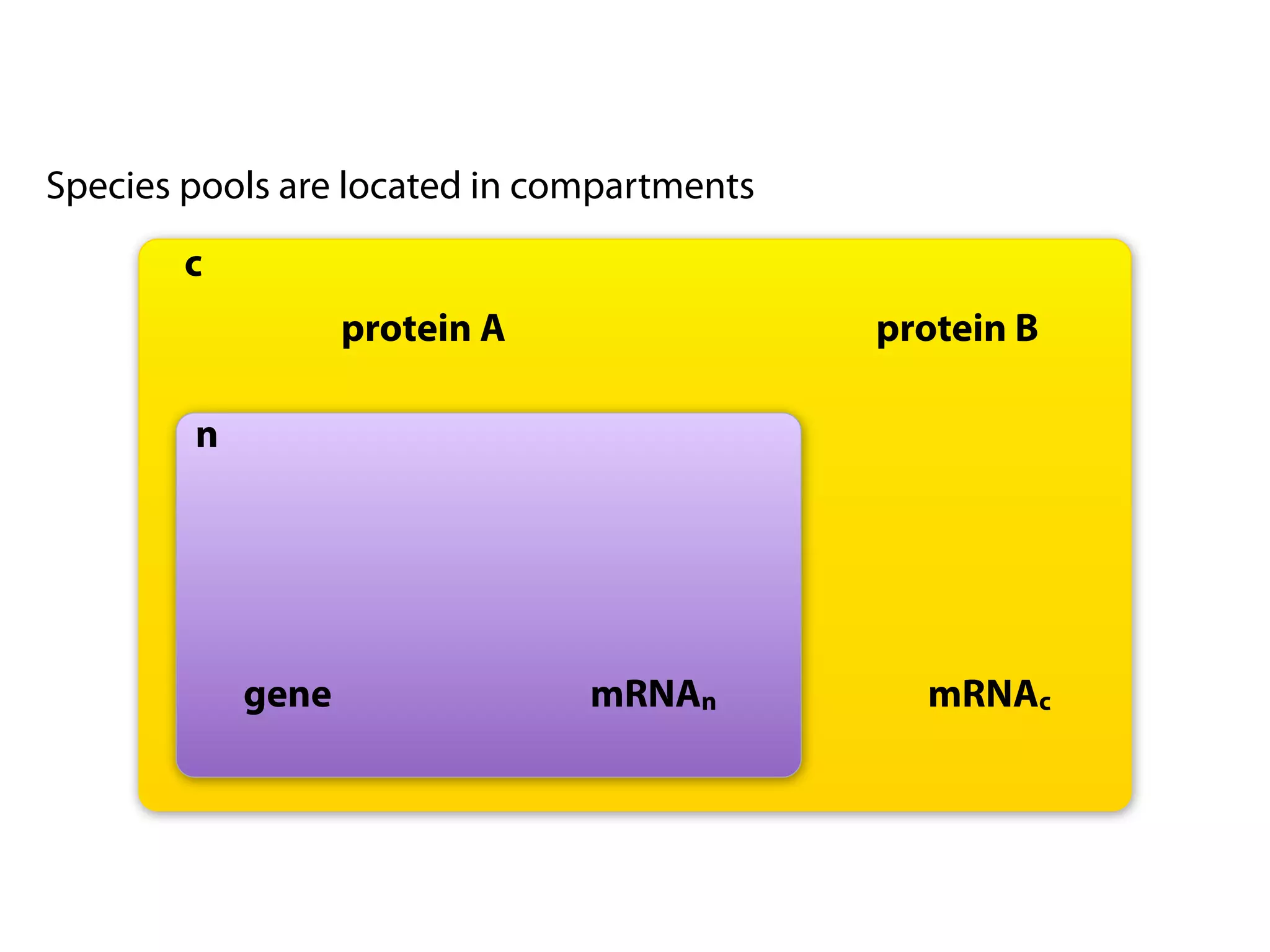
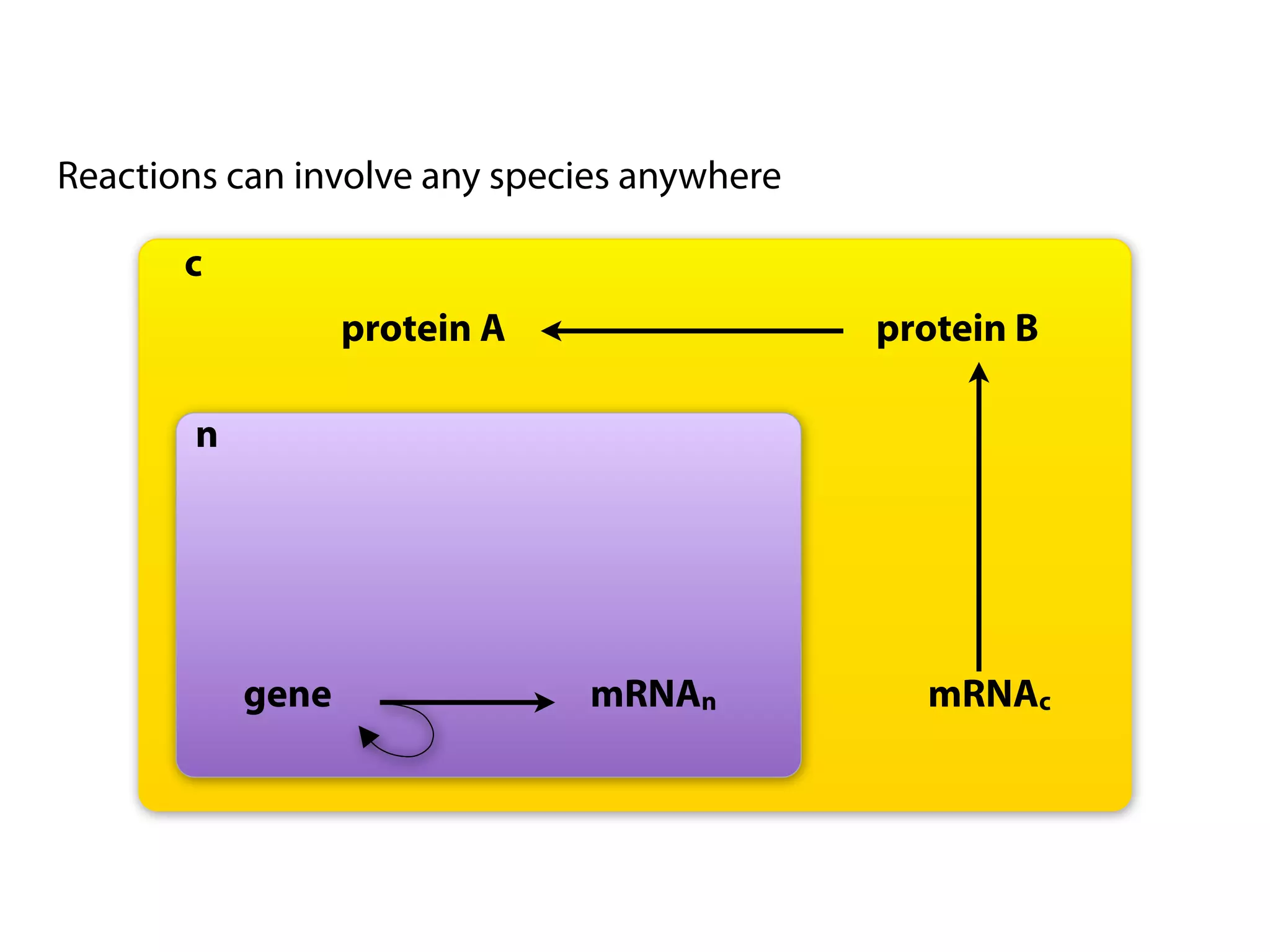
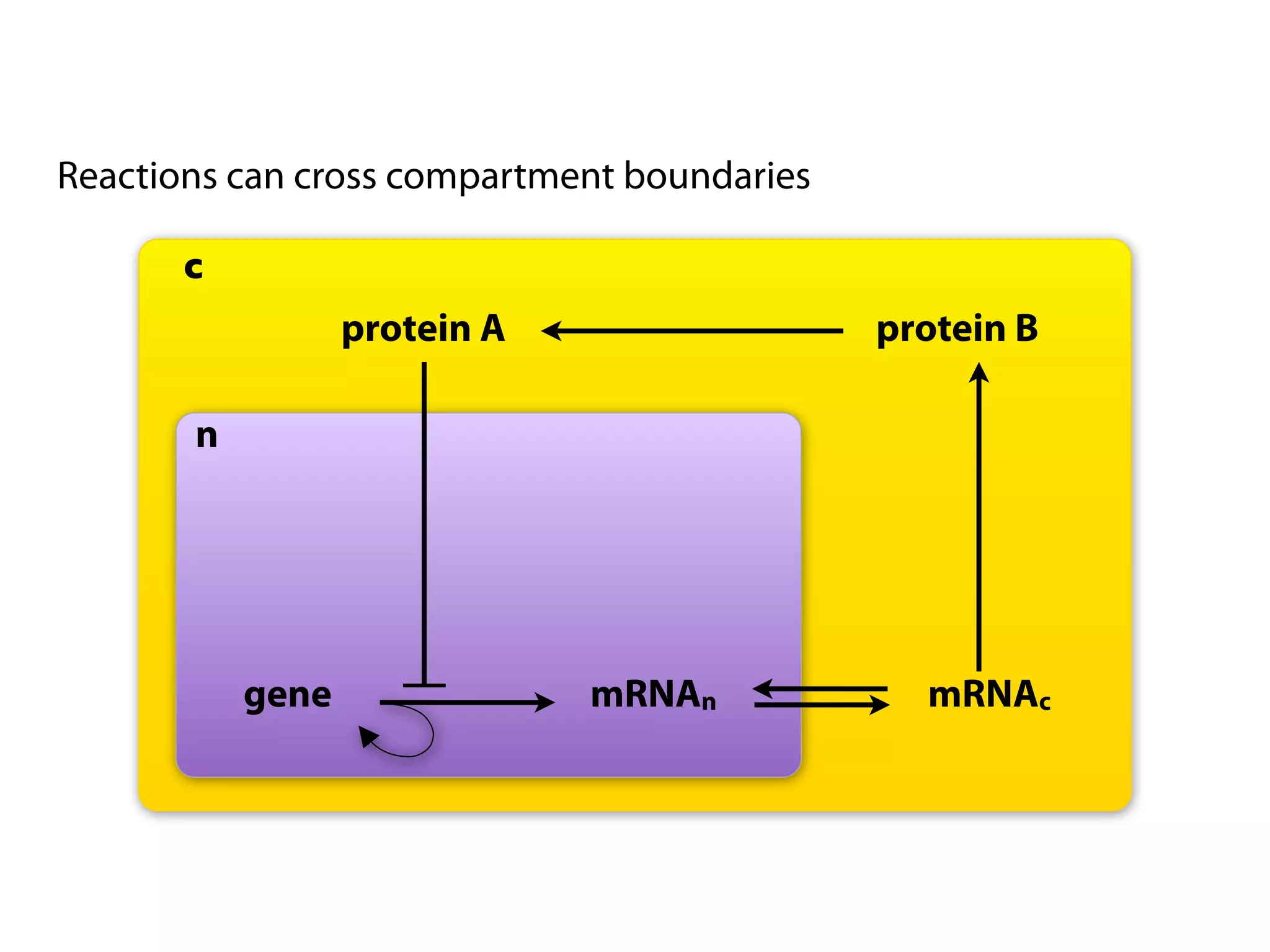
