Spago is an open-source machine learning library written in Go, designed to facilitate natural language processing tasks with a focus on simplicity and production readiness. It includes features like automatic differentiation, various neural network architectures, and optimizers, though currently lacks GPU support. The project aims to provide high-quality implementations for educational and practical production use, catering to developers interested in machine learning within the Go ecosystem.










![Building Blocks
Lightweight internal Machine Learning framework
1. mat package (pkg/mat)
a. 2D dense and sparse matrices (vectors and scalars as a subset of).
i. go test dense 80% statements
ii. go test sparse 77% statements
b. []float64 as data storage.
c. BLAS linear algebra asm/f64 (implementation by GONUM Authors).
d. sync.Pool for an efficient reuse of allocated memory.](https://image.slidesharecdn.com/spagogowayfest2020-201001111157/75/spaGO-A-self-contained-ML-NLP-library-in-GO-11-2048.jpg)



![Toy Example (c = a+b)
g := ag.NewGraph()
// create a new node of type variable with a scalar
a := g.NewVariable(mat.NewScalar(2.0), true)
// create another node of type variable with a scalar
b := g.NewVariable(mat.NewScalar(5.0), true)
// create an addition operator (the calculation is actually performed here)
c := g.Add(a, b)
// print the result
fmt.Printf("c = %vn", c.Value()) // c = [7]
