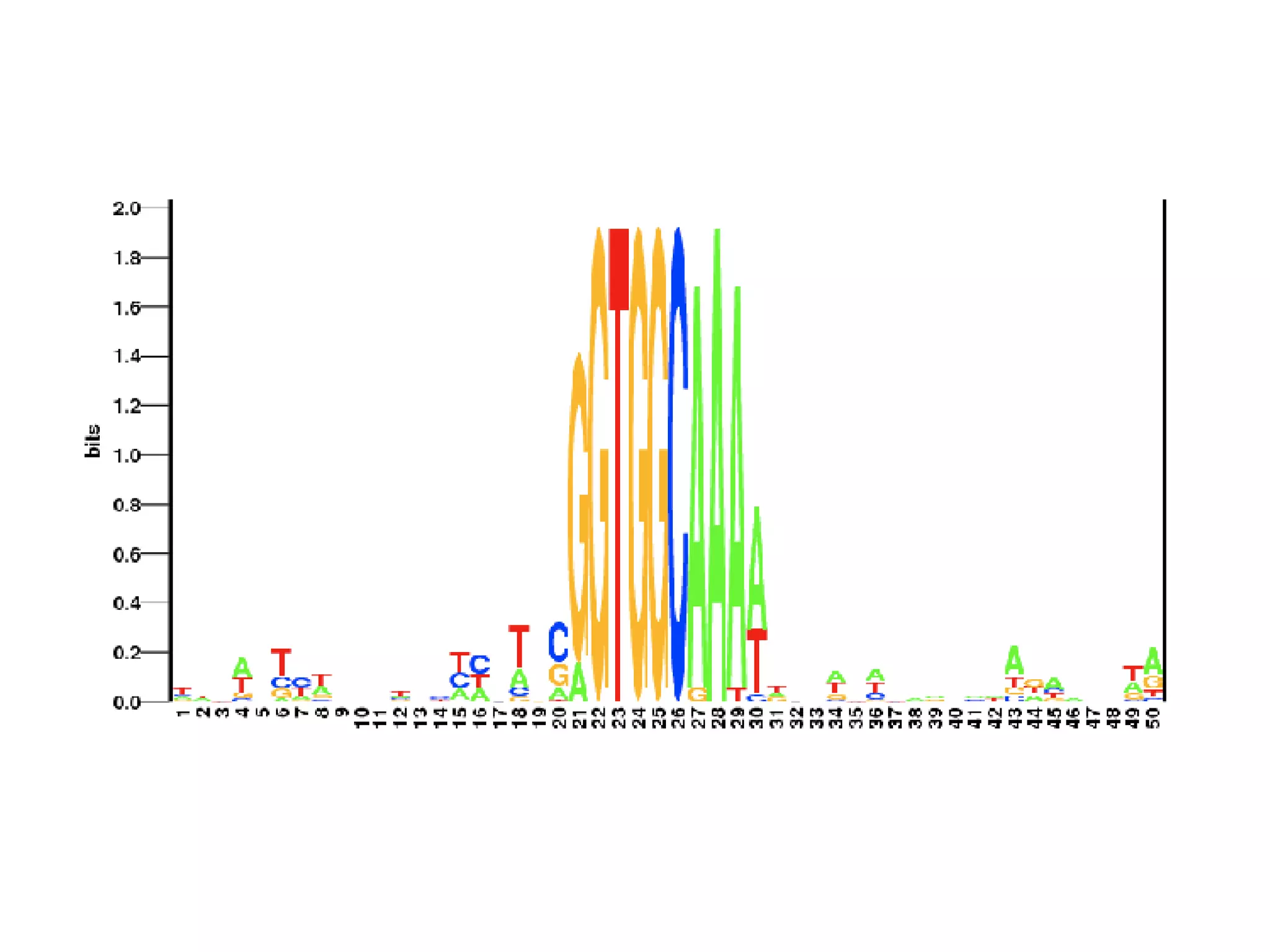
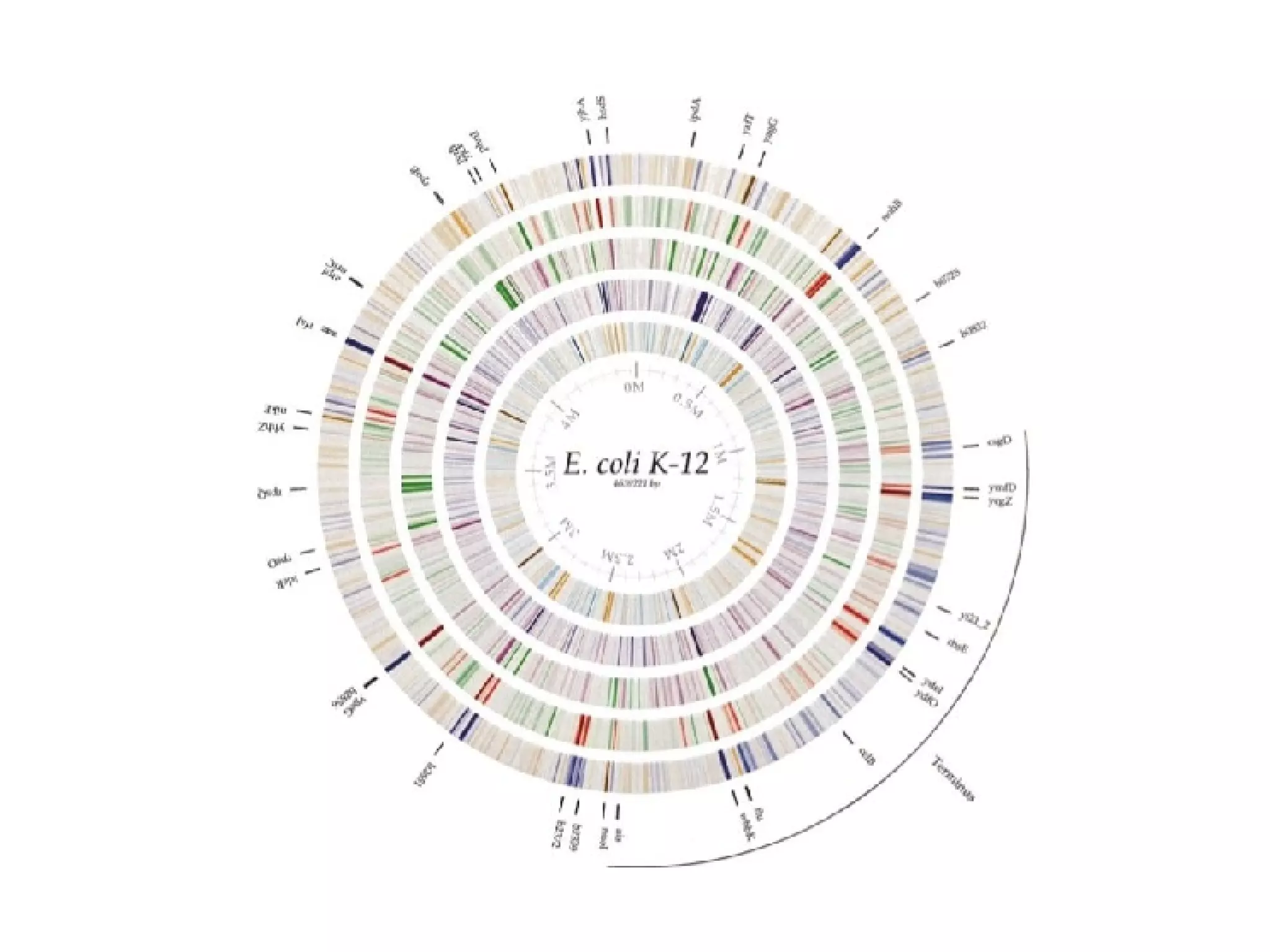
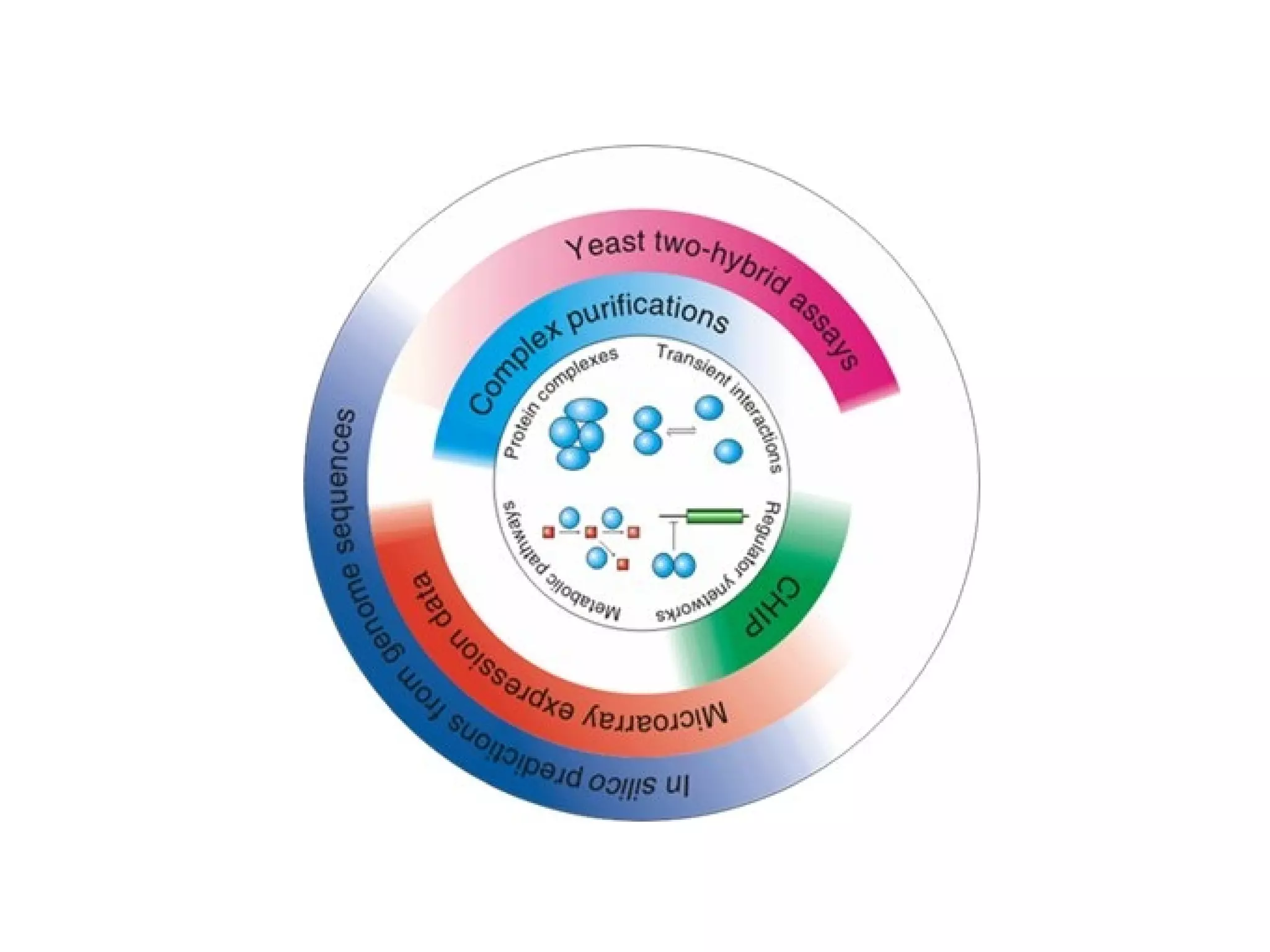
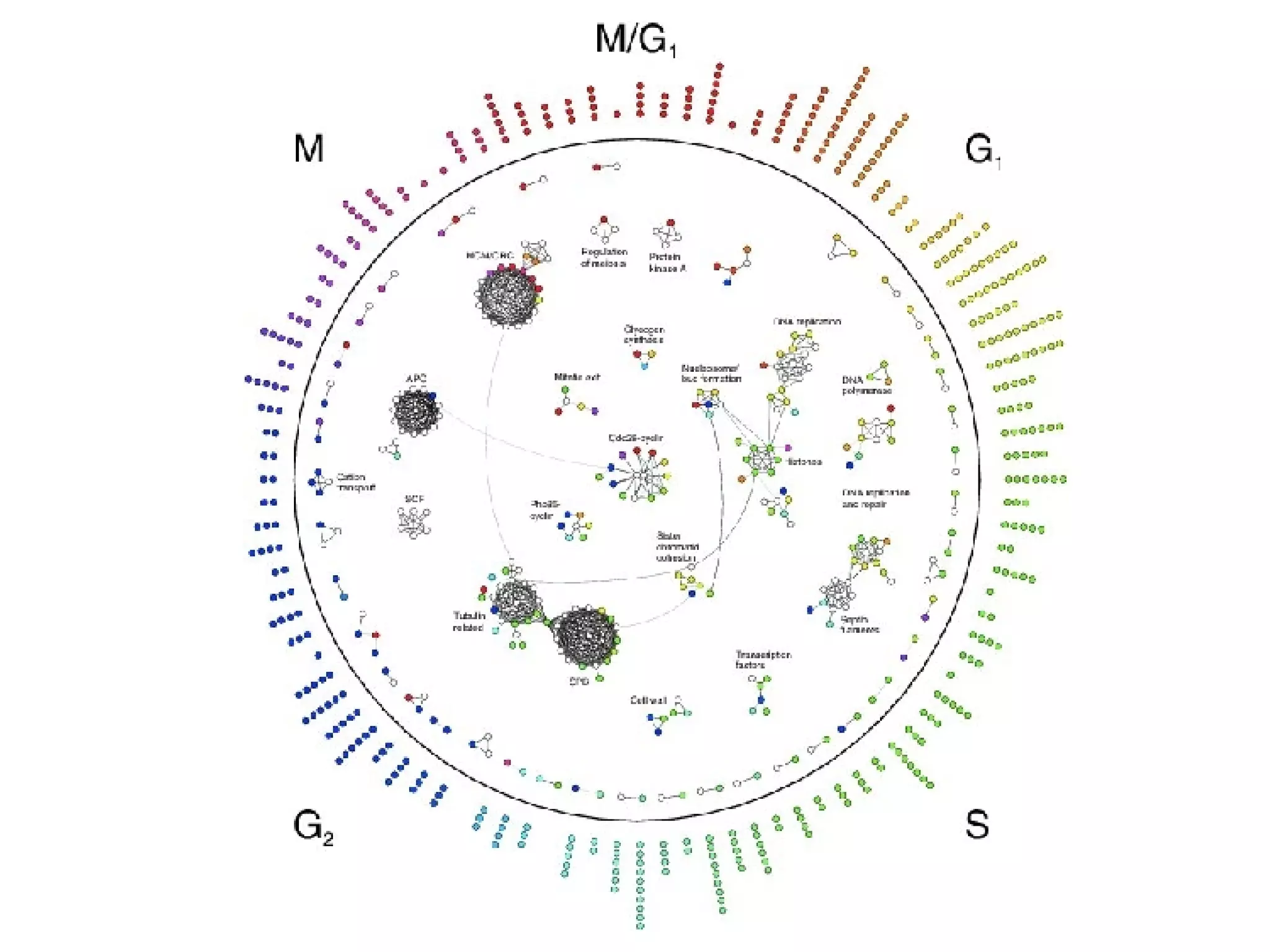
