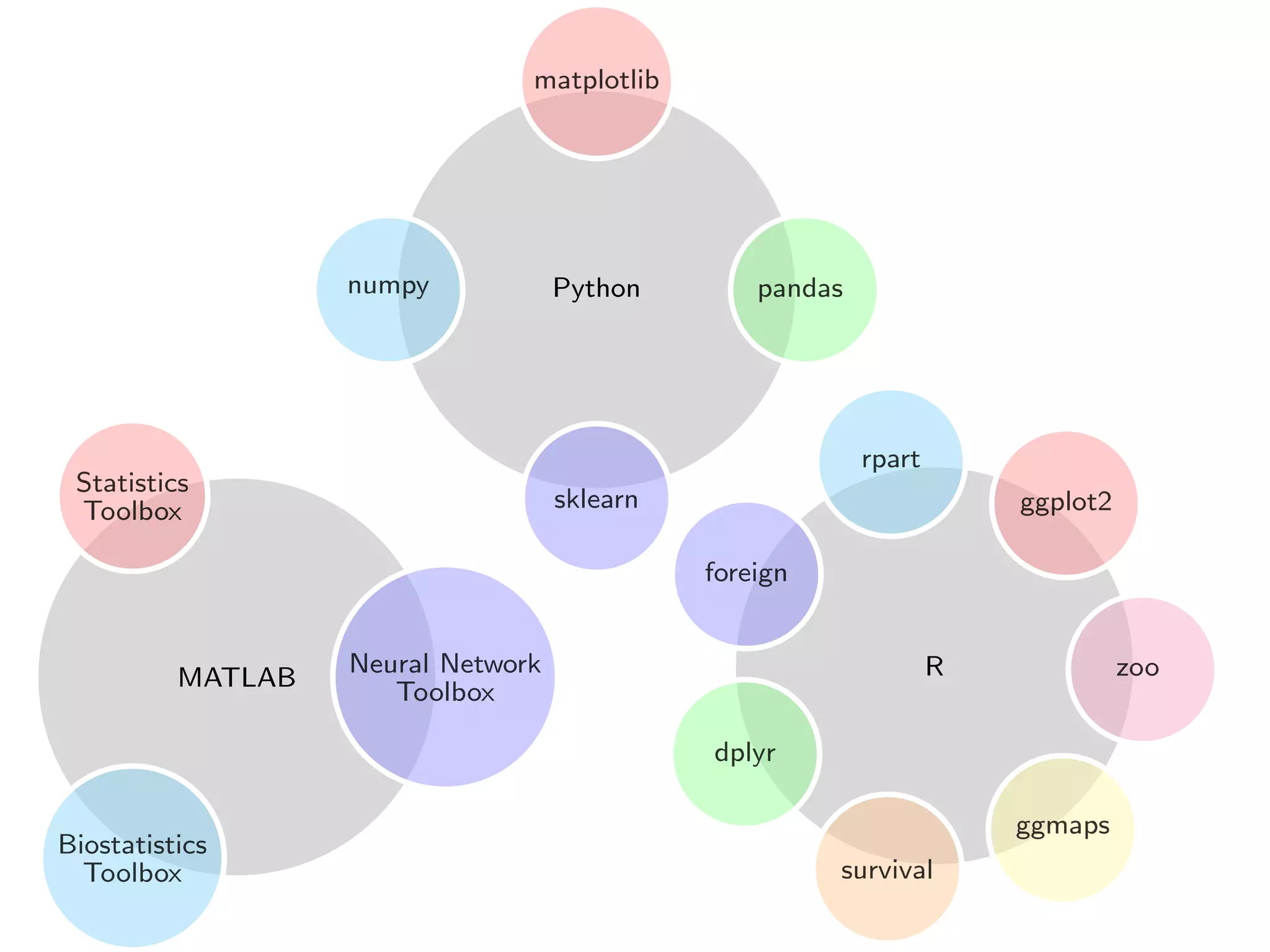
This document discusses Python for scientific computing. It provides notes on NumPy, the fundamental package for scientific computing in Python. NumPy allows vectorized mathematical operations on multidimensional arrays in a simple and efficient manner. The notes cover common NumPy operations and syntax as compared to MATLAB and R. Pandas is also introduced as a package for data manipulation and analysis based on the concept of data frames from R. Examples are given of generating fake data to demonstrate modeling capabilities in Python.