g.Backward(c, ag.OutputGrad(mat.NewScalar(0.5)))
// print the gradients
fmt.Printf("ga = %vn", a.Grad()) // ga = [0.5]
fmt.Printf("gb = %vn", b.Grad()) // gb = [0.5]](https://image.slidesharecdn.com/spagogowayfest2020-201001111157/75/spaGO-A-self-contained-ML-NLP-library-in-GO-15-2048.jpg)
![Affine transformation
func Affine(g *ag.Graph, xs ...ag.Node) ag.Node {
y := g.Add(xs[0], g.Mul(xs[1], xs[2])) // y = b + Wx
for i := 3; i < len(xs)-1; i += 2 {
w, x := xs[i], xs[i+1]
if x != nil {
y = g.Add(y, g.Mul(w, x)) // y += Wx
}
}
return y
}
// y = b + W1x1 + W2x2 + ... + WnXn](https://image.slidesharecdn.com/spagogowayfest2020-201001111157/75/spaGO-A-self-contained-ML-NLP-library-in-GO-16-2048.jpg)
![Linear Model (pkg/ml/nn/linear)
var (
_ nn.Model = &Model{}
_ nn.Processor = &Processor{}
)
type Model struct {
W *nn.Param `type:"weights"`
B *nn.Param `type:"biases"`
}
// New returns a new Linear model
func New(in, out int) *Model {
return &Model{
W: nn.NewParam(mat.NewEmptyDense(out, in)),
B: nn.NewParam(mat.NewEmptyVecDense(out)),
}
}
type Processor struct {
nn.BaseProcessor
w, b ag.Node
}
// NewProc returns Linear processor operating to the graph
func (m *Model) NewProc(g *ag.Graph) nn.Processor {
return &Processor{
BaseProcessor: nn.BaseProcessor{
Model: m,
Mode: nn.Training,
Graph: g,
},
// insert the weights and biases into the graph
w: g.NewWrap(m.W),
b: g.NewWrap(m.B),
}
}
// Forward computes y[i] = w (dot) x[i] + b