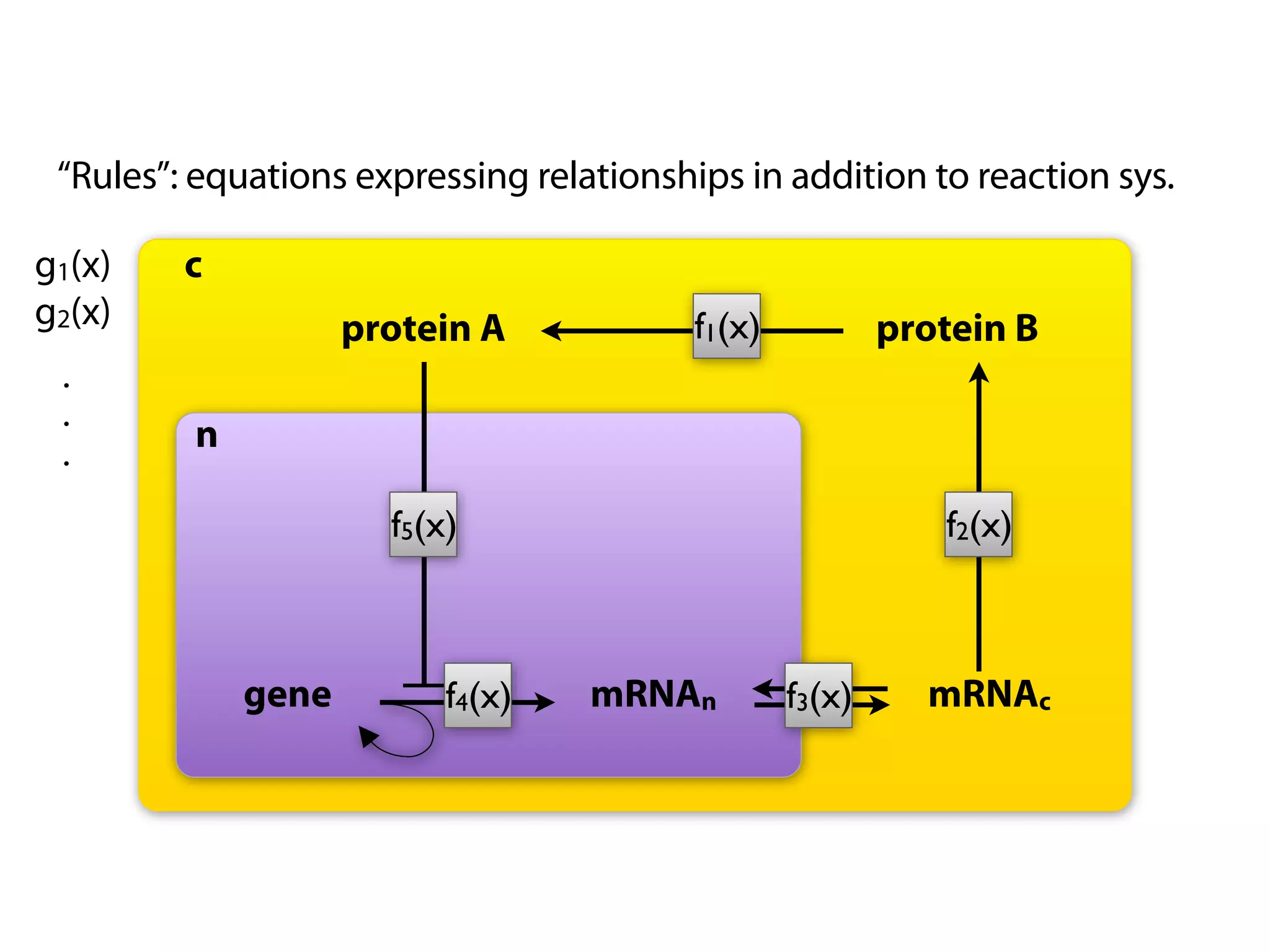
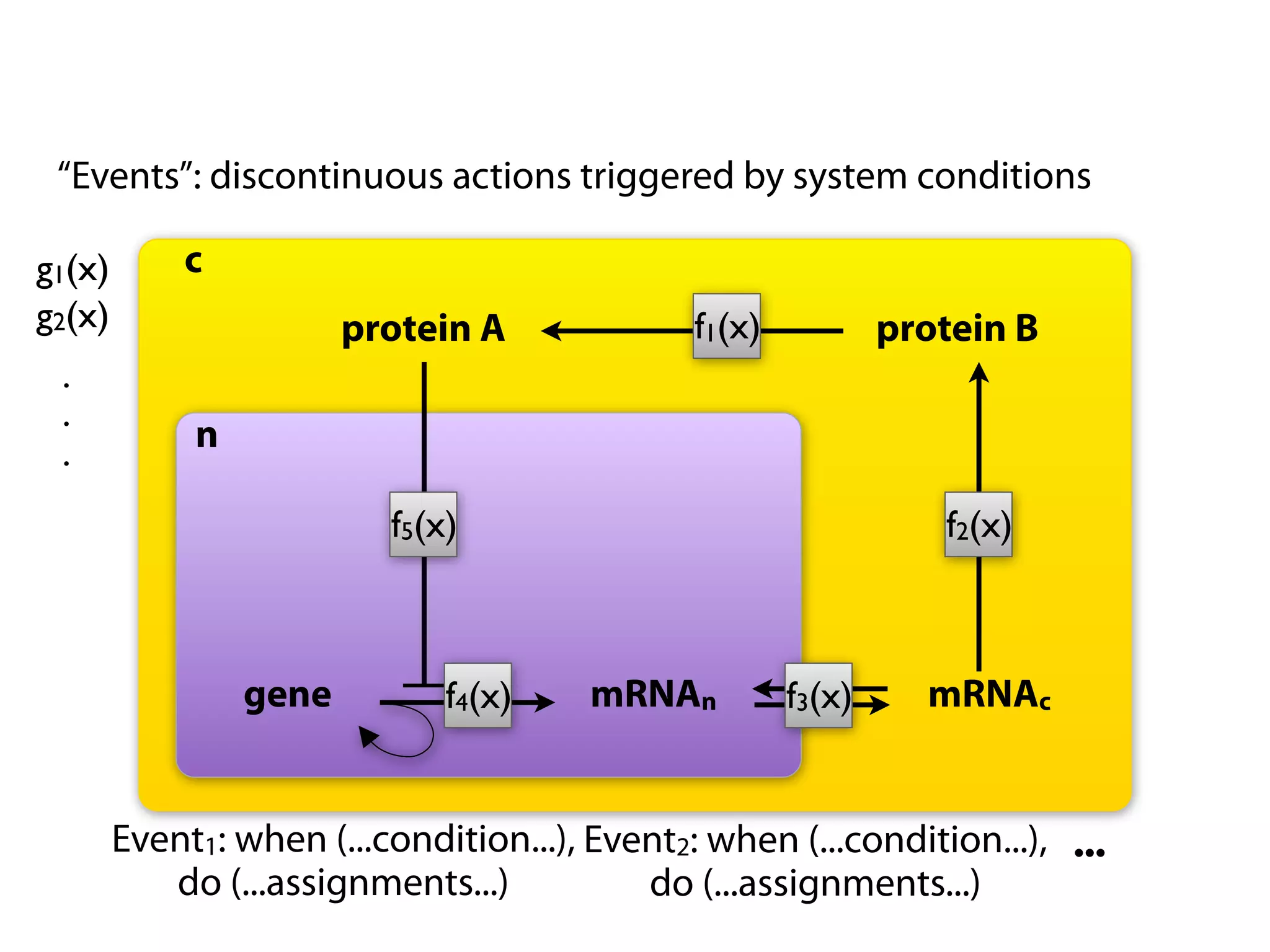
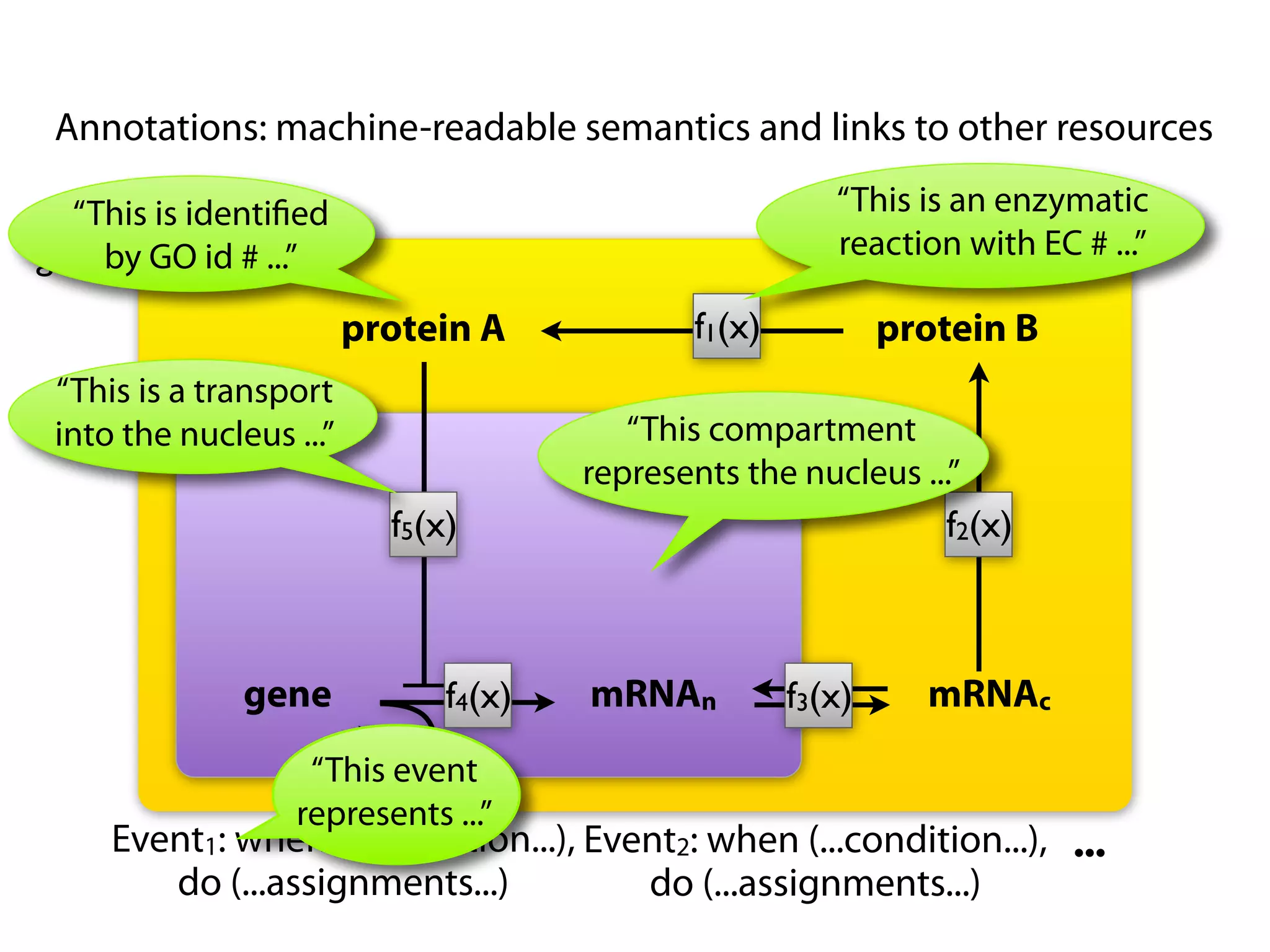
