The document provides an overview of RDKit, an open-source cheminformatics toolkit used for molecular operations and integration with various software like KNIME and PostgreSQL. It details the features and functionalities of RDKit, including its capabilities in molecular descriptor generation, substructure searching, and data integration, as well as case studies from its application at NIBR. Additionally, it highlights the community involvement and contributions from various stakeholders within academia and industry.




![Some features
§ Input/Output: SMILES/SMARTS, SDF, TDT, PDB,
SLN [1], Corina mol2 [1]
§ “Cheminformatics”:
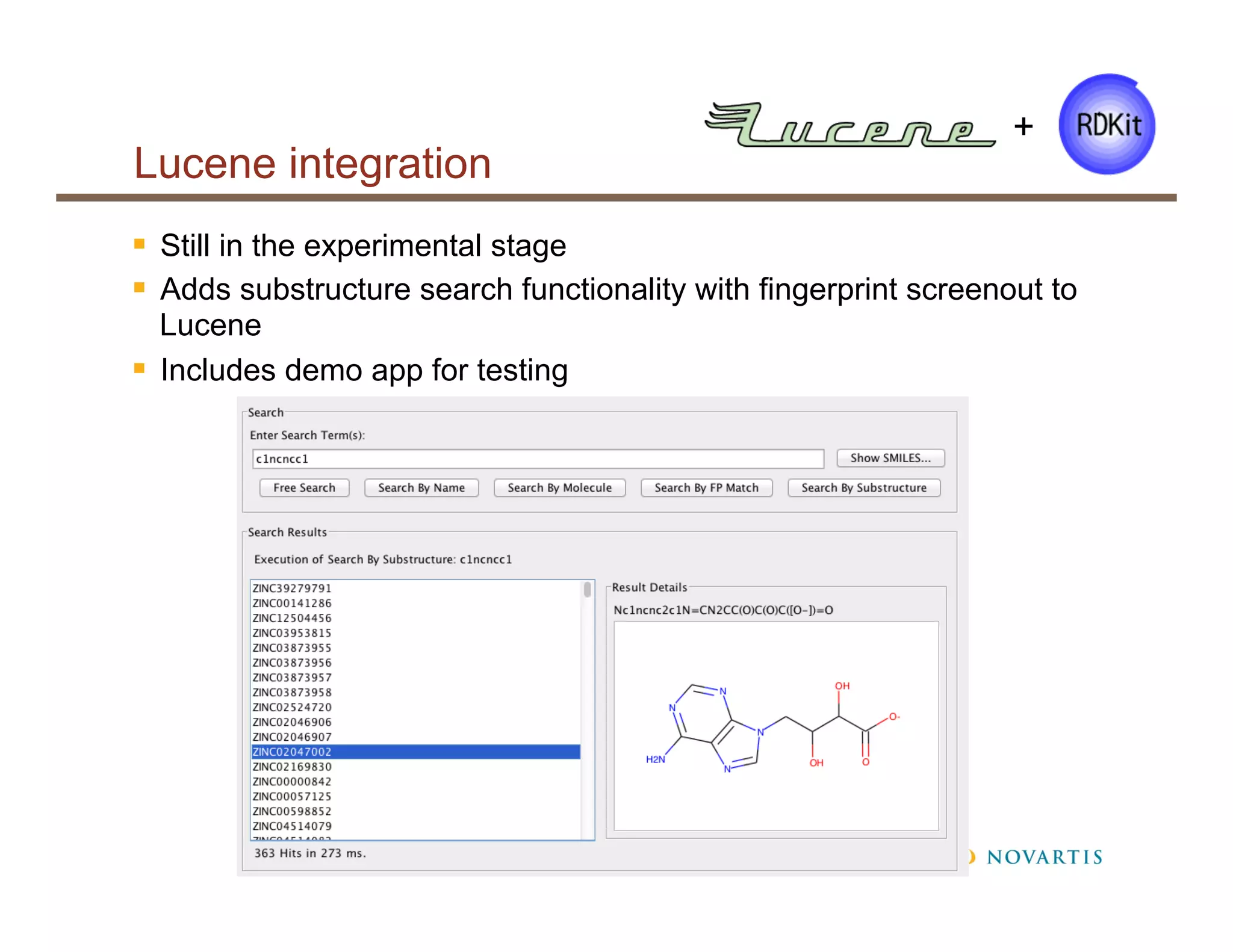
• Substructure searching
• Canonical SMILES
• Chirality support (i.e. R/S or E/Z labeling)
• Chemical transformations (e.g. remove matching
substructures)
• Chemical reactions
§ 2D depiction, including constrained depiction
§ 2D->3D conversion/conformational analysis via
distance geometry
§ UFF and MMFF94 implementation for cleaning up
structures
§ Fingerprinting: Daylight-like, atom pairs, topological
torsions, Morgan algorithm, “MACCS keys”, etc.