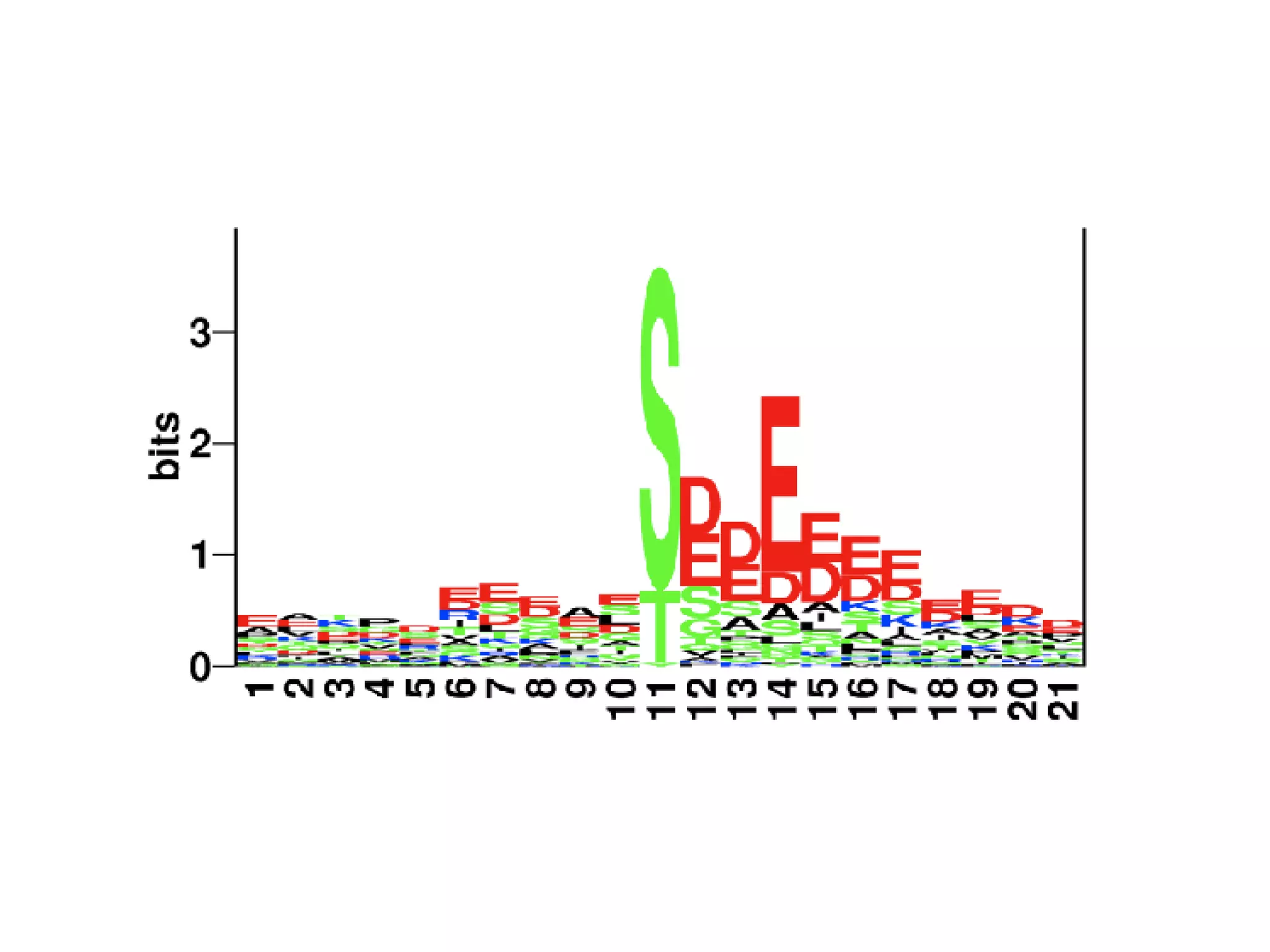
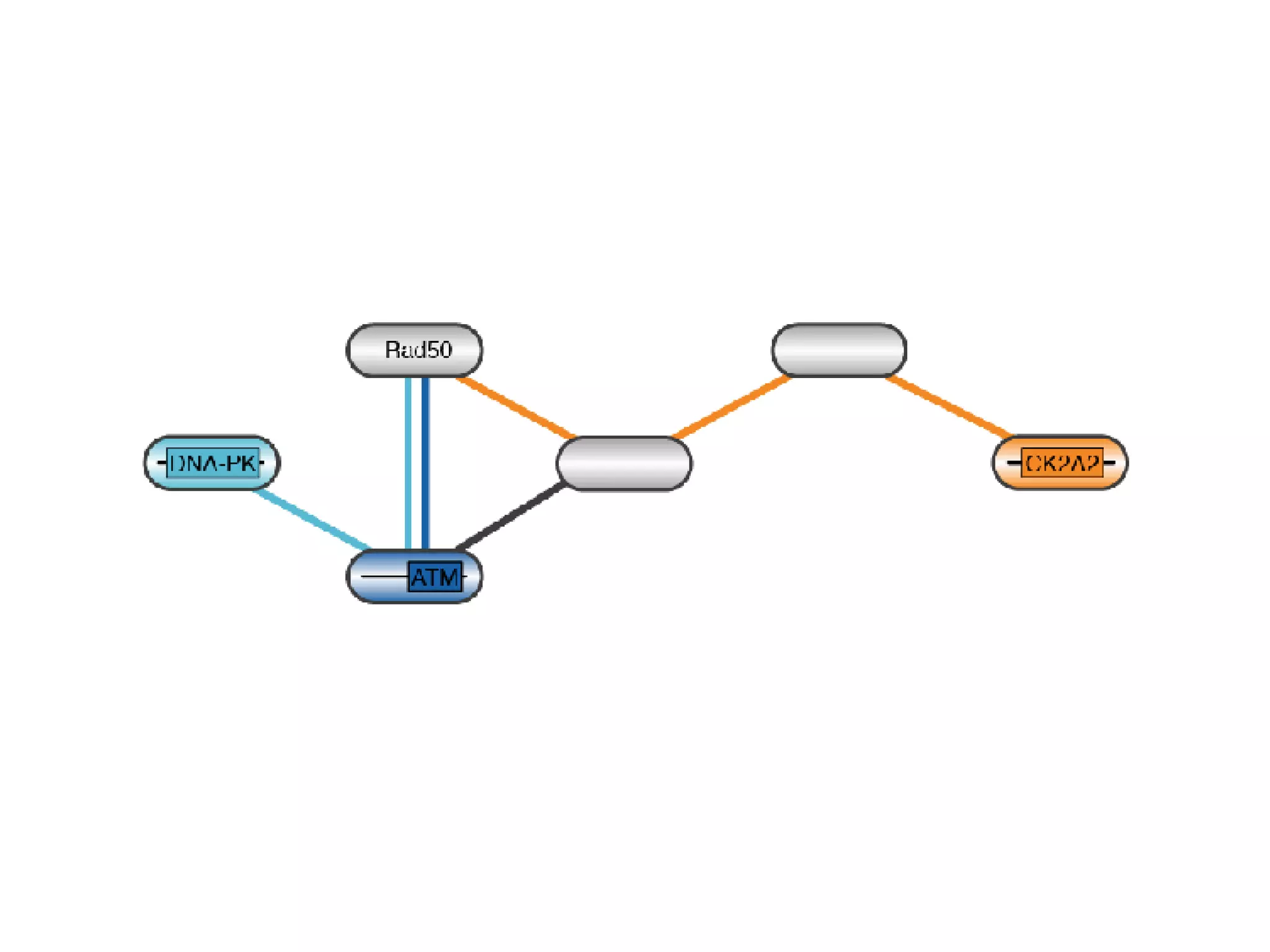
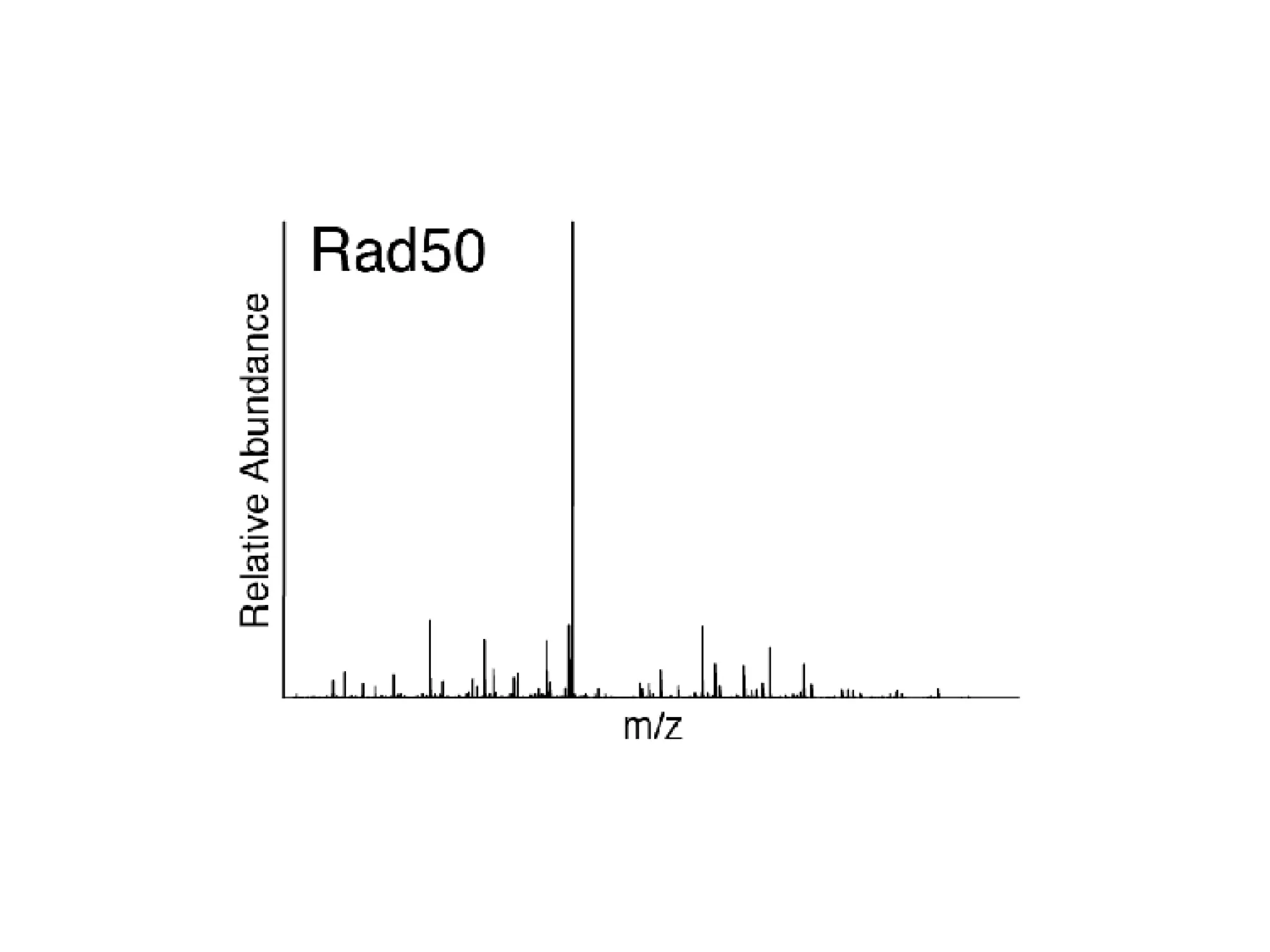
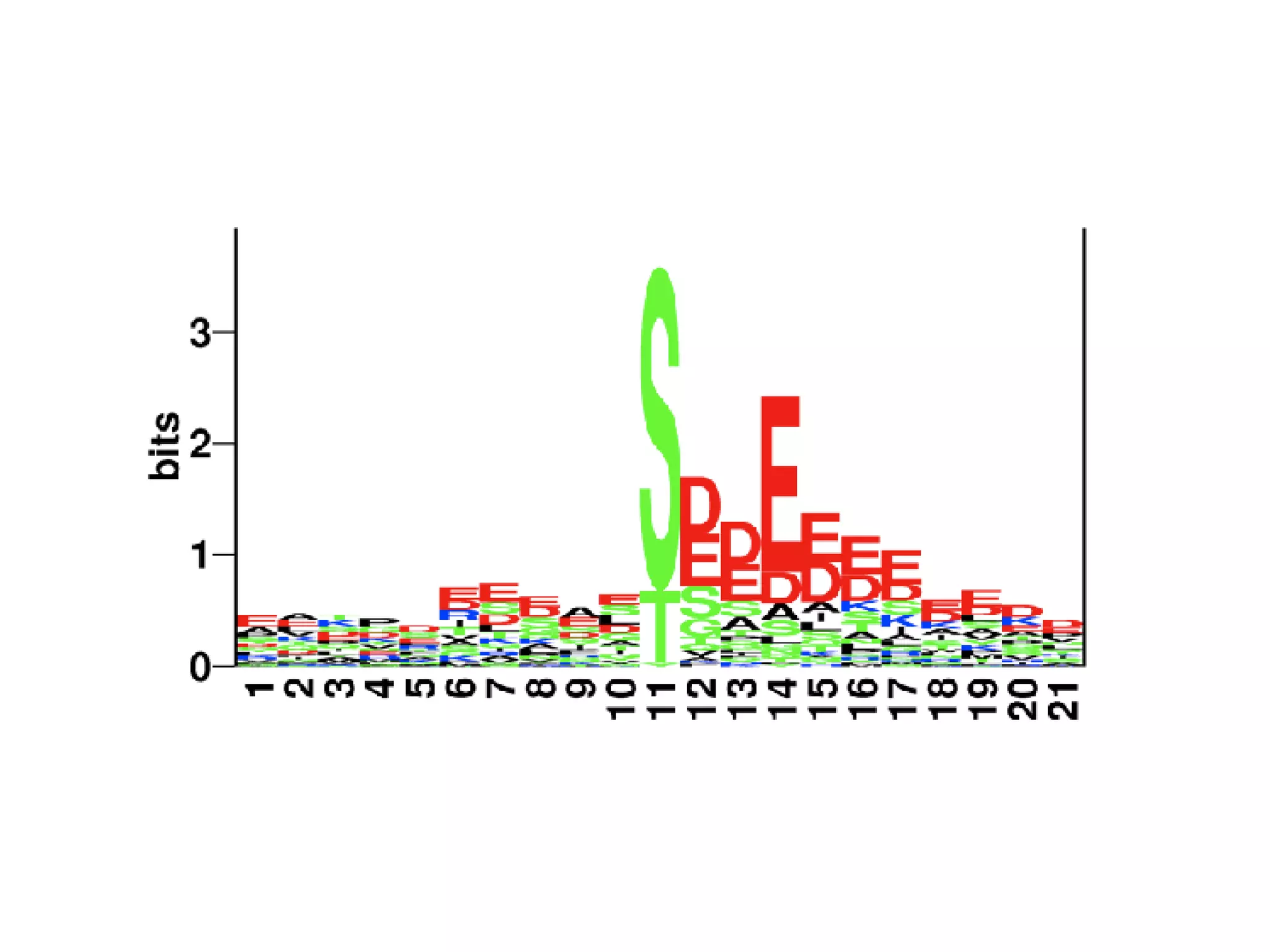
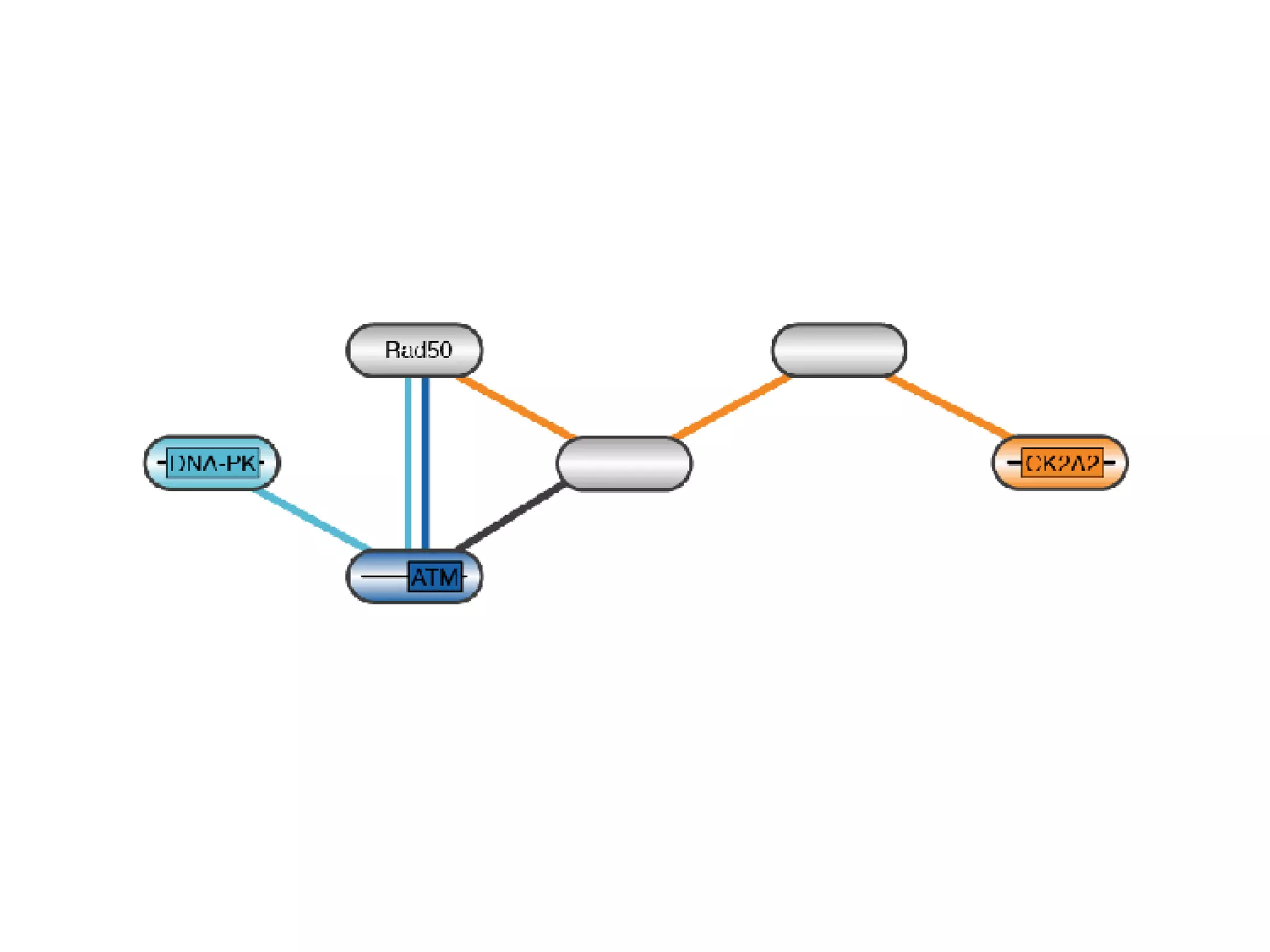