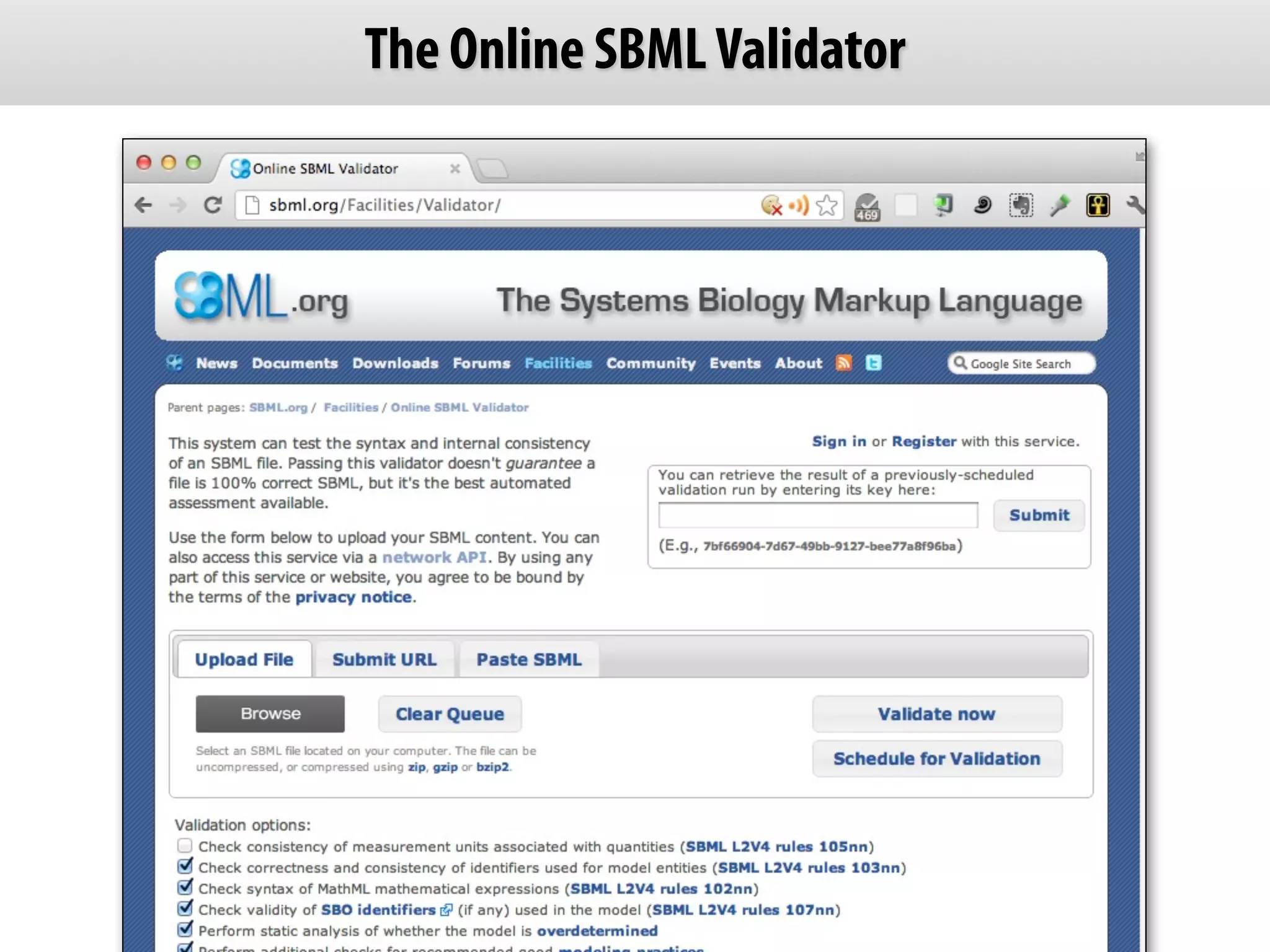
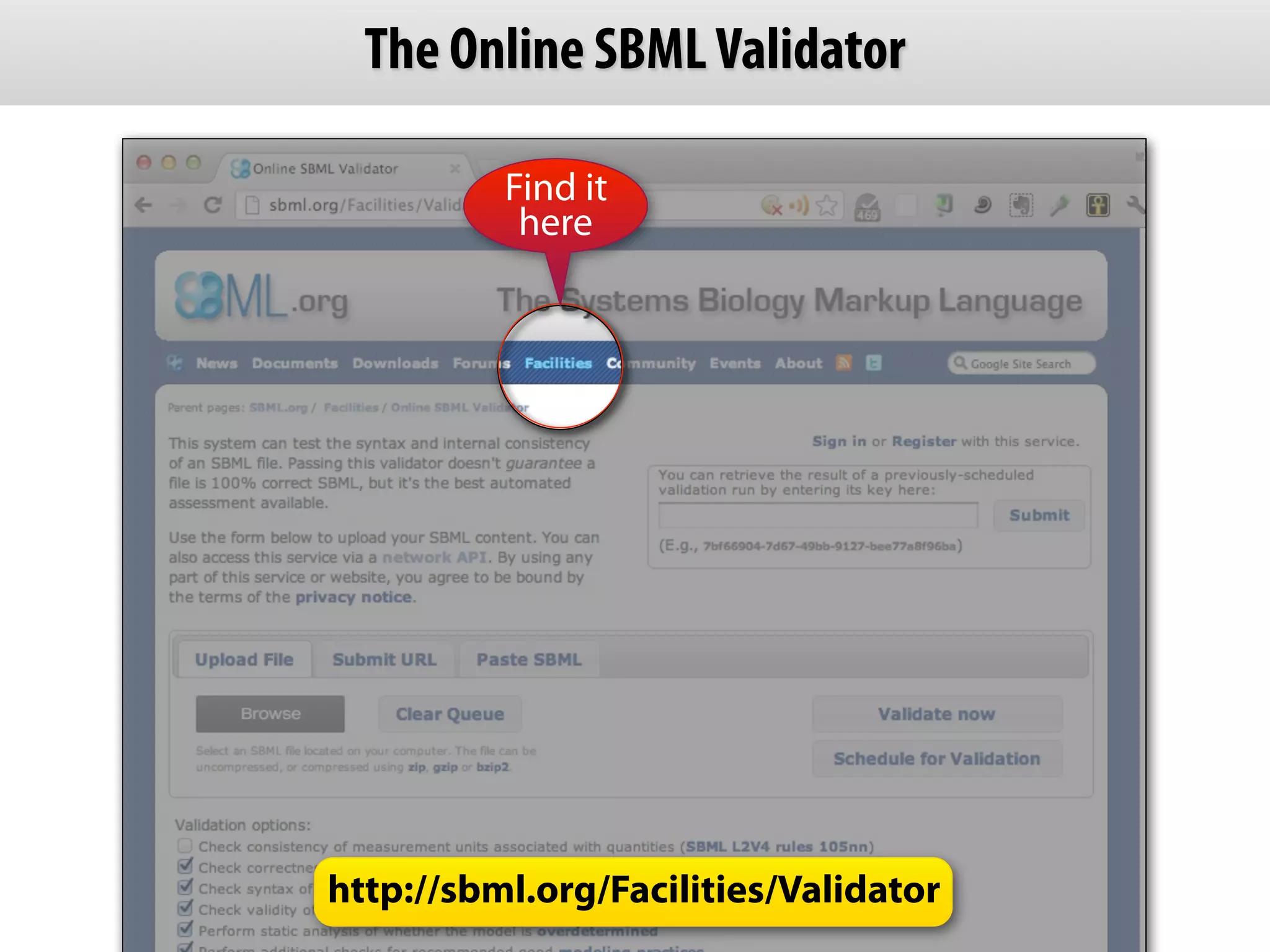