![Numpy Notes
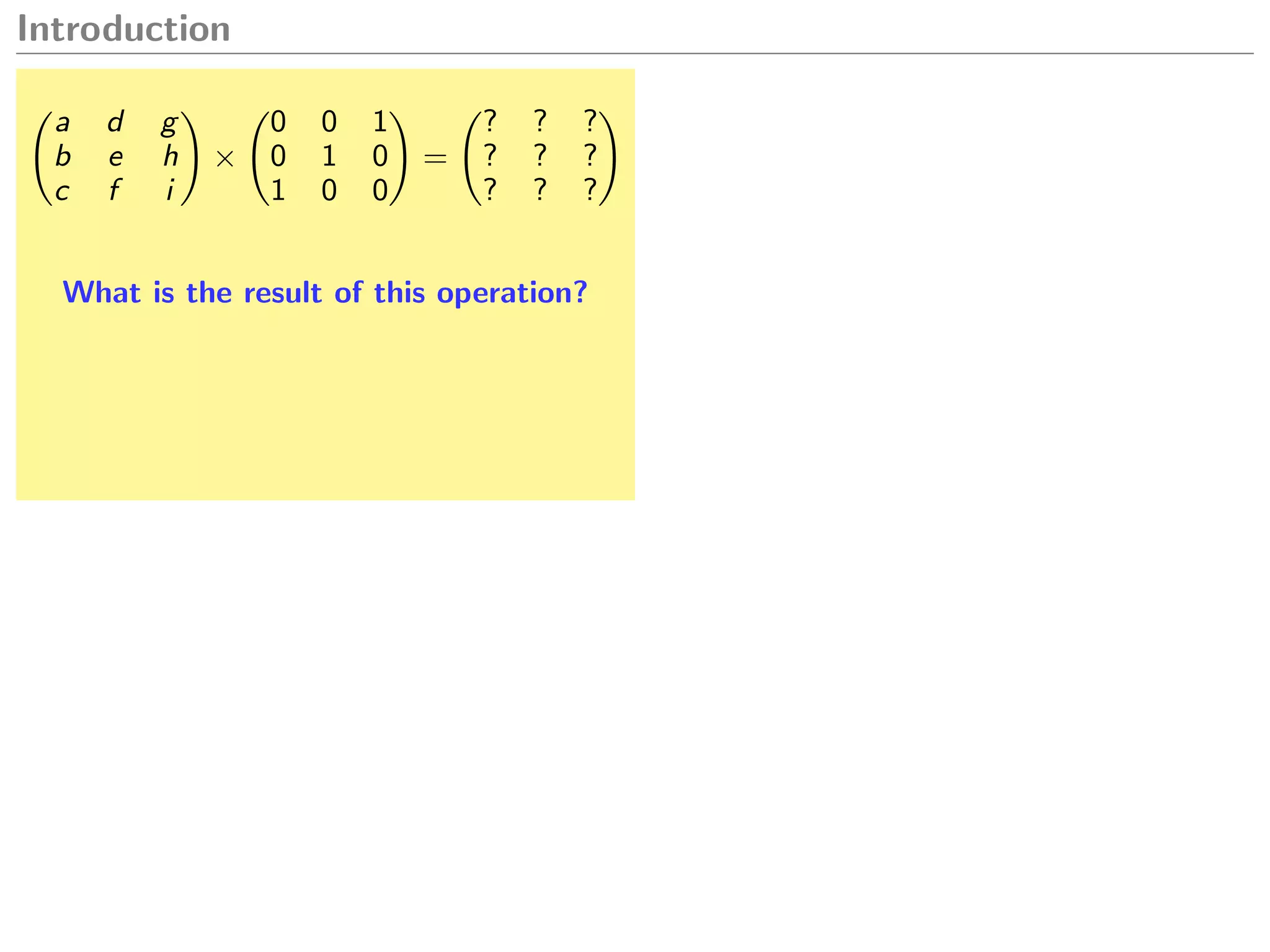