§ Similarity/diversity picking
§ 2D pharmacophores [1]
§ Gasteiger-Marsili charges
§ Hierarchical subgraph/fragment analysis
§ Bemis and Murcko scaffold determination
§ RECAP and BRICS implementations
§ Multi-molecule maximum common substructure
§ Feature maps
§ Shape-based similarity
§ Fraggle similarity (from GSK)
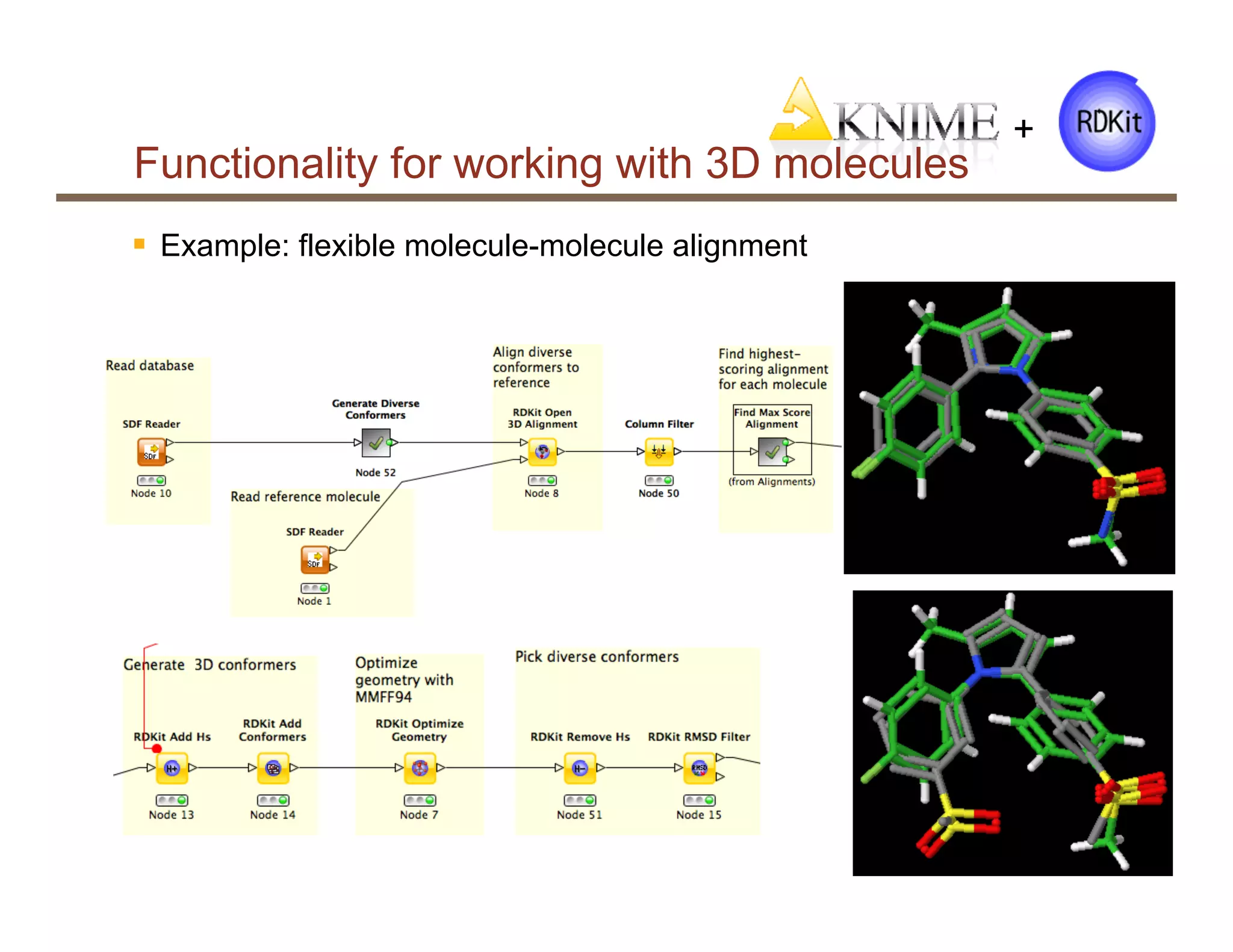
§ Molecule-molecule alignment
§ Open3DAlign implementation
§ Integration with PyMOL for 3D visualization
§ Functional group filtering
§ Salt stripping
§ Molecular descriptor library:
Topological (κ3, Balaban J, etc.), Compositional (Number
of Rings, Number of Aromatic Heterocycles, etc.),
EState, SlogP/SMR (Wildman and Crippen approach),
“MOE like” VSA descriptors, Feature-map vectors
§ Machine Learning:
• Clustering (hierarchical)
• Information theory (Shannon entropy, information
gain, etc.)