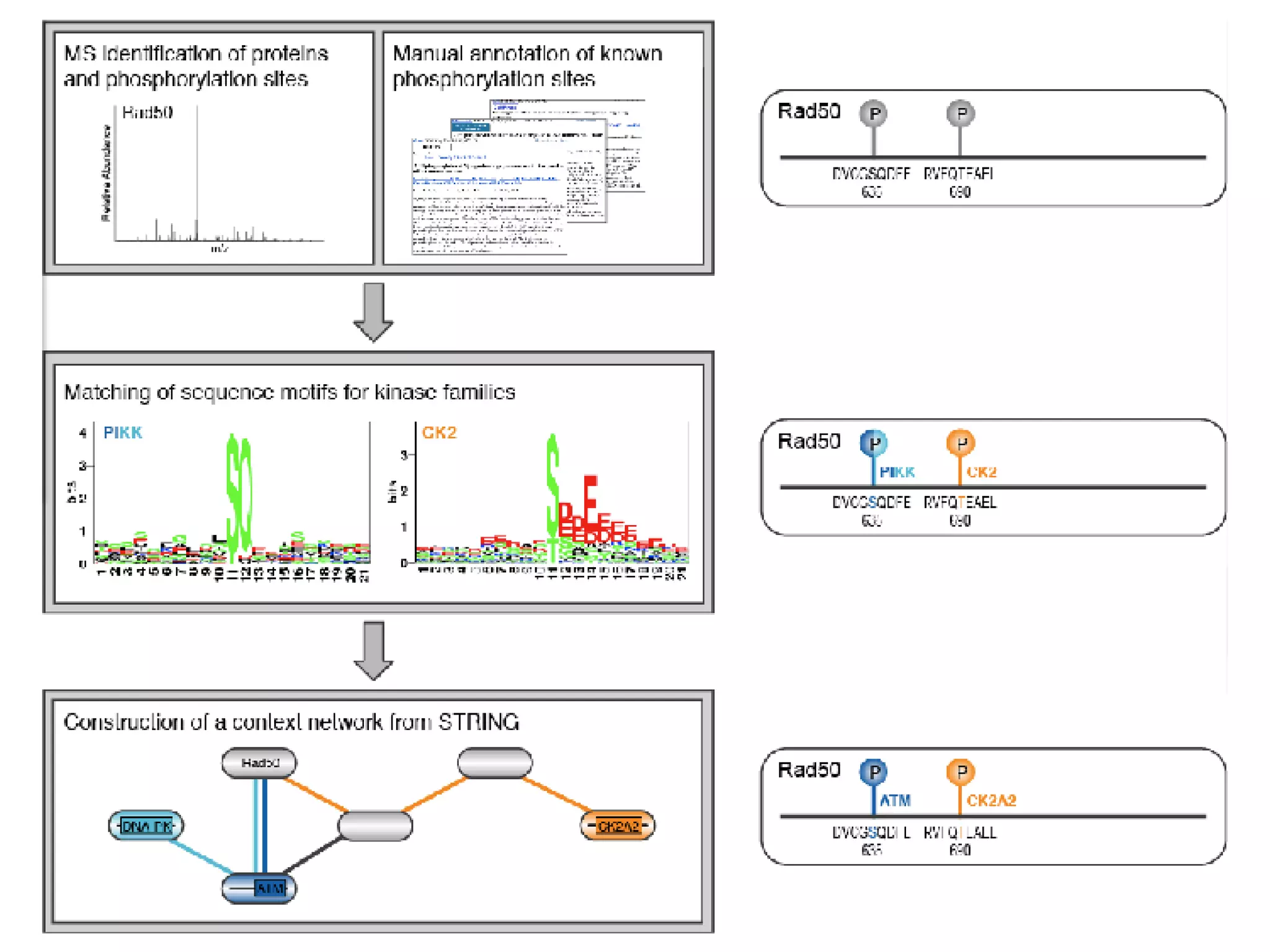
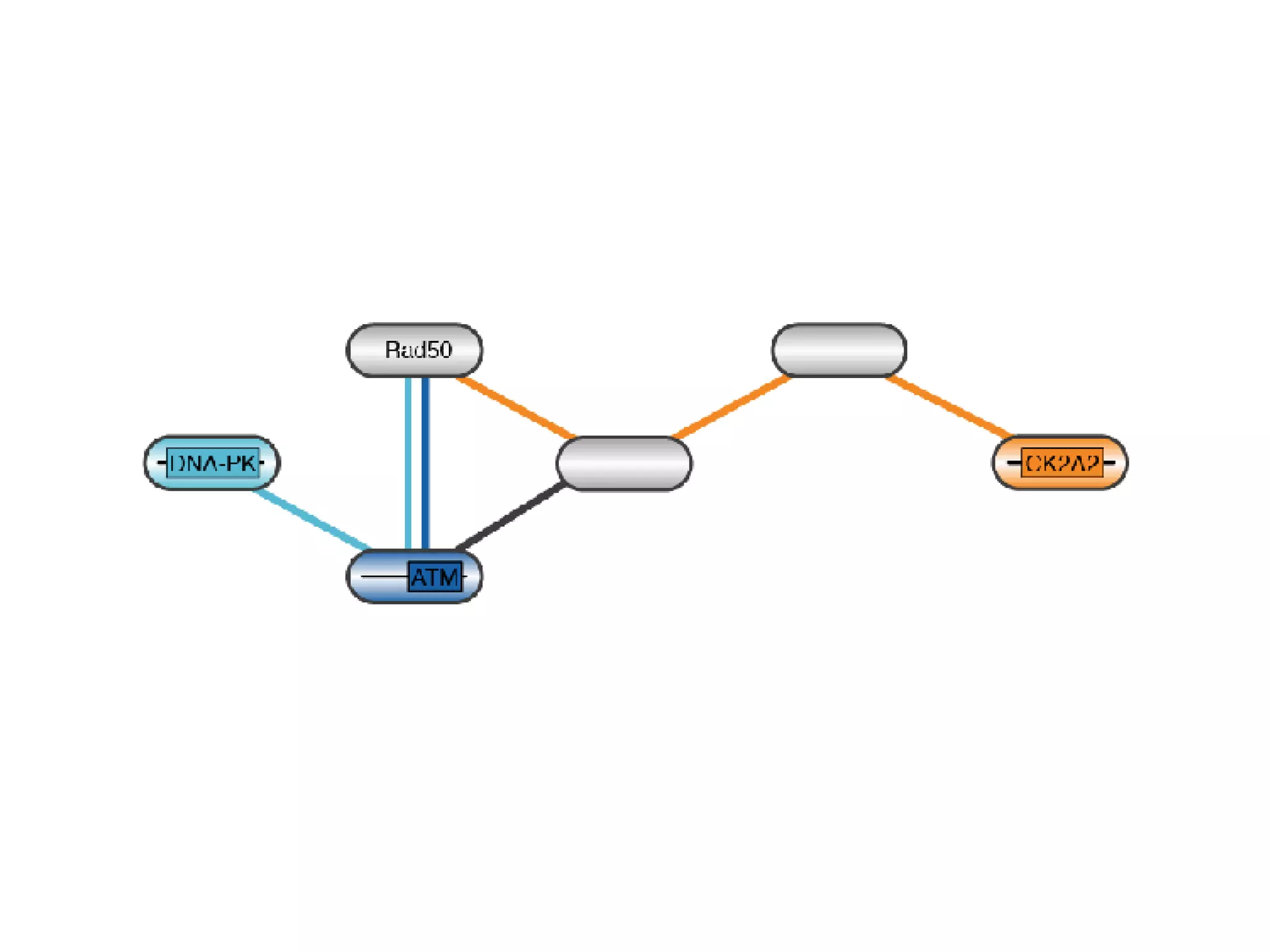
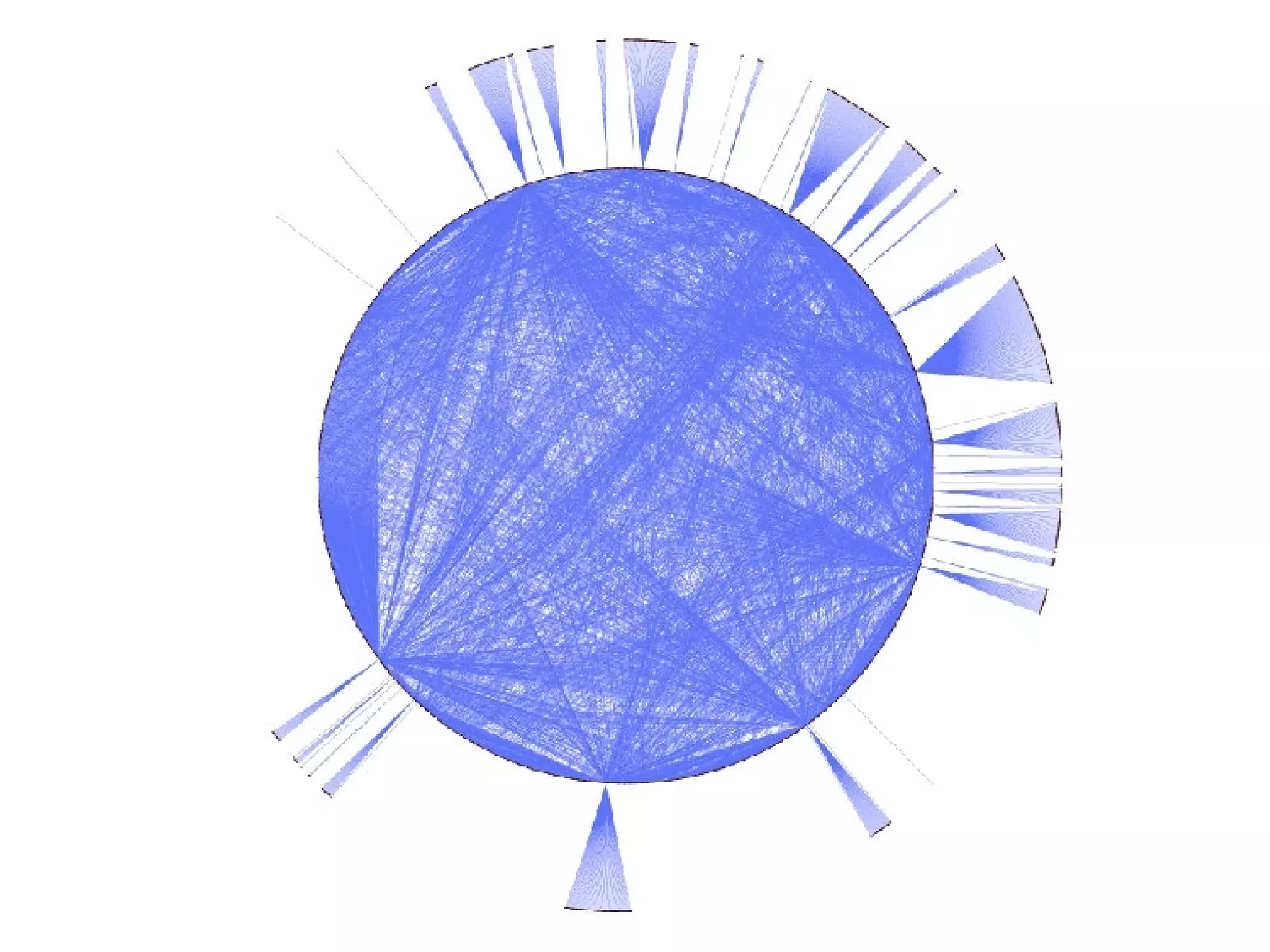
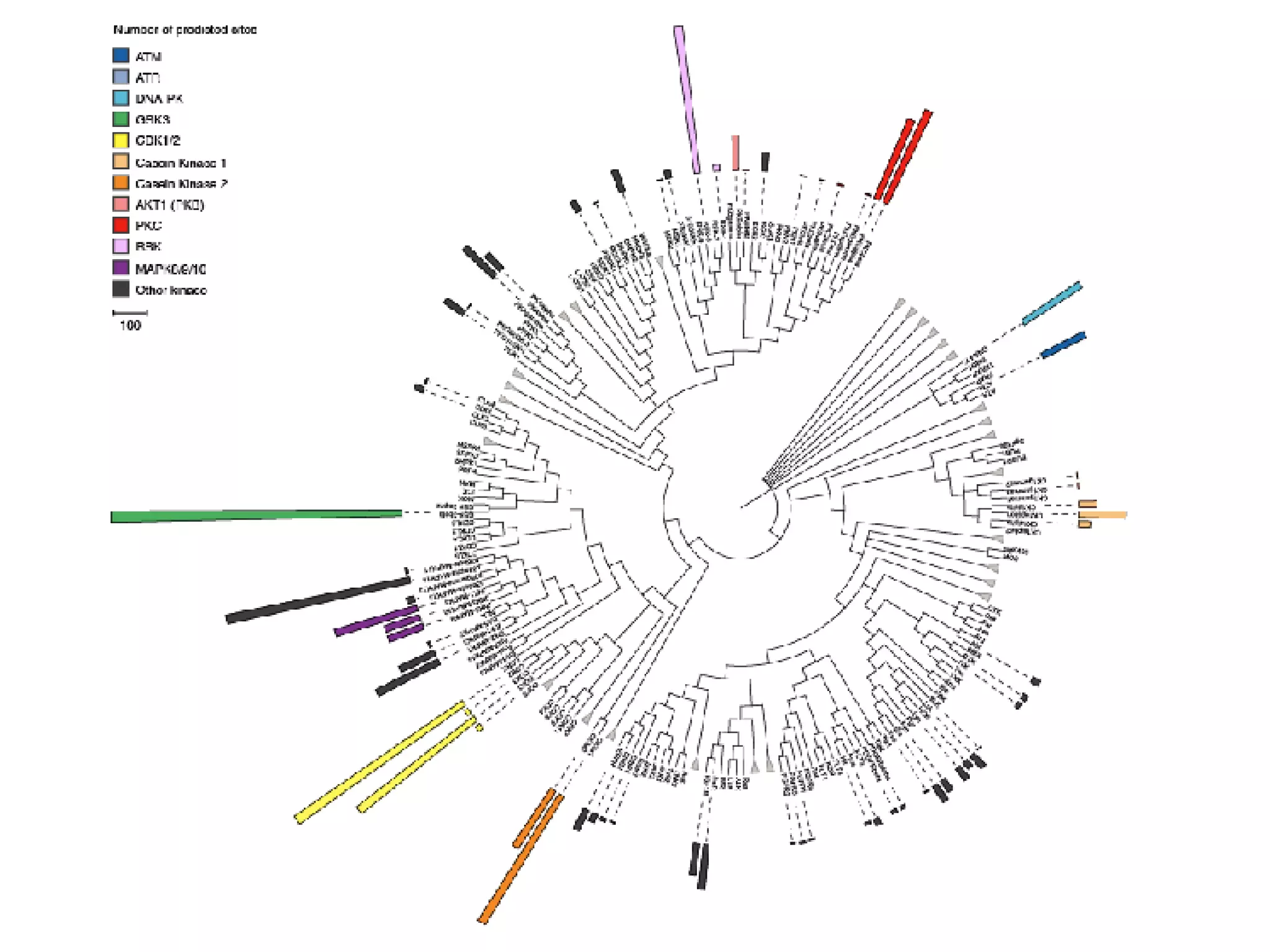
