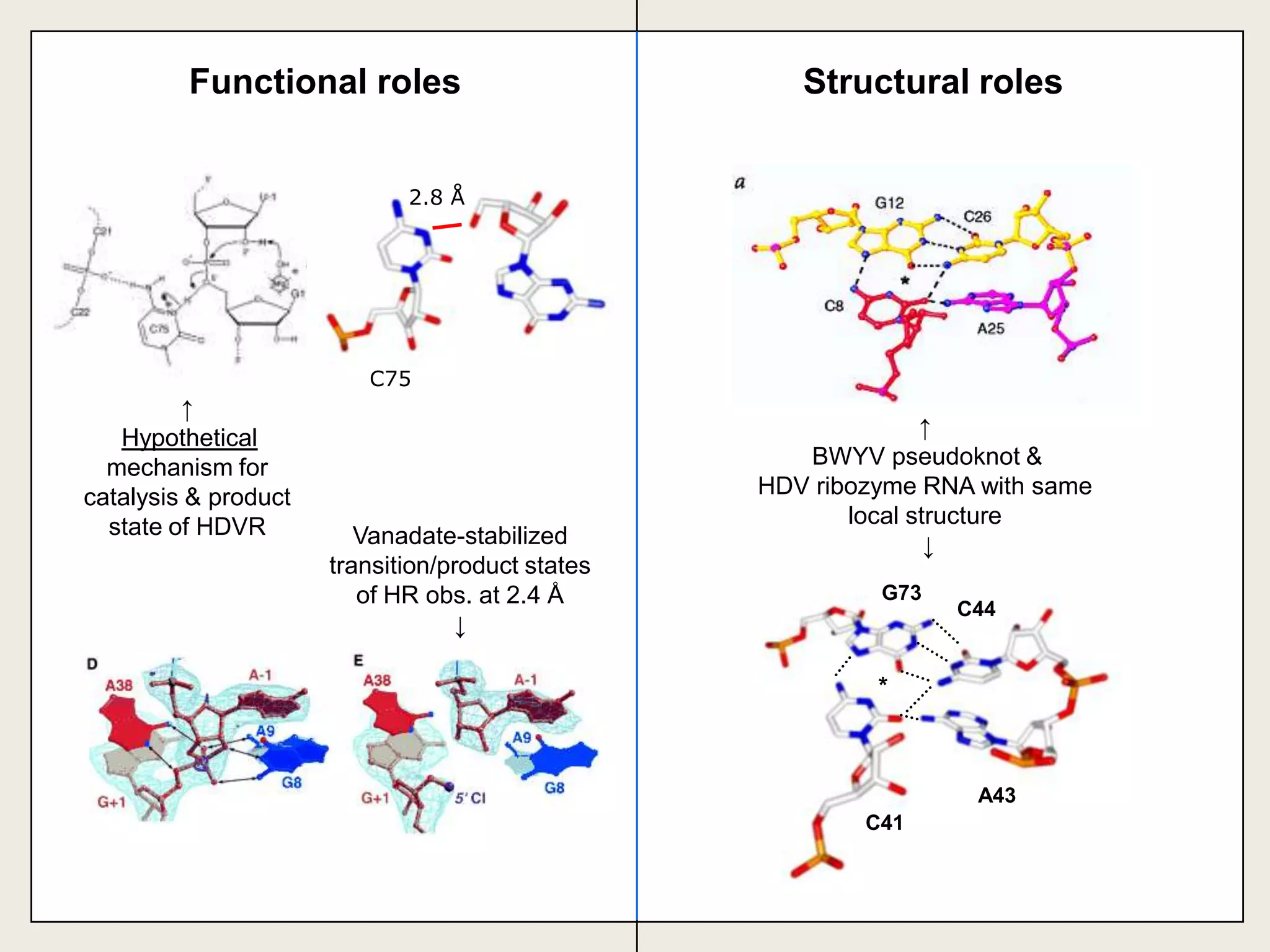
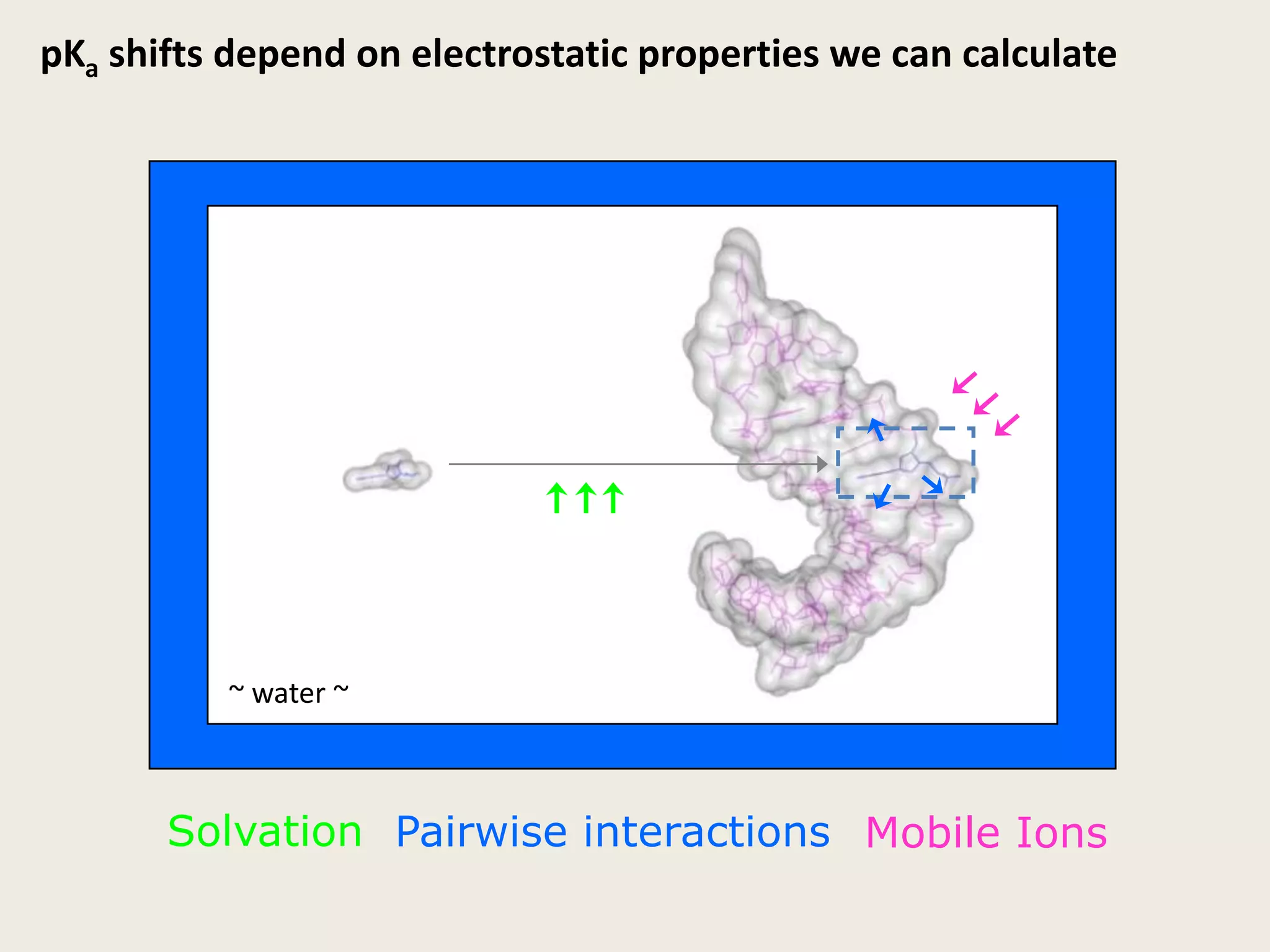
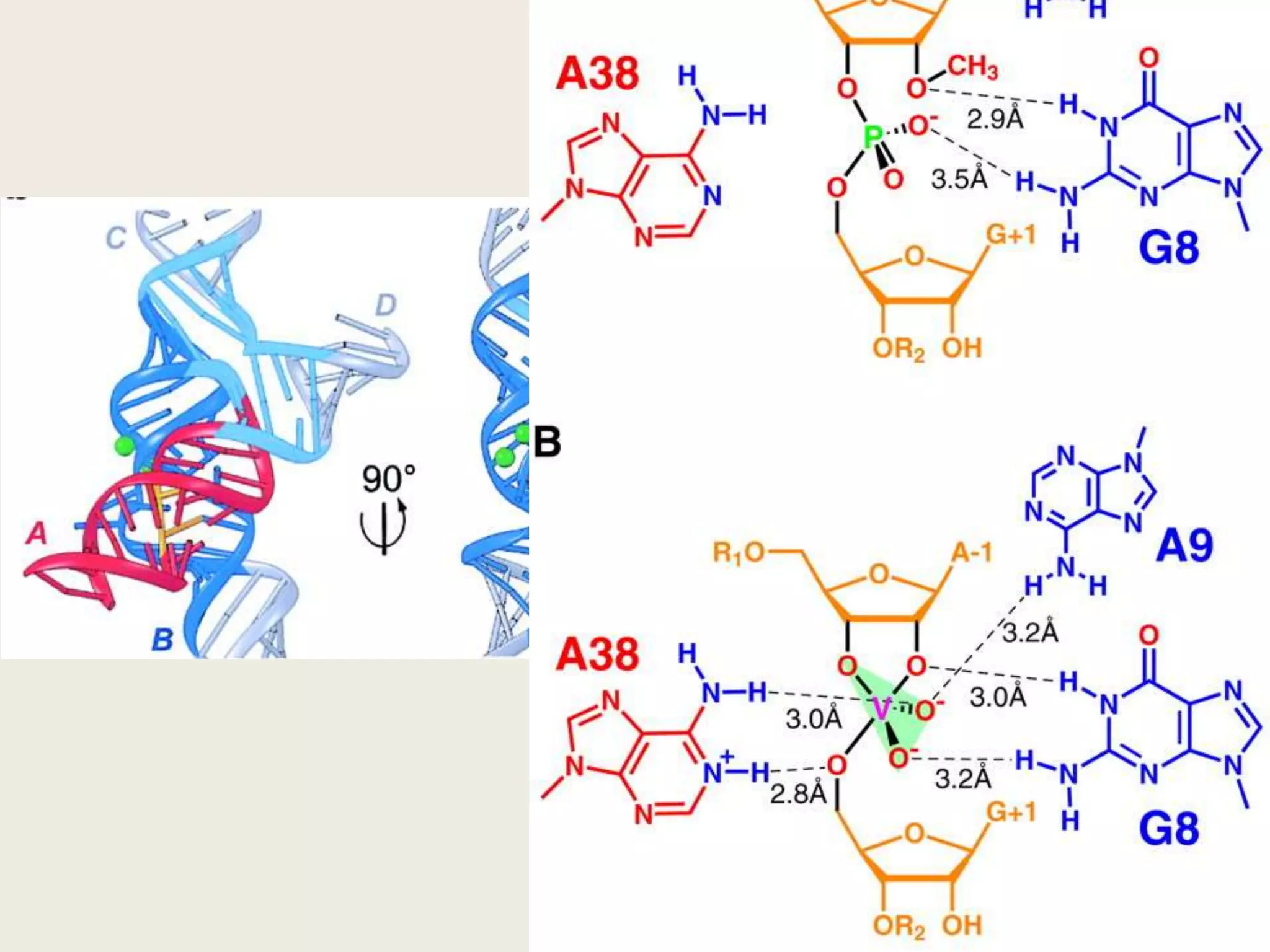