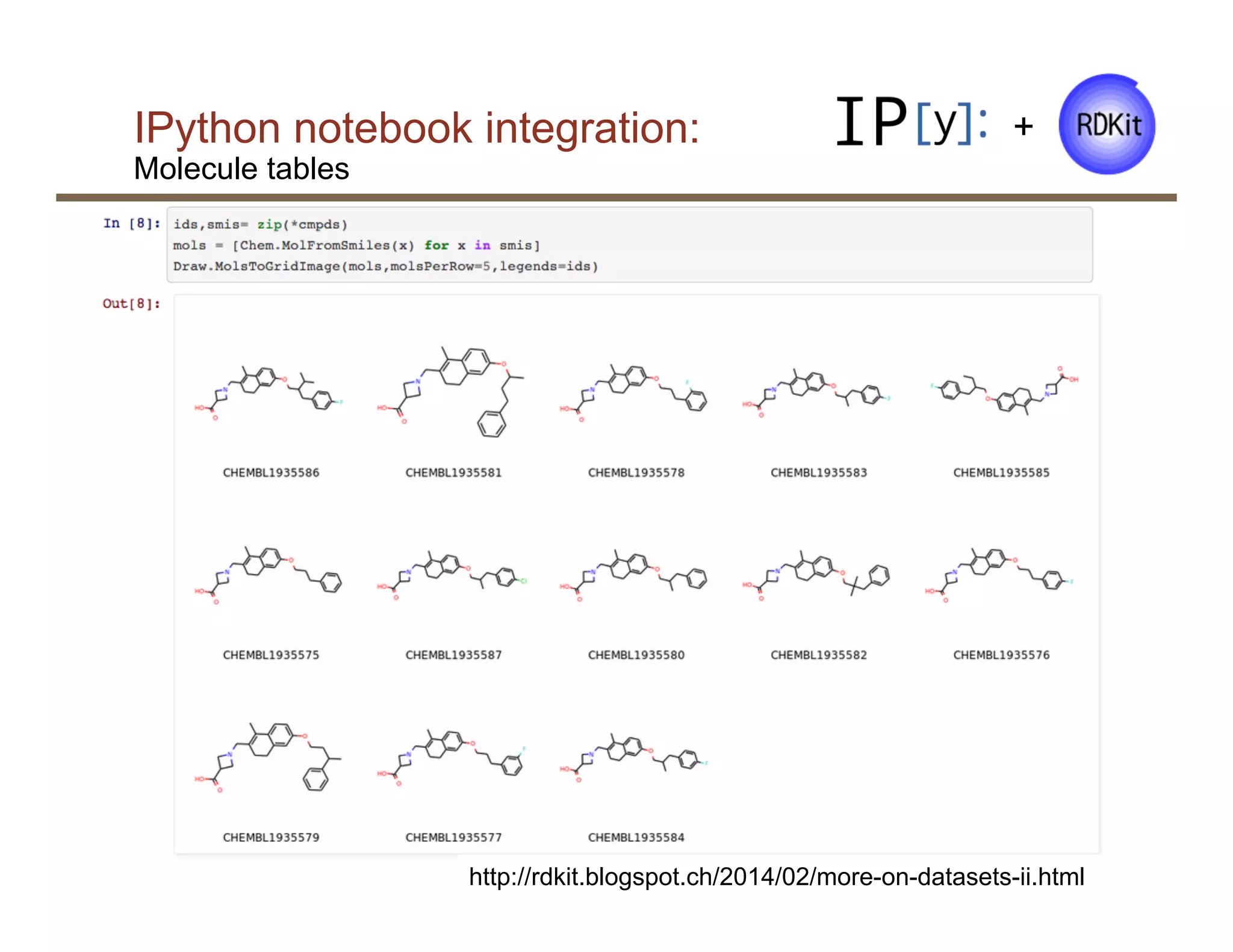
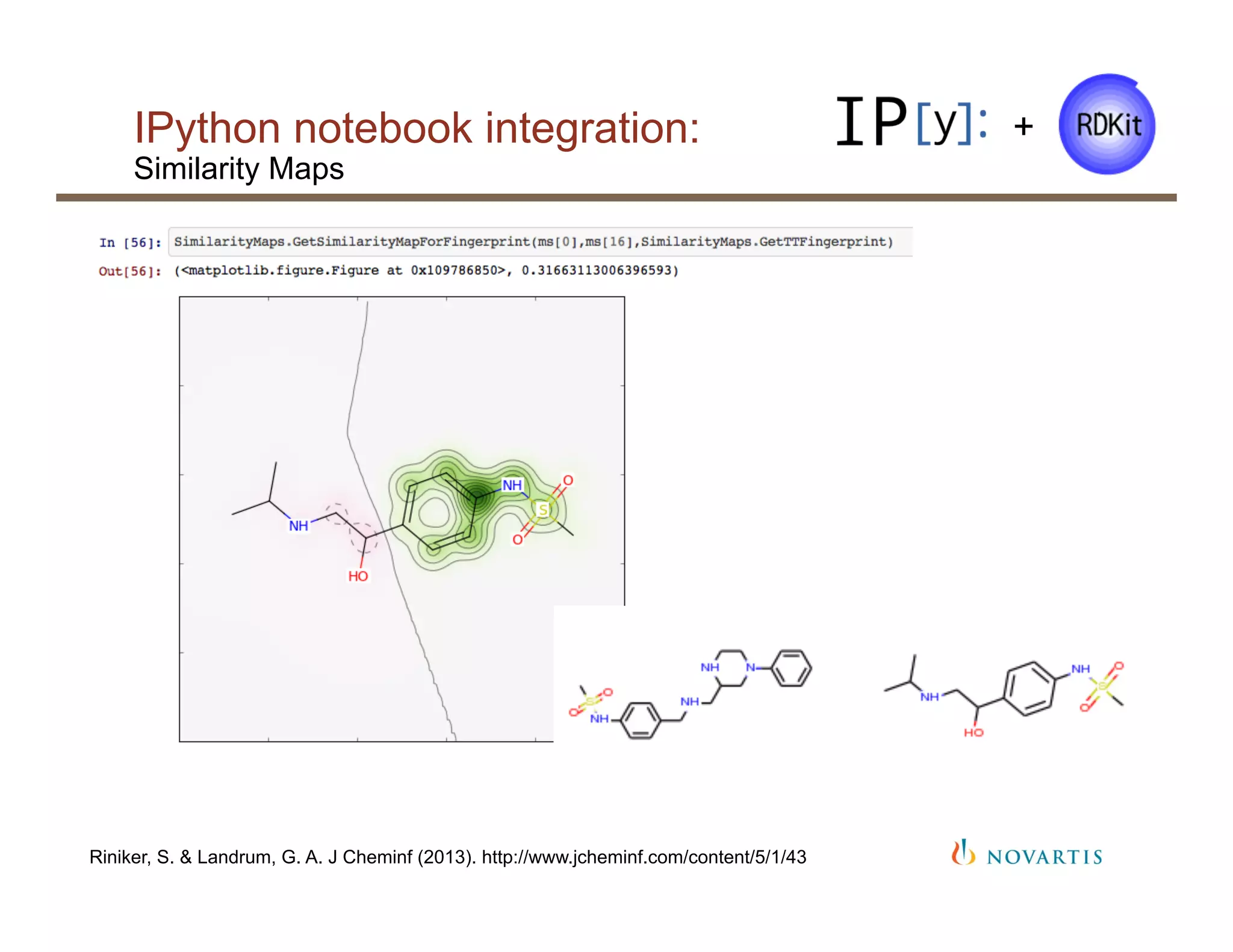
§ Tight integration with the IPython notebook and
pandas
§ Integration with the InChI library
[1] These implementations are functional but are not necessarily
the best, fastest, or most complete.](https://image.slidesharecdn.com/rdkit-140509054918-phpapp02/75/Open-source-from-in-the-enterprise-the-RDKit-5-2048.jpg)