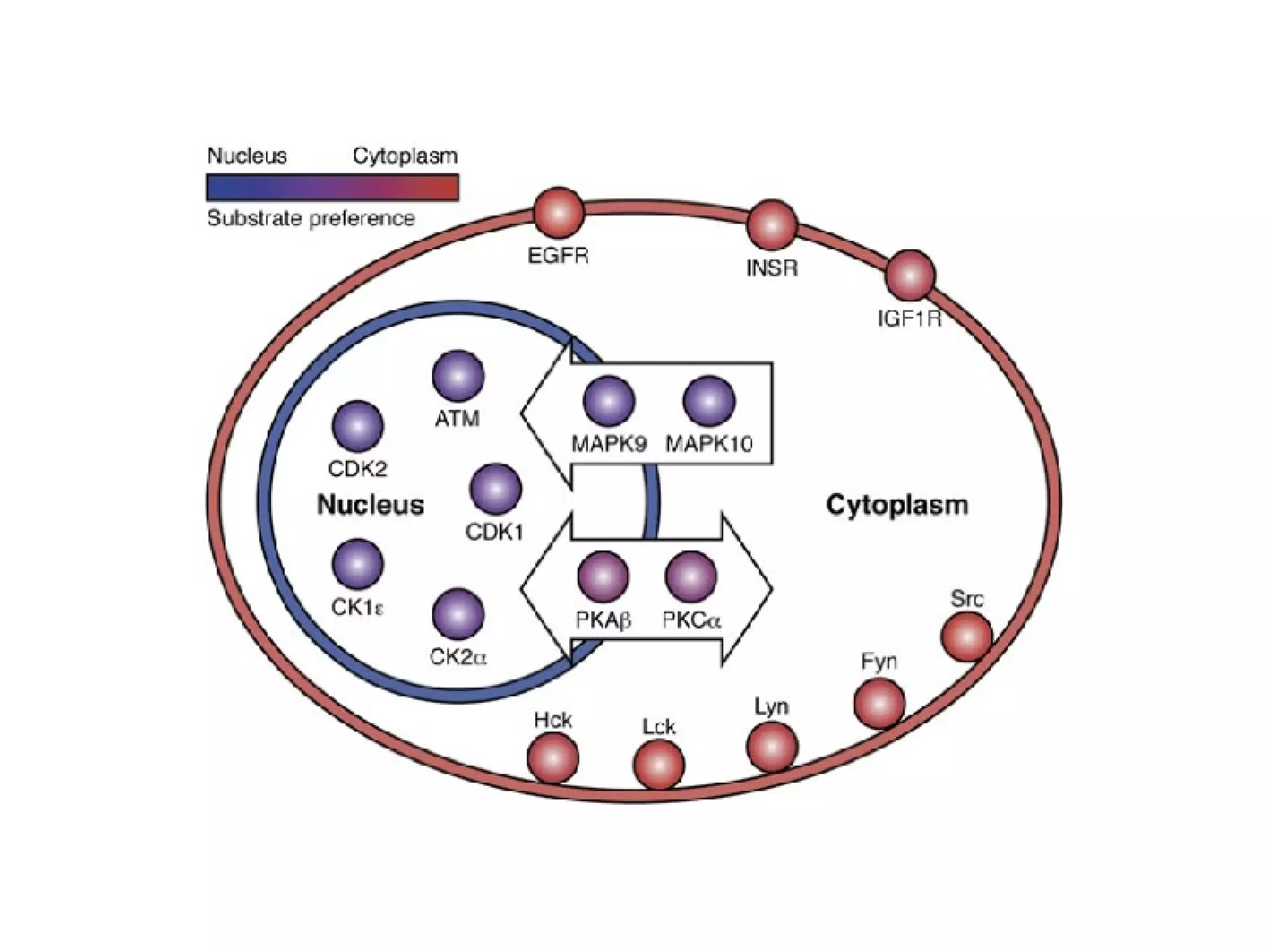
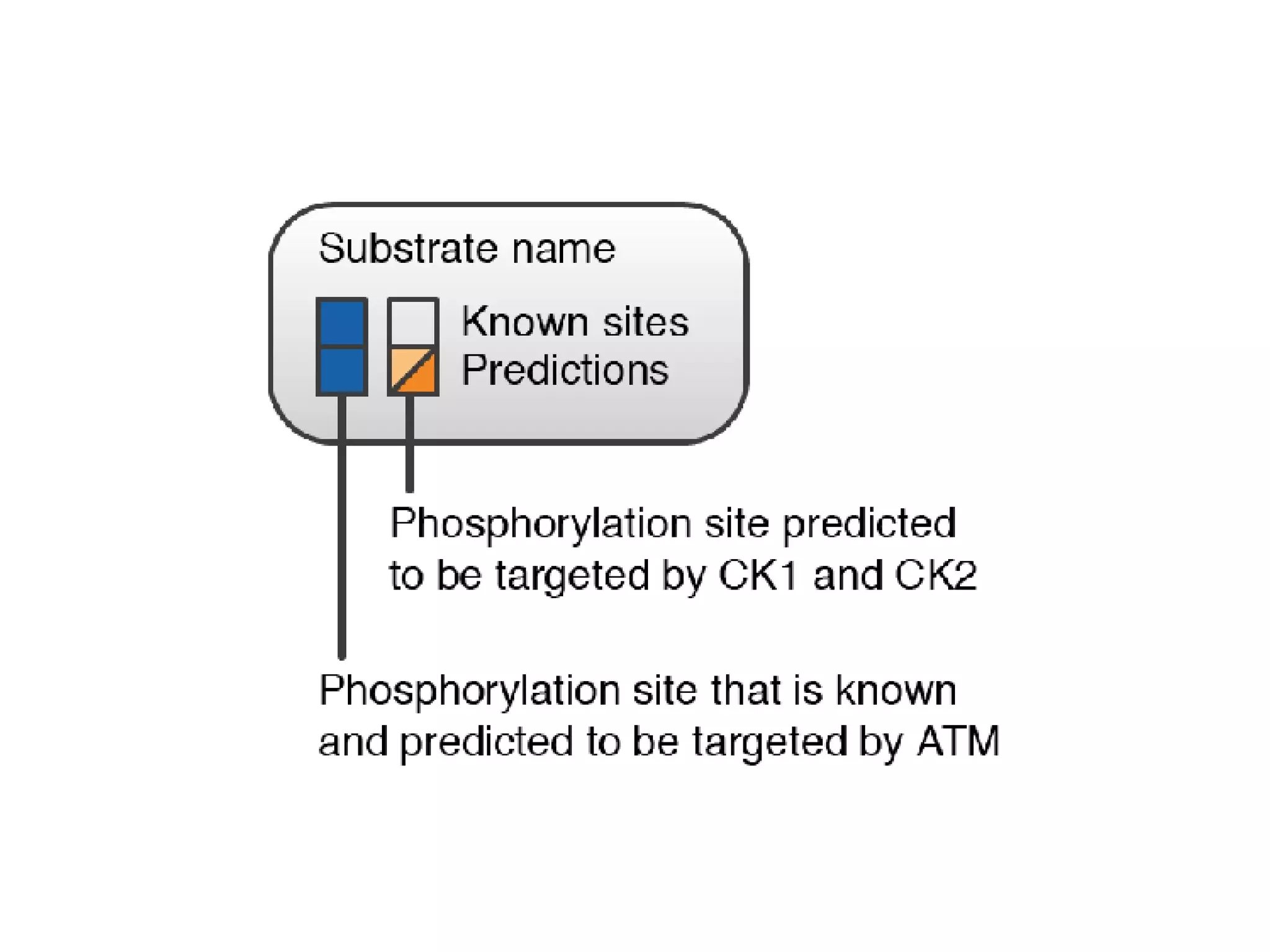
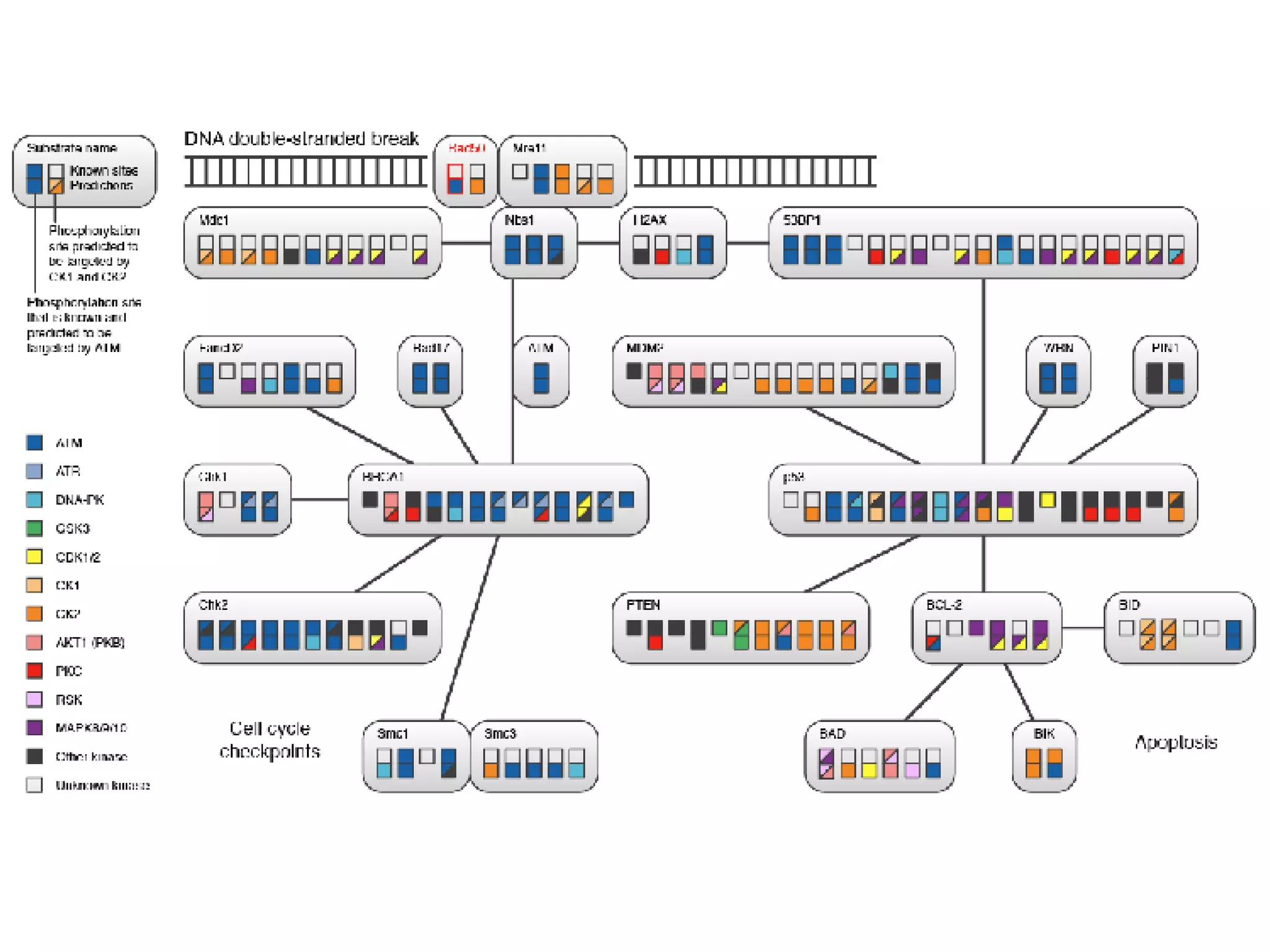
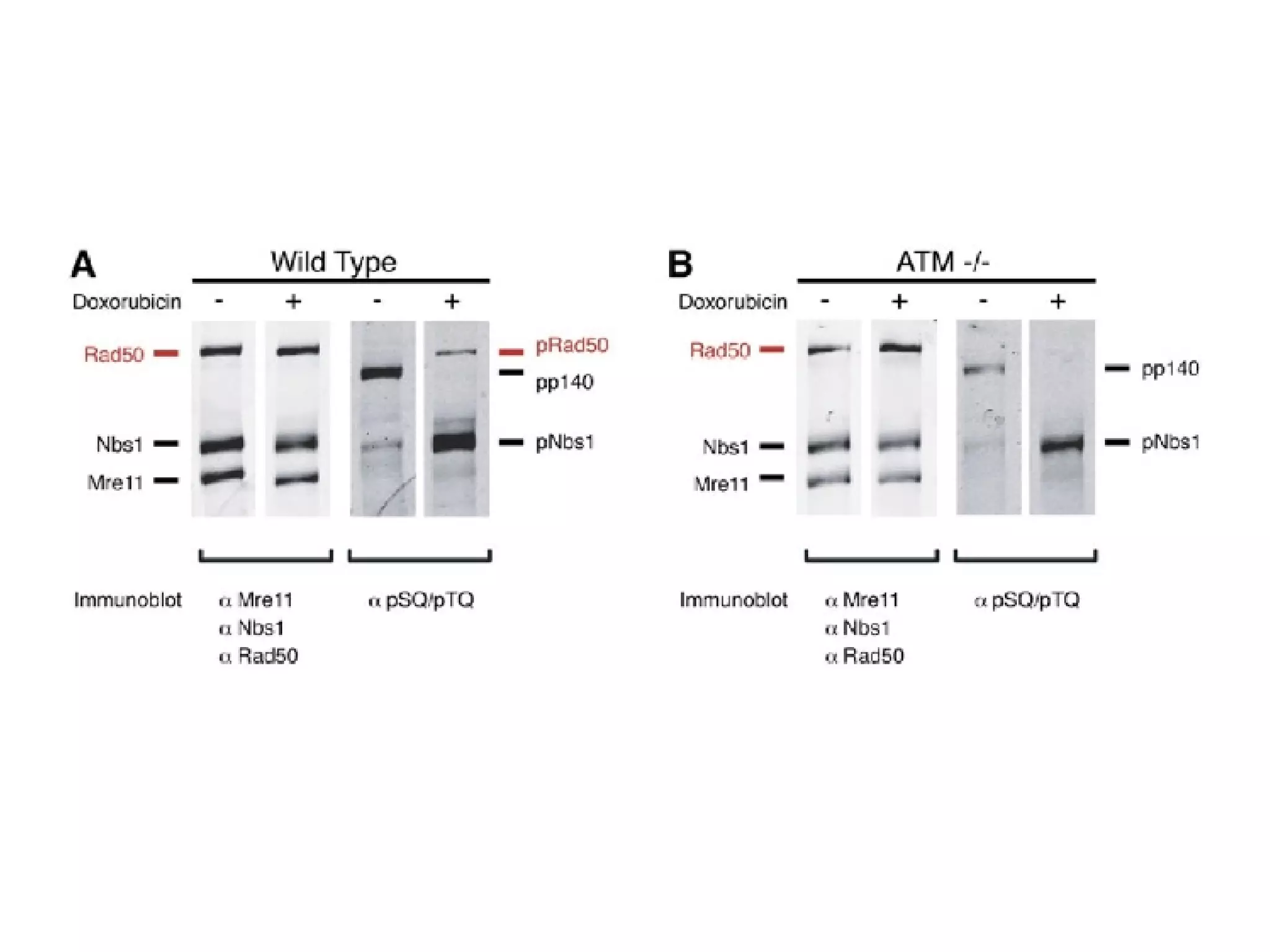