func (p *Processor) Forward(xs ...ag.Node) []ag.Node
ys := make([]ag.Node, len(xs))
for i, x := range xs {
ys[i] = nn.Affine(p.Graph, p.b, p.w, x)
}
return ys
}](https://image.slidesharecdn.com/spagogowayfest2020-201001111157/75/spaGO-A-self-contained-ML-NLP-library-in-GO-17-2048.jpg)
![Linear Model (pkg/ml/nn/linear)
var (
_ nn.Model = &Model{}
_ nn.Processor = &Processor{}
)
type Model struct {
W *nn.Param `type:"weights"`
B *nn.Param `type:"biases"`
}
// New returns a new Linear model
func New(in, out int) *Model {
return &Model{
W: nn.NewParam(mat.NewEmptyDense(out, in)),
B: nn.NewParam(mat.NewEmptyVecDense(out)),
}
}
type Processor struct {
nn.BaseProcessor
w, b ag.Node
}
// NewProc returns Linear processor operating to the graph
func (m *Model) NewProc(g *ag.Graph) nn.Processor {
return &Processor{
BaseProcessor: nn.BaseProcessor{
Model: m,
Mode: nn.Training,
Graph: g,
},
// insert the weights and biases into the graph
w: g.NewWrap(m.W),
b: g.NewWrap(m.B),
}
}
// Forward computes y[i] = w (dot) x[i] + b
func (p *Processor) Forward(xs ...ag.Node) []ag.Node
ys := make([]ag.Node, len(xs))
for i, x := range xs {
ys[i] = nn.Affine(p.Graph, p.b, p.w, x)
}
return ys
}](https://image.slidesharecdn.com/spagogowayfest2020-201001111157/75/spaGO-A-self-contained-ML-NLP-library-in-GO-18-2048.jpg)
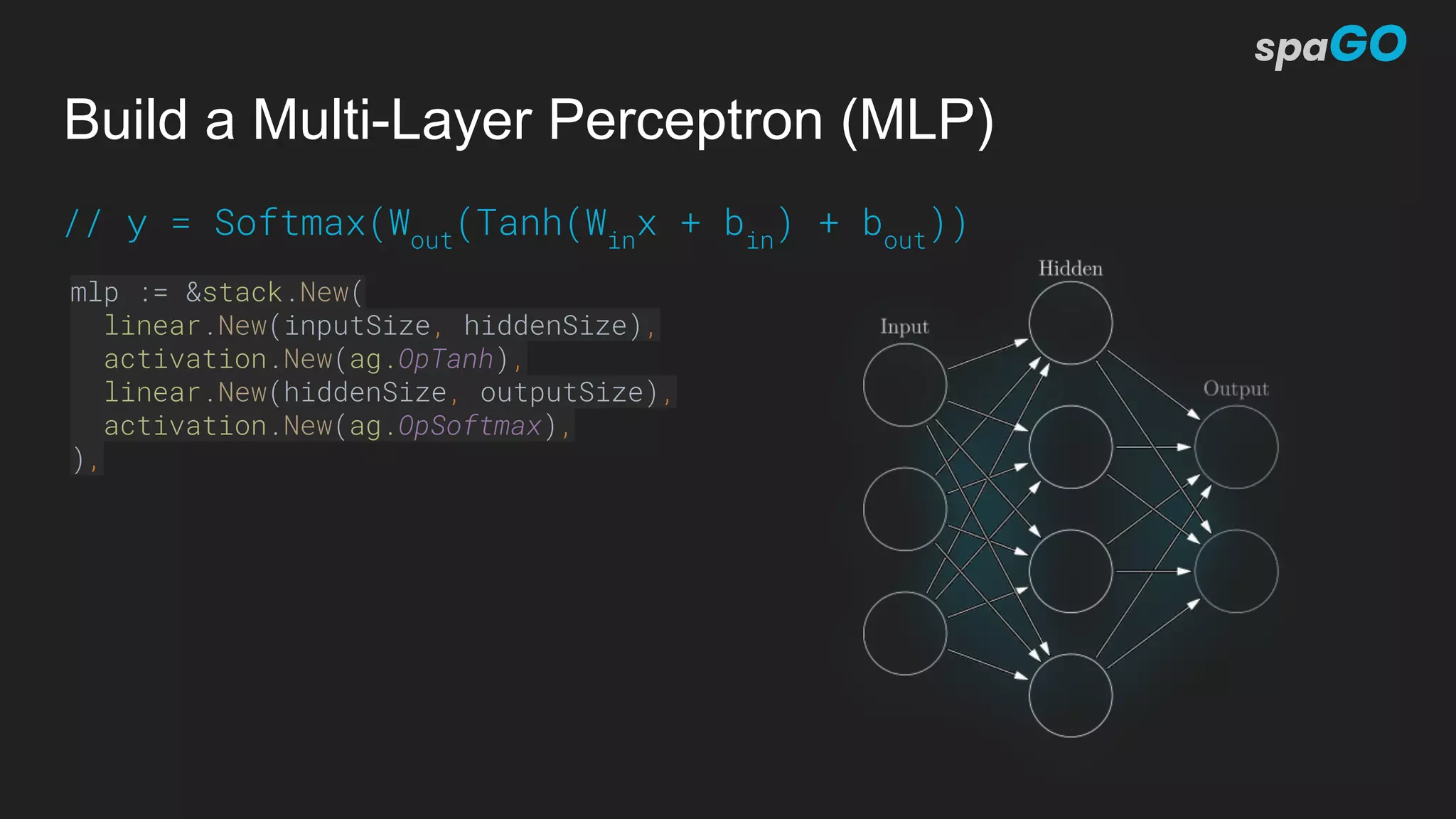


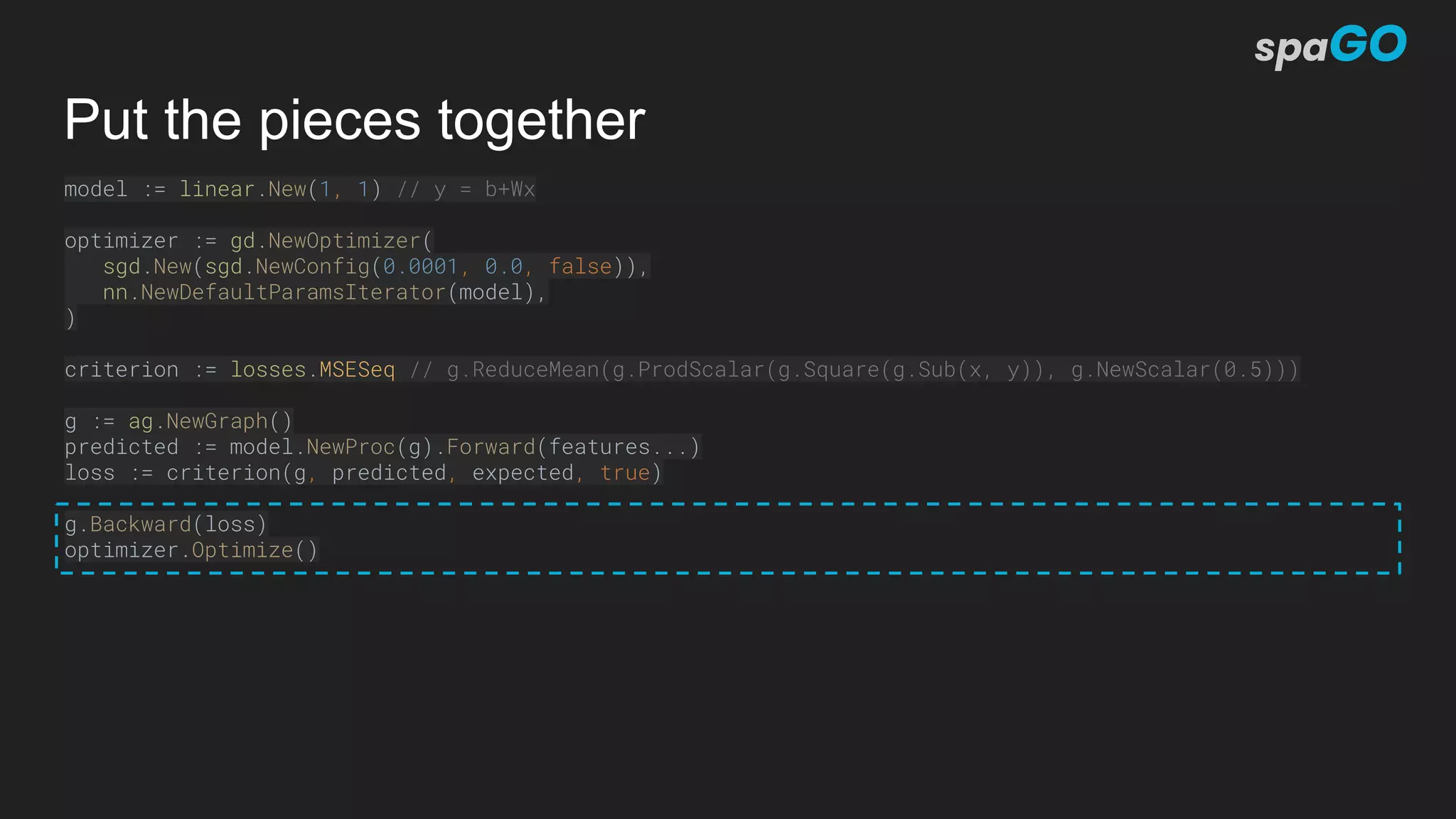