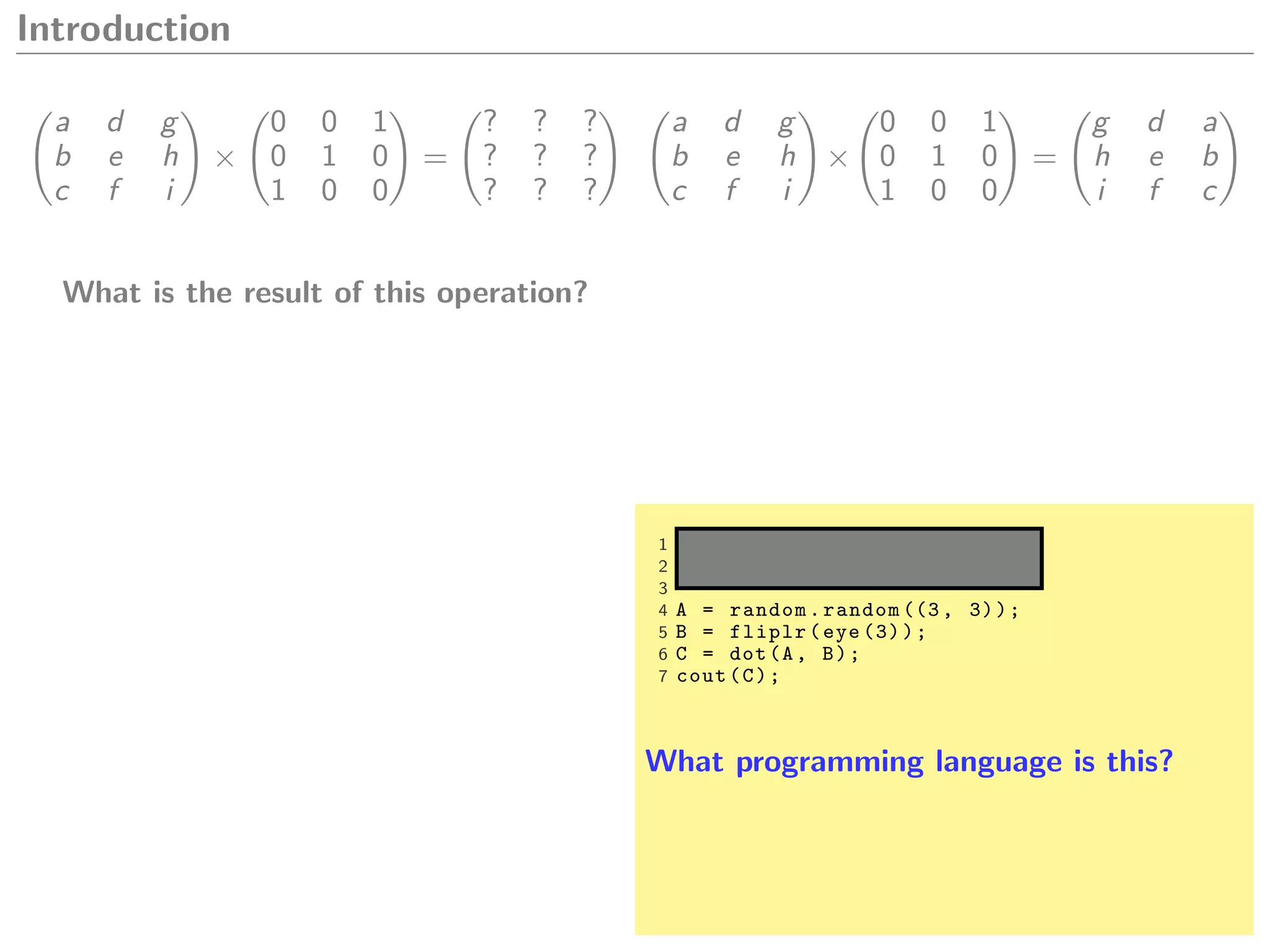