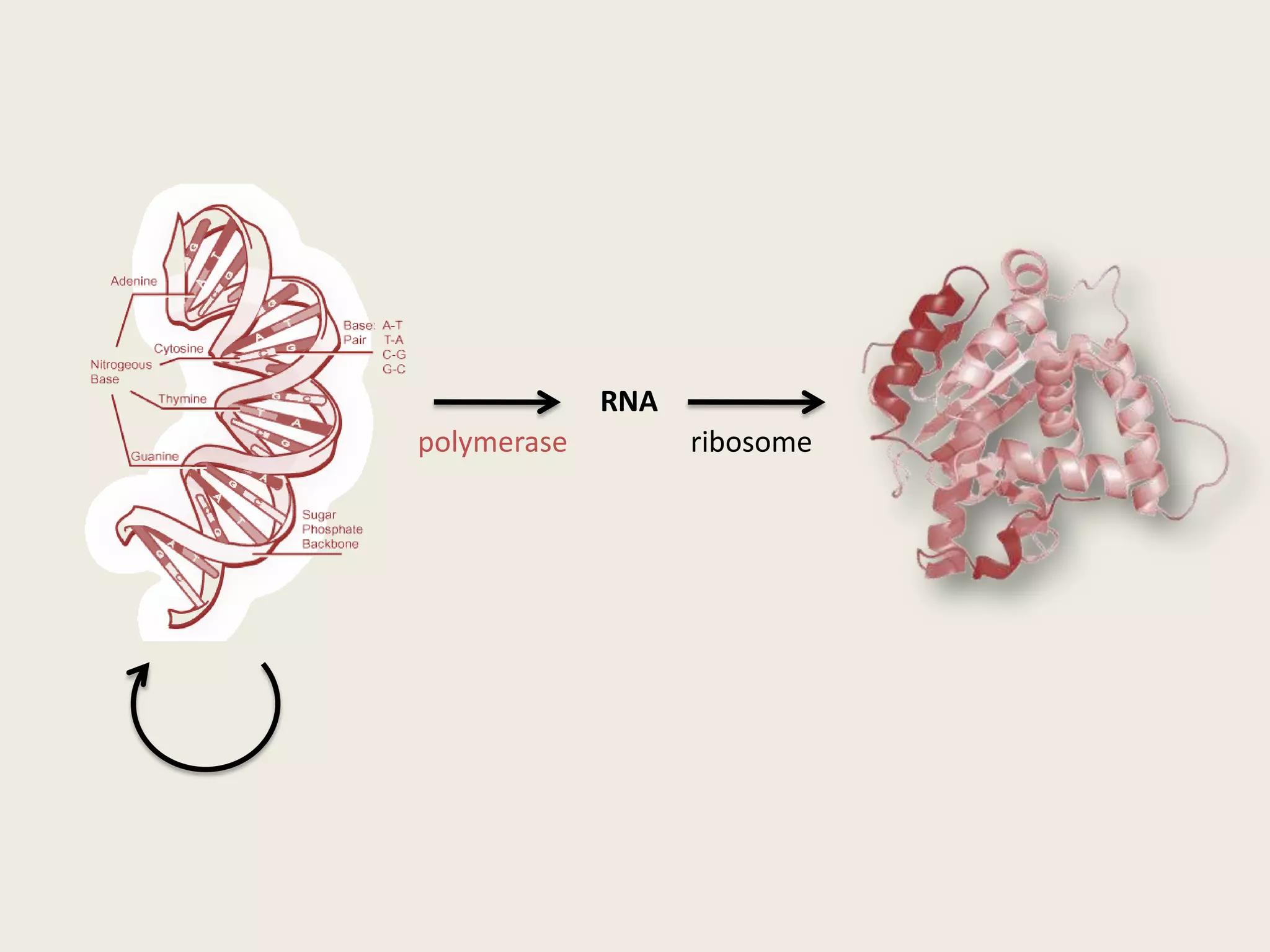
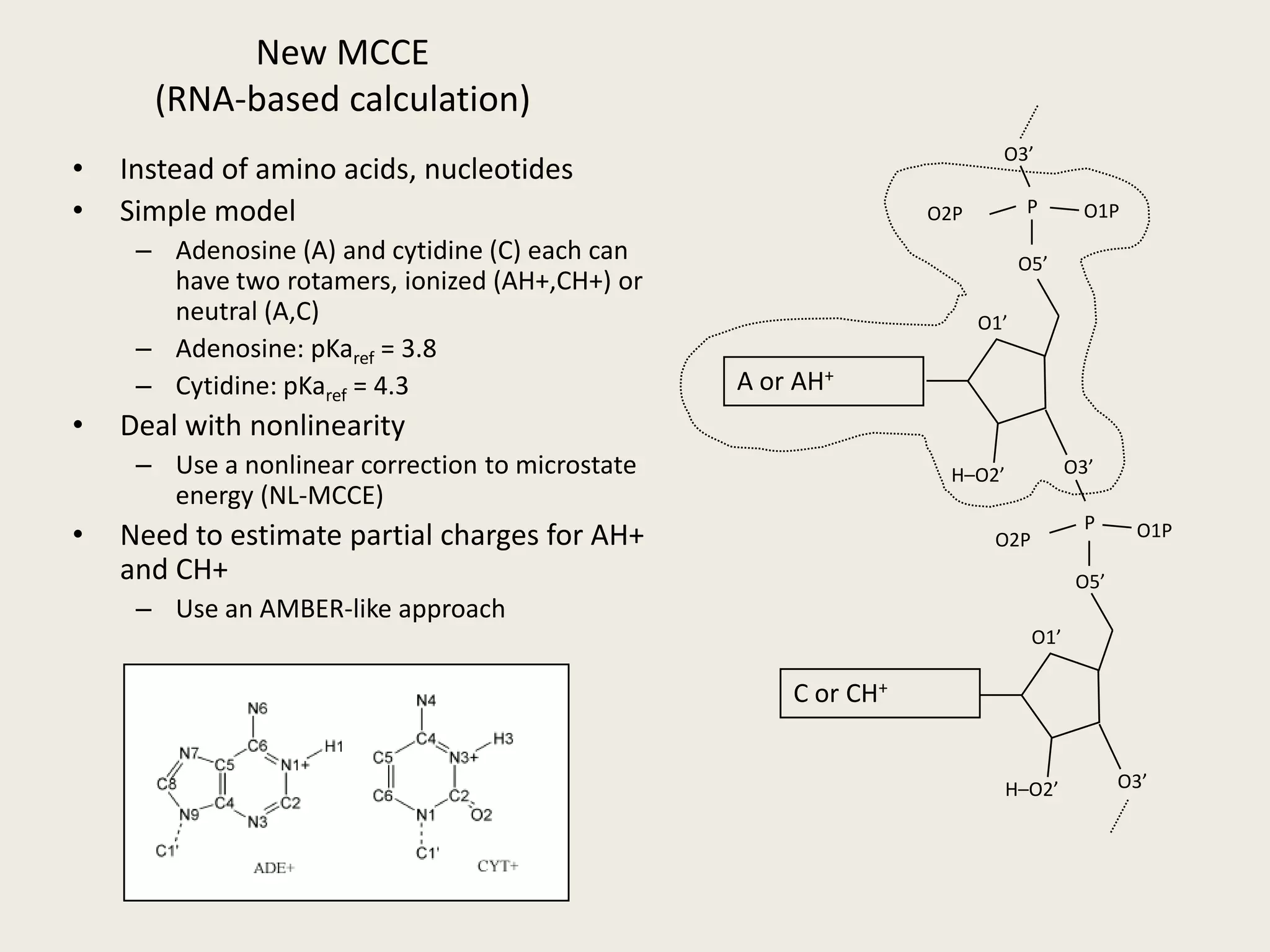