![NLP: Sequence Labeler (Flair Architecture)
flair := &sequencelabeler.Model{
Labels: config.Labels,
TaggerLayer: &birnncrf.Model{
BiRNN: birnn.New(
lstm.New(...),
lstm.New(...),
birnn.Concat,
),
Scorer: linear.New(...),
CRF: crf.New(...),
},
EmbeddingsLayer: &stackedembeddings.Model{
WordsEncoders: []nn.Model{
embeddings.New(...), // glovo
contextualstringembeddings.New(
charlm.New(...), // left to right
charlm.New(...), // right to left
contextualstringembeddings.Concat,
),
},
ProjectionLayer: linear.New(...),
},
}](https://image.slidesharecdn.com/spagogowayfest2020-201001111157/75/spaGO-A-self-contained-ML-NLP-library-in-GO-25-2048.jpg)
![NLP: BERT Layer
type Layer struct {
MultiHeadAttention *multiheadattention.Model
NormAttention *layernorm.Model
FFN *stack.Model
NormFFN *layernorm.Model
}
func (m *Layer) NewProc(g *ag.Graph) nn.Processor {
return &Processor{
BaseProcessor: nn.BaseProcessor{Model: m, Mode: nn.Training, Graph: g},
MultiHeadAttention: m.MultiHeadAttention.NewProc(g).(*multiheadattention.Processor),
NormAttention: m.NormAttention.NewProc(g).(*layernorm.Processor),
FFN: m.FFN.NewProc(g).(*stack.Processor),
NormFFN: m.NormFFN.NewProc(g).(*layernorm.Processor),
}
}
func (p *Processor) Forward(xs ...ag.Node) []ag.Node {
subLayer1 := rc.PostNorm(p.Graph, p.MultiHeadAttention.Forward, p.NormAttention.Forward, xs...)
subLayer2 := rc.PostNorm(p.Graph, p.FFN.Forward, p.NormFFN.Forward, subLayer1...)