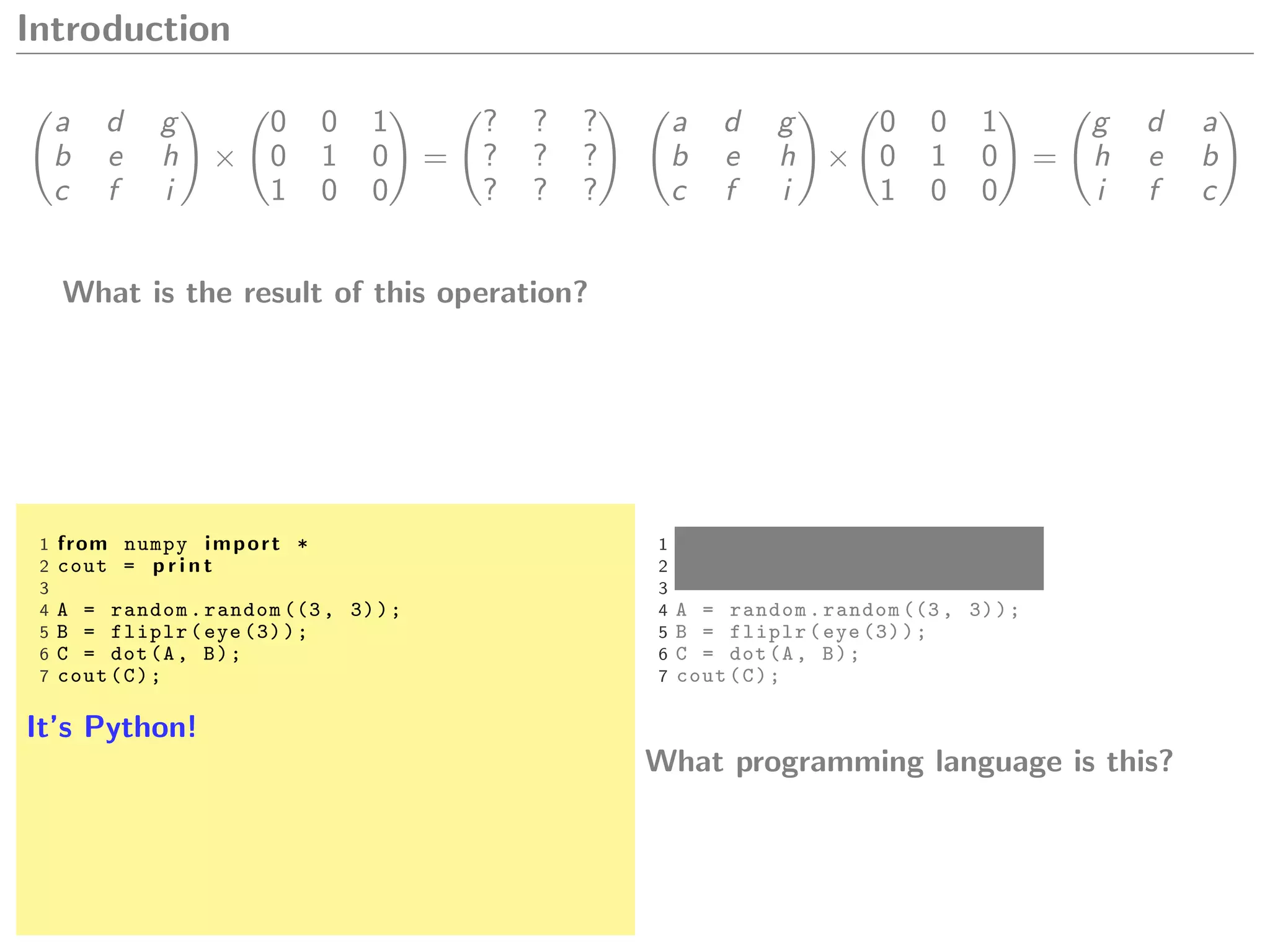