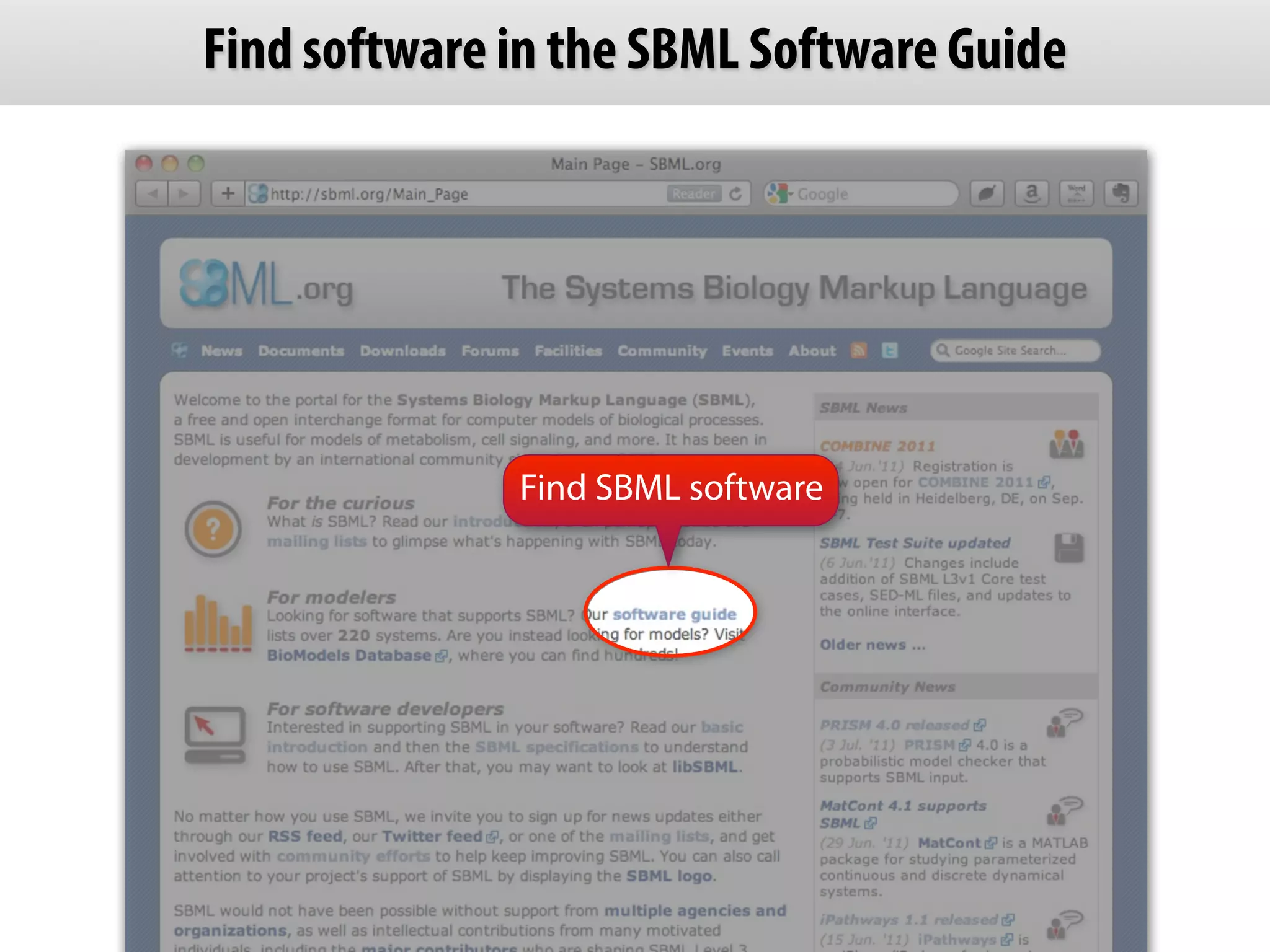
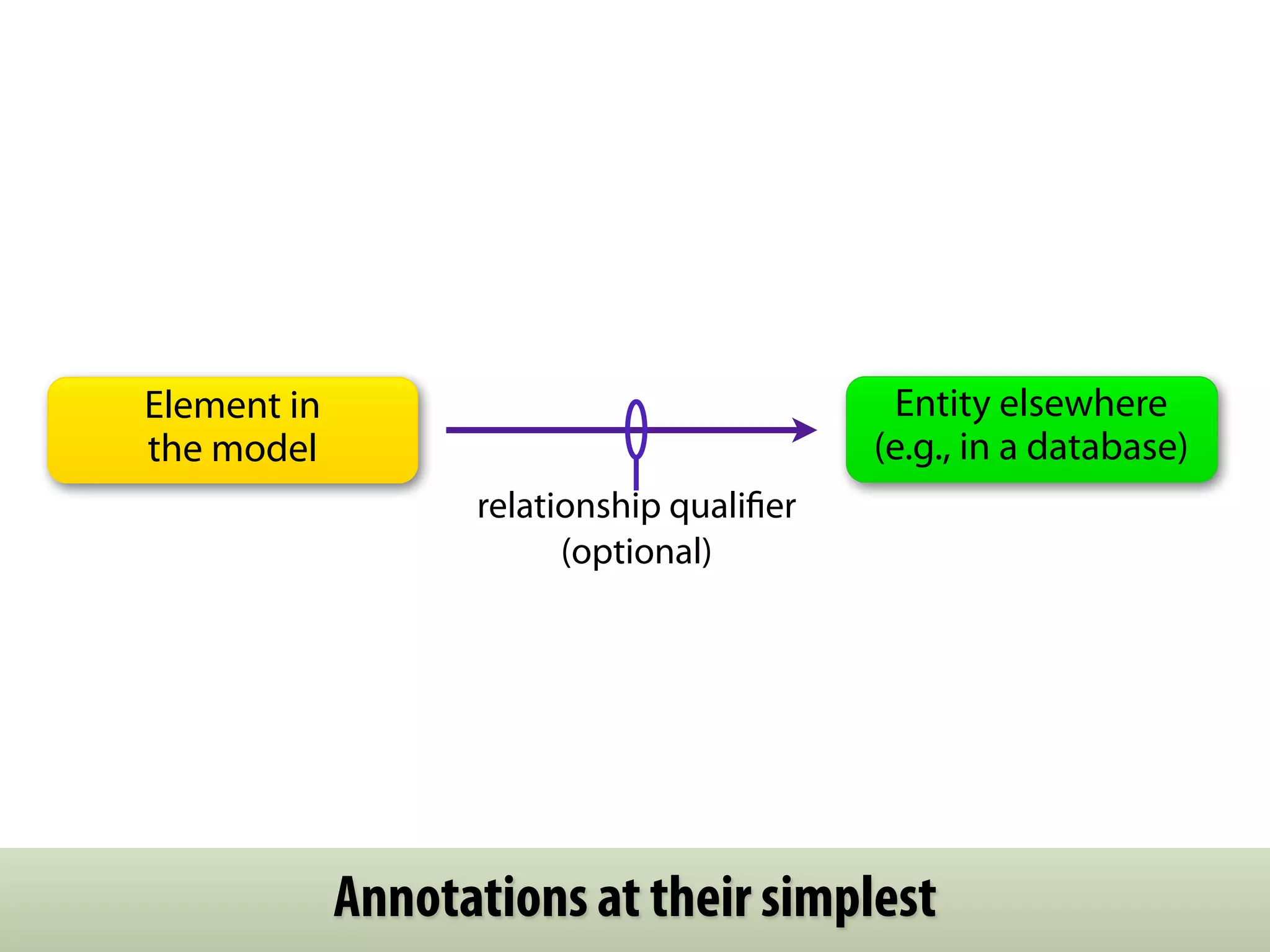
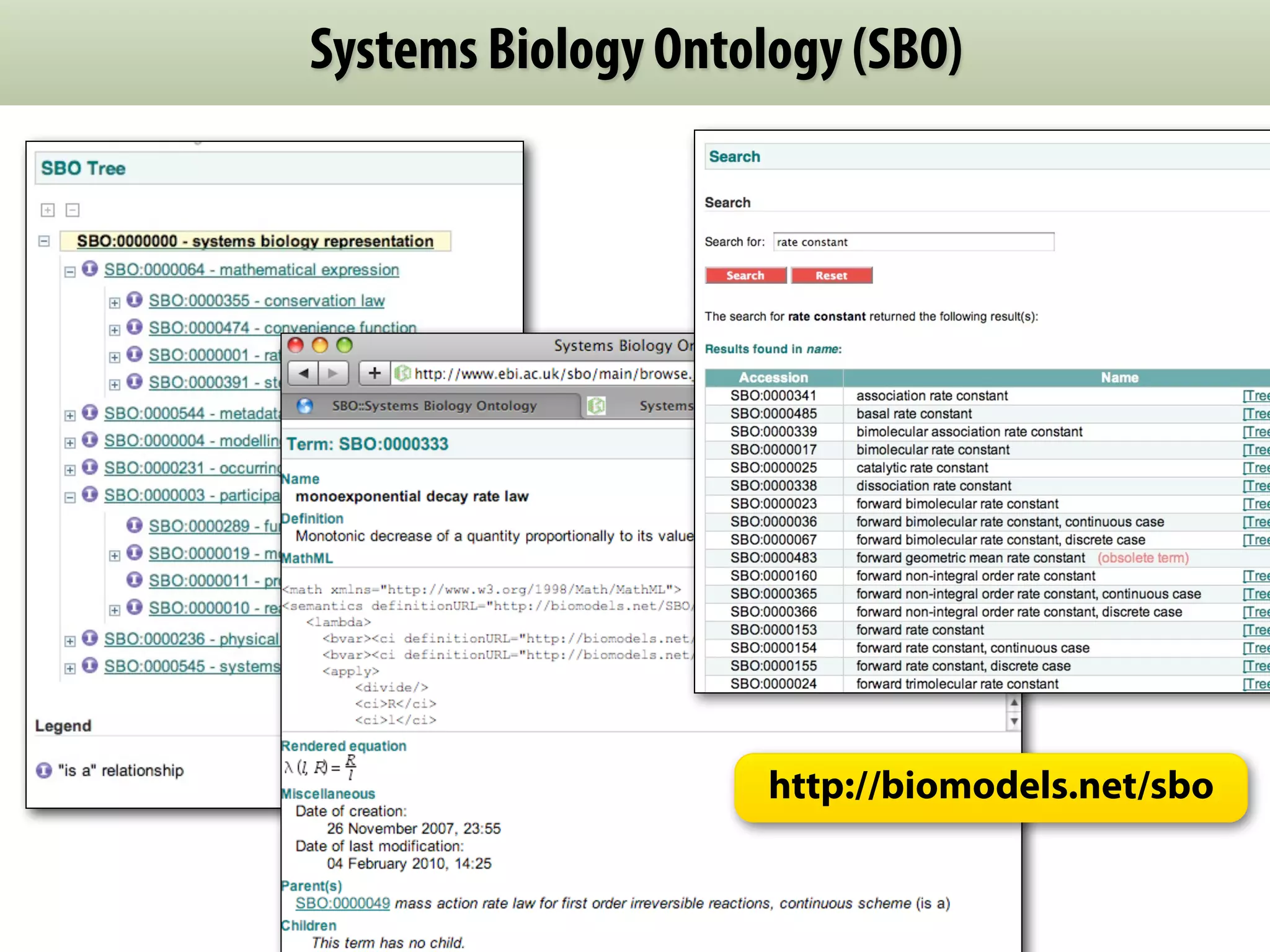