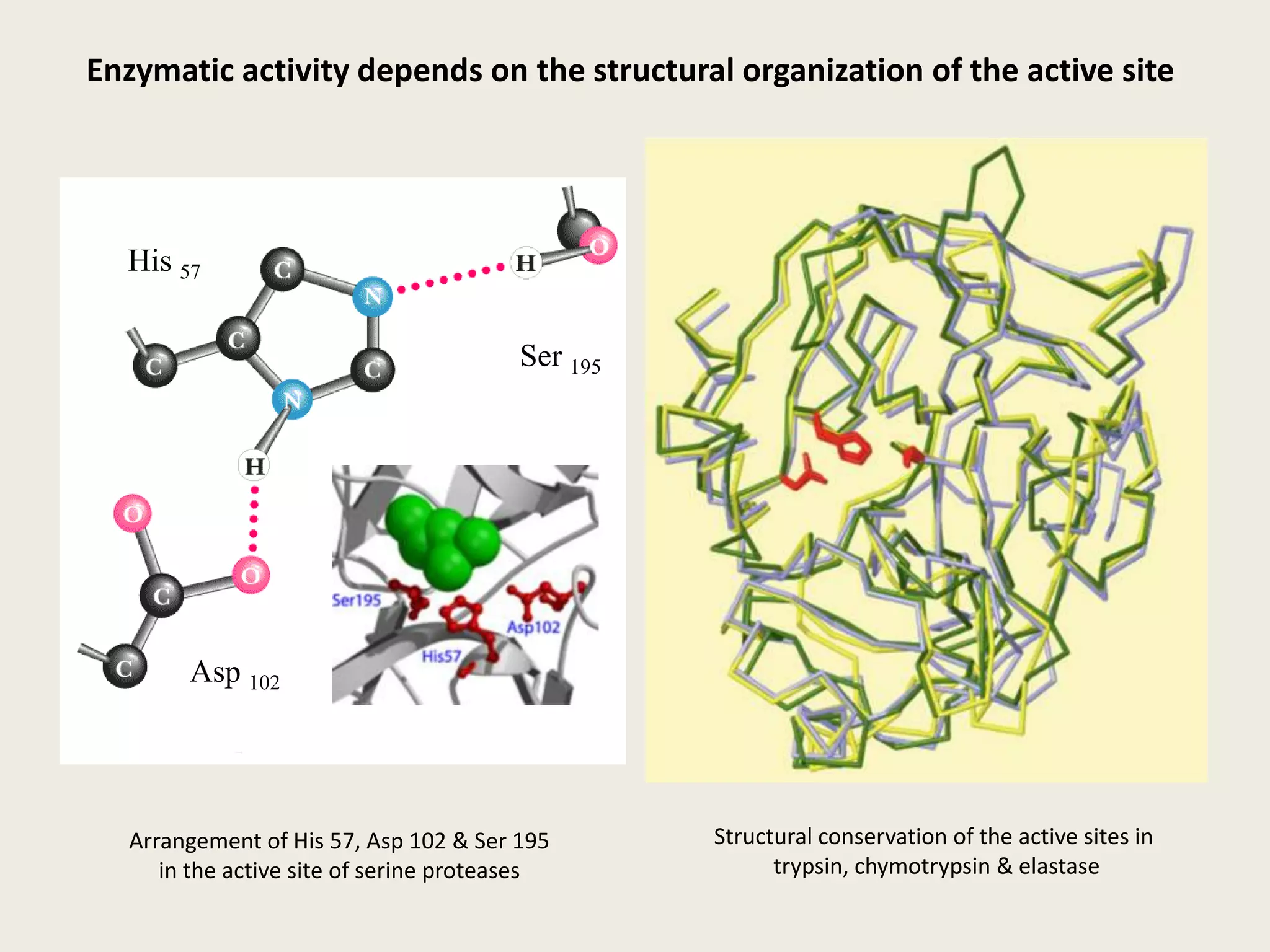
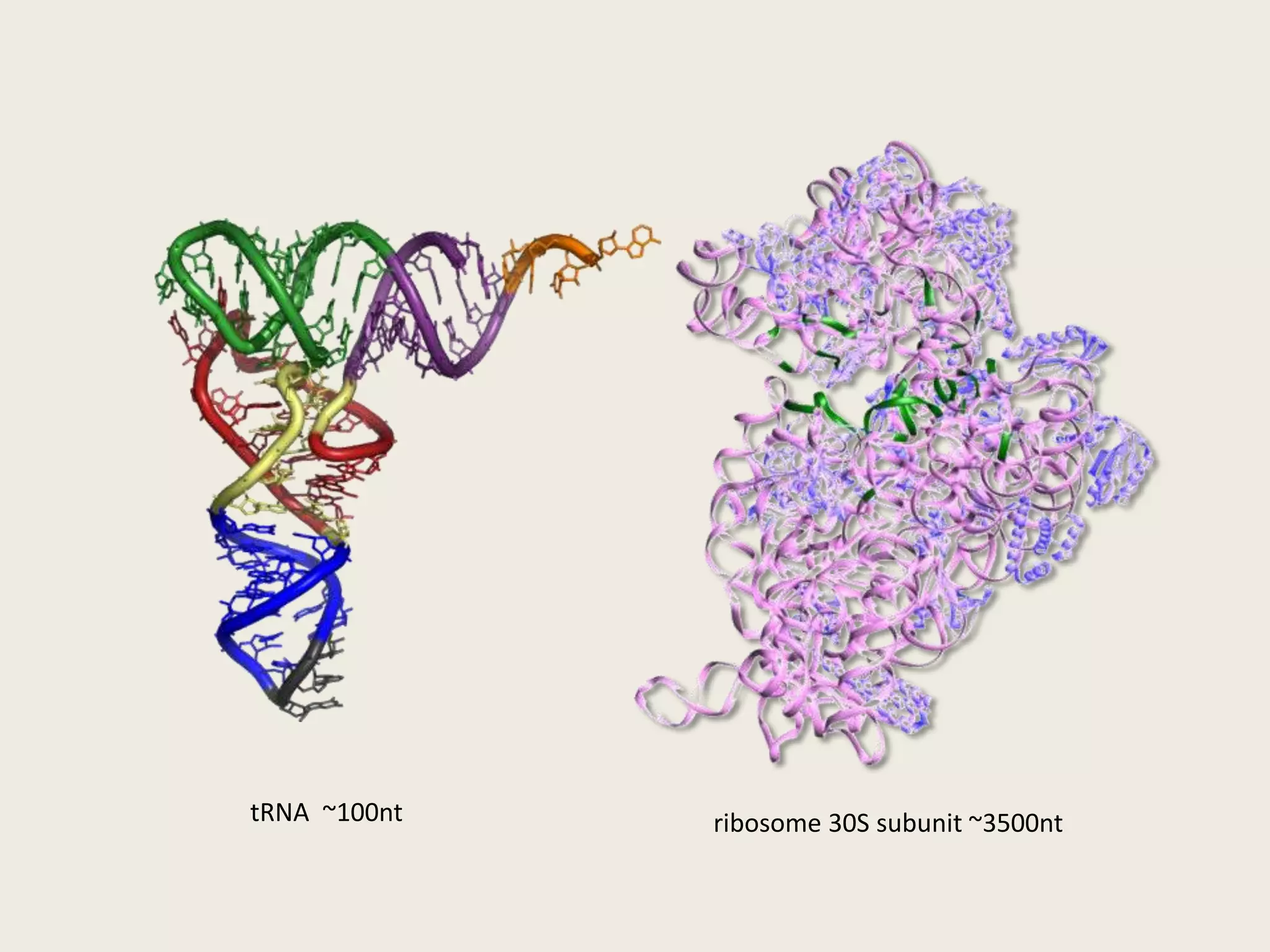
The document discusses computational studies related to pKa calculations of proteins and nucleic acids, focusing on the structural aspects that improve sequence alignments. It highlights the complexity of ribozymes, particularly in the context of the hepatitis delta virus (HDV), which relies on specific catalytic mechanisms. The findings emphasize the importance of structural geometry in understanding pKa shifts and the role of electrostatics in determining nucleotide interactions within RNA.