return subLayer2
}](https://image.slidesharecdn.com/spagogowayfest2020-201001111157/75/spaGO-A-self-contained-ML-NLP-library-in-GO-26-2048.jpg)
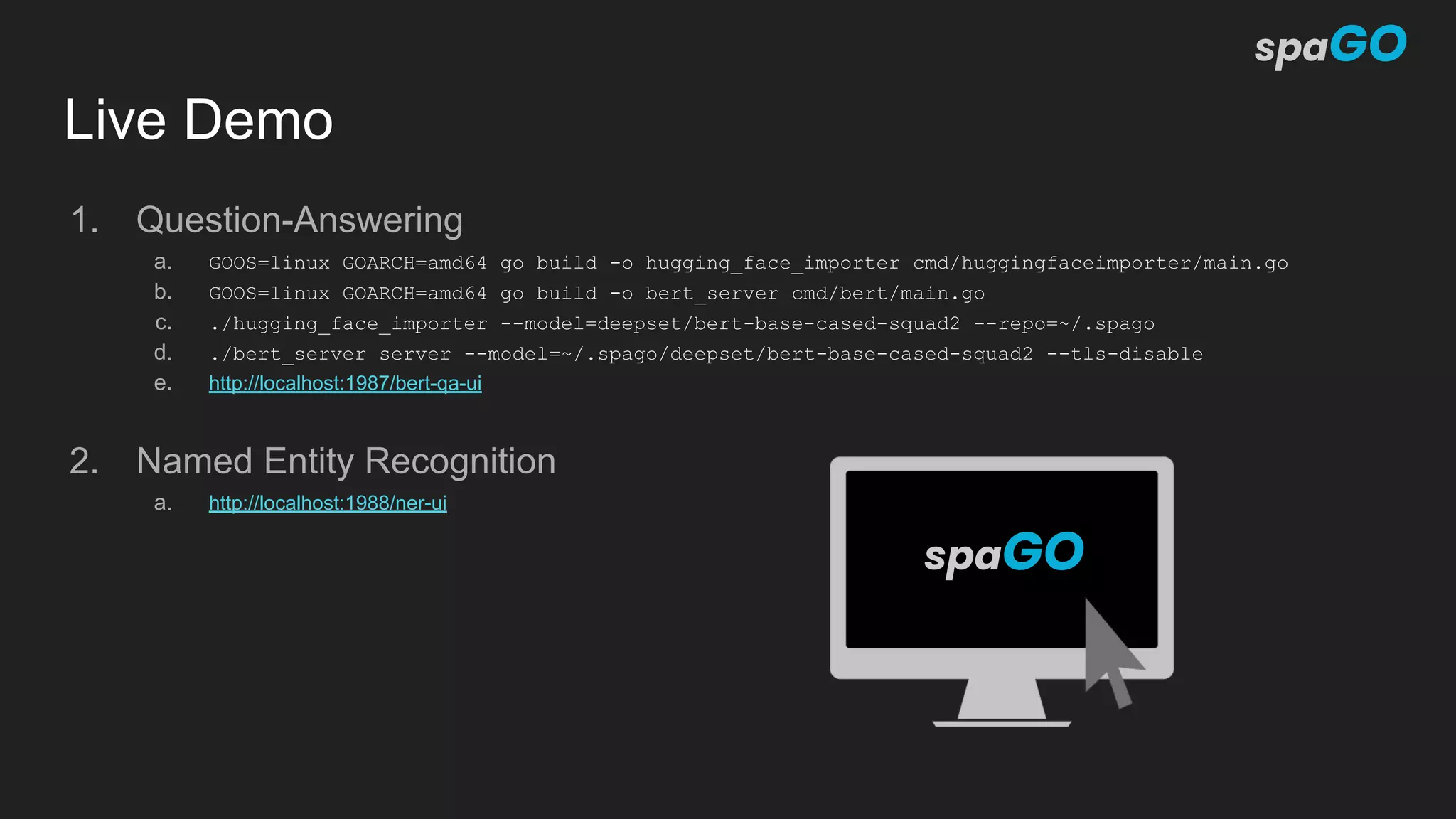

![Current status (v 0.0.z)
{
“Stars”: >600,
“Forks”: 23,
“Pull Requests”: 19,
“Issues”: 17,
“Main Contributors”: [
“M. Grella”, “E. Brambilla”, ”S. Cangialosi”, “M. Nicola”,
“E. McClure”, “J. Viana”,
],
“Building”: “passing”
“Codecov”: “60%”,
}](https://image.slidesharecdn.com/spagogowayfest2020-201001111157/75/spaGO-A-self-contained-ML-NLP-library-in-GO-29-2048.jpg)