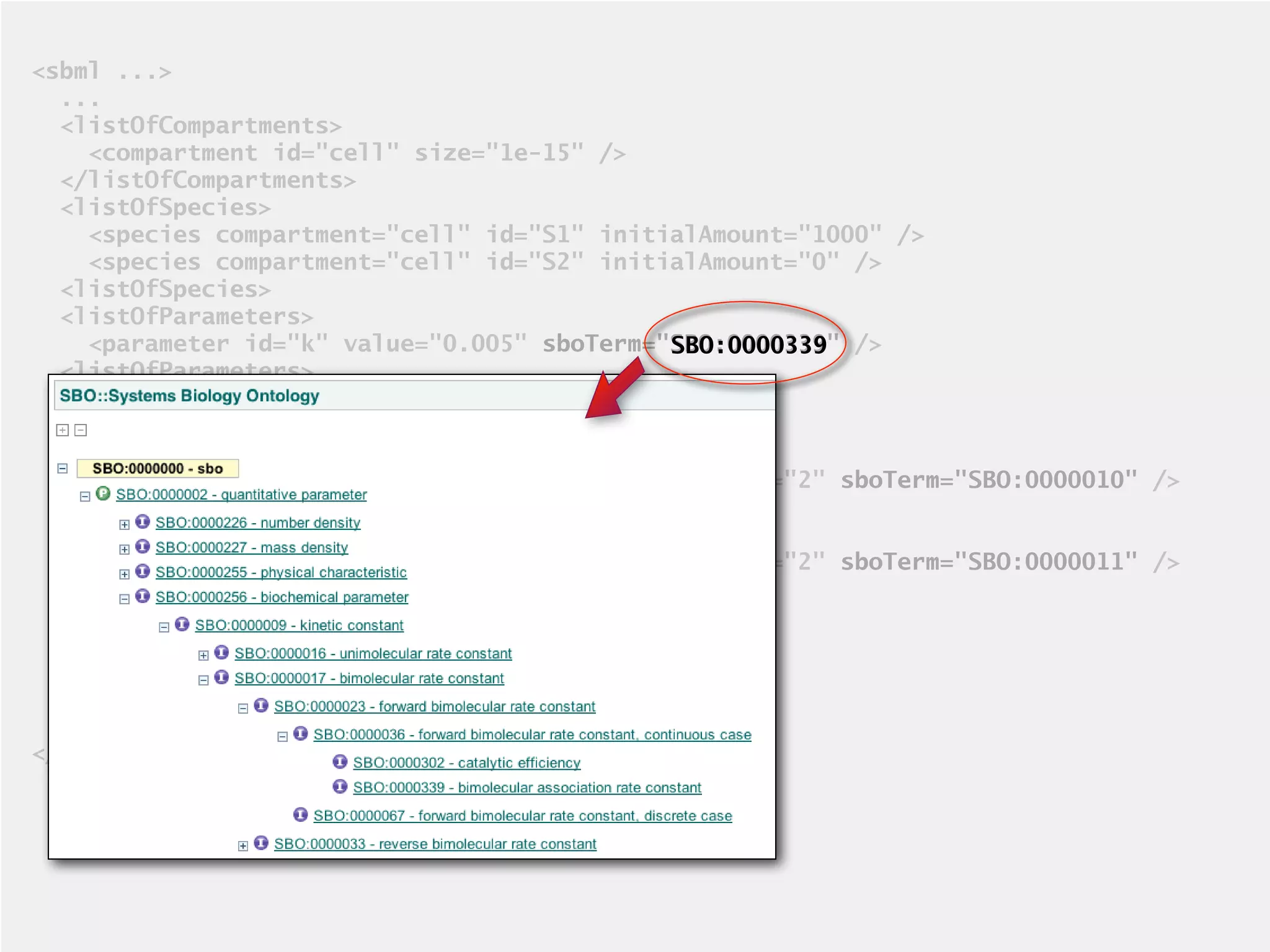
The document provides an overview of the Systems Biology Markup Language (SBML), its features, and its importance for modeling biological processes. It outlines key concepts, resources for modelers, and emphasizes the need for reproducibility and interoperability in biological modeling. Additionally, it discusses upcoming developments in SBML standards and the significance of annotations for enhancing model communication.