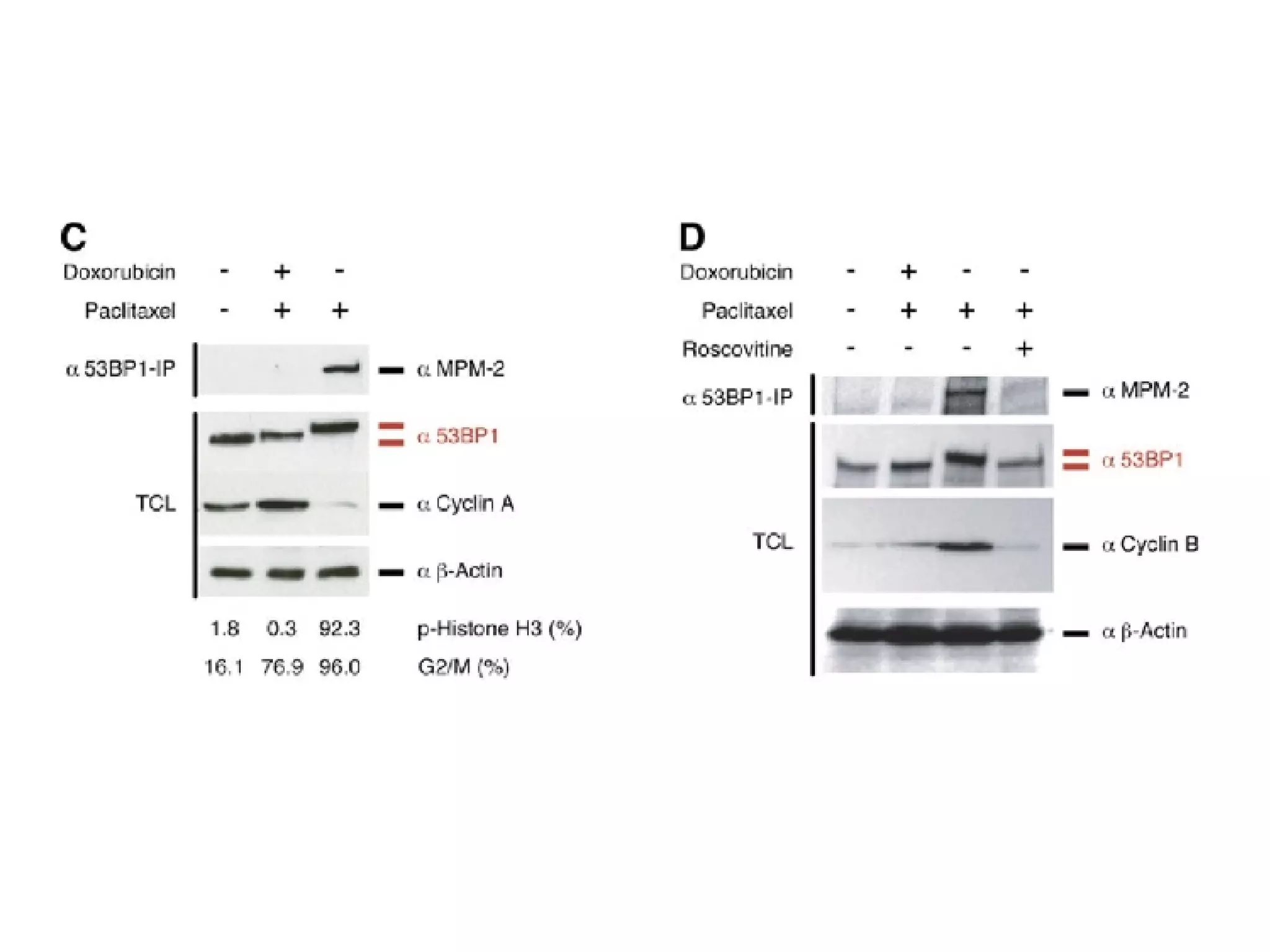
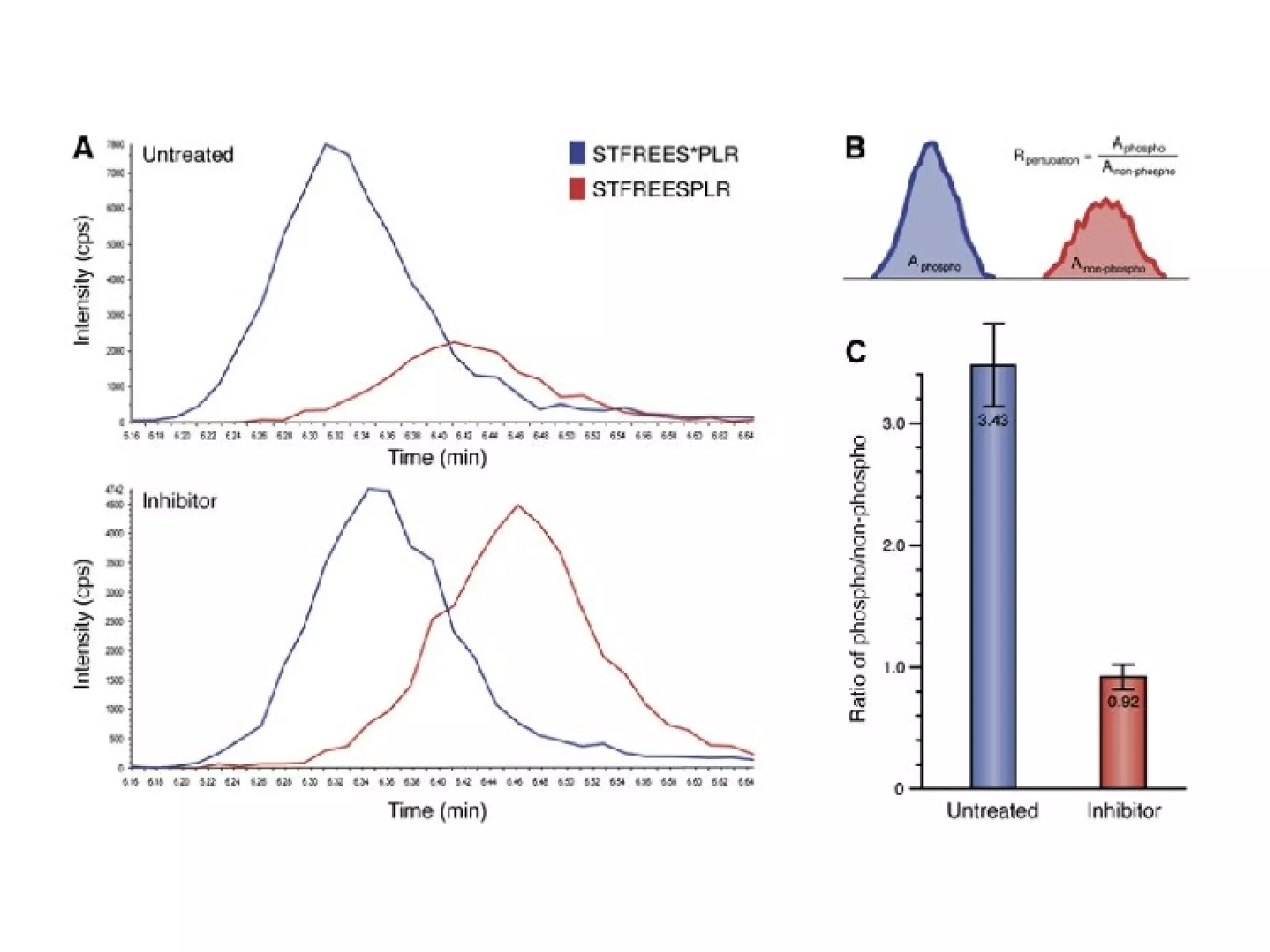
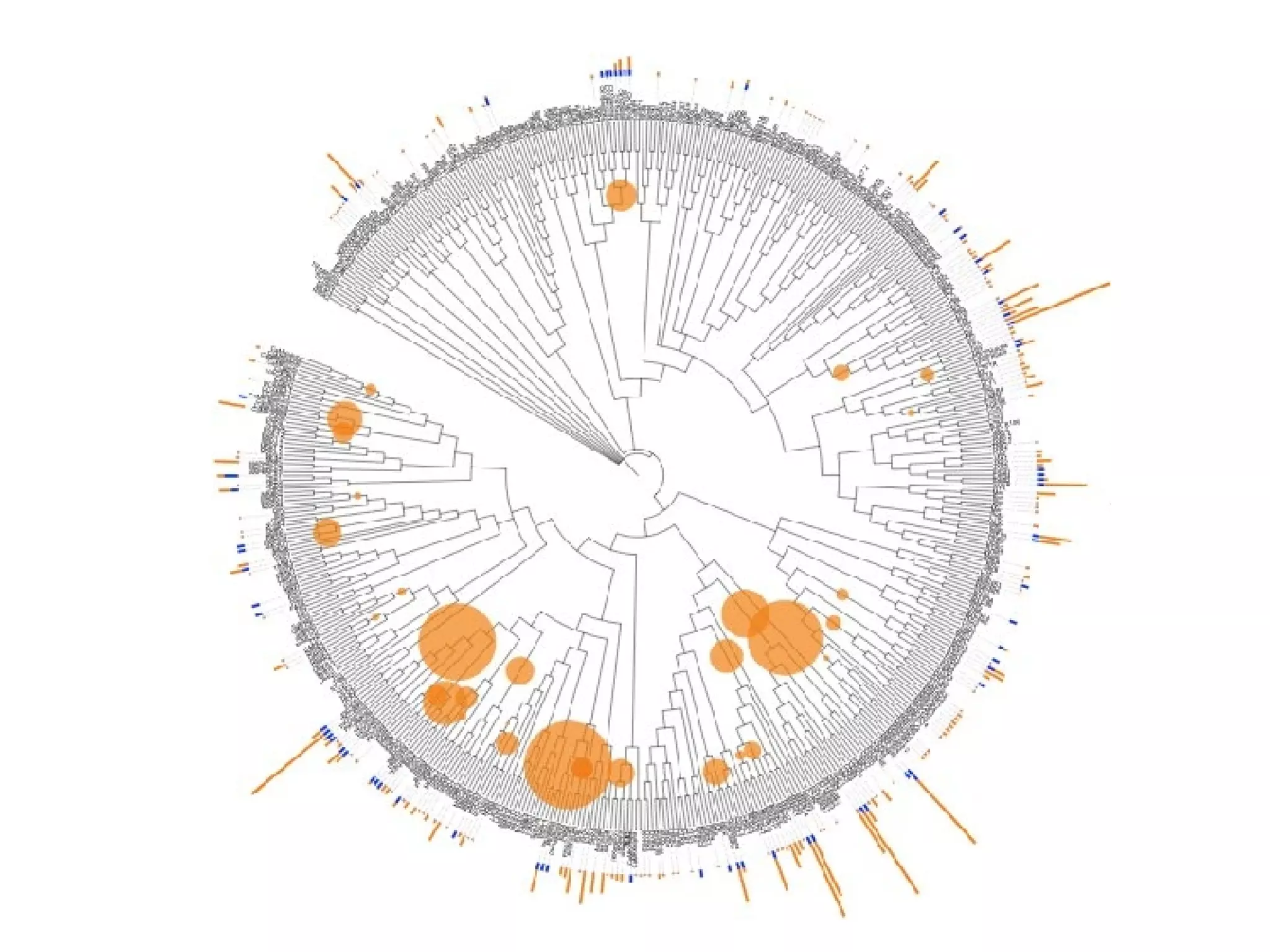
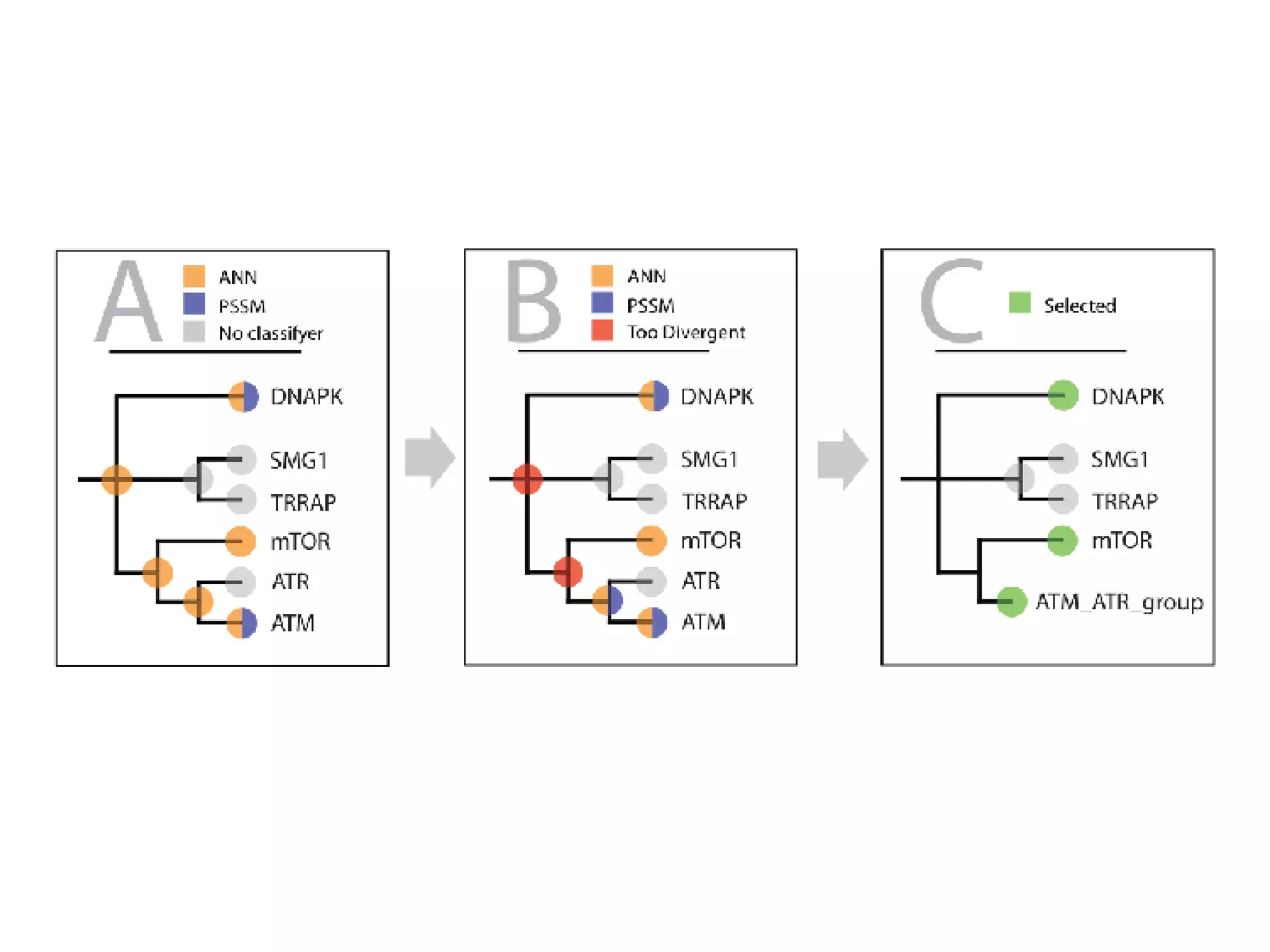
The document discusses the systematic discovery of phosphorylation networks by combining linear motifs and protein interactions, highlighting the role of kinases and the importance of context for understanding phosphorylation sites. It outlines methods such as mass spectrometry, gene integration, and various computational approaches for interaction prediction, resulting in thousands of predictions related to kinases and substrates. The future of this research includes improved data organization, automatic pipelines, and further exploration of signaling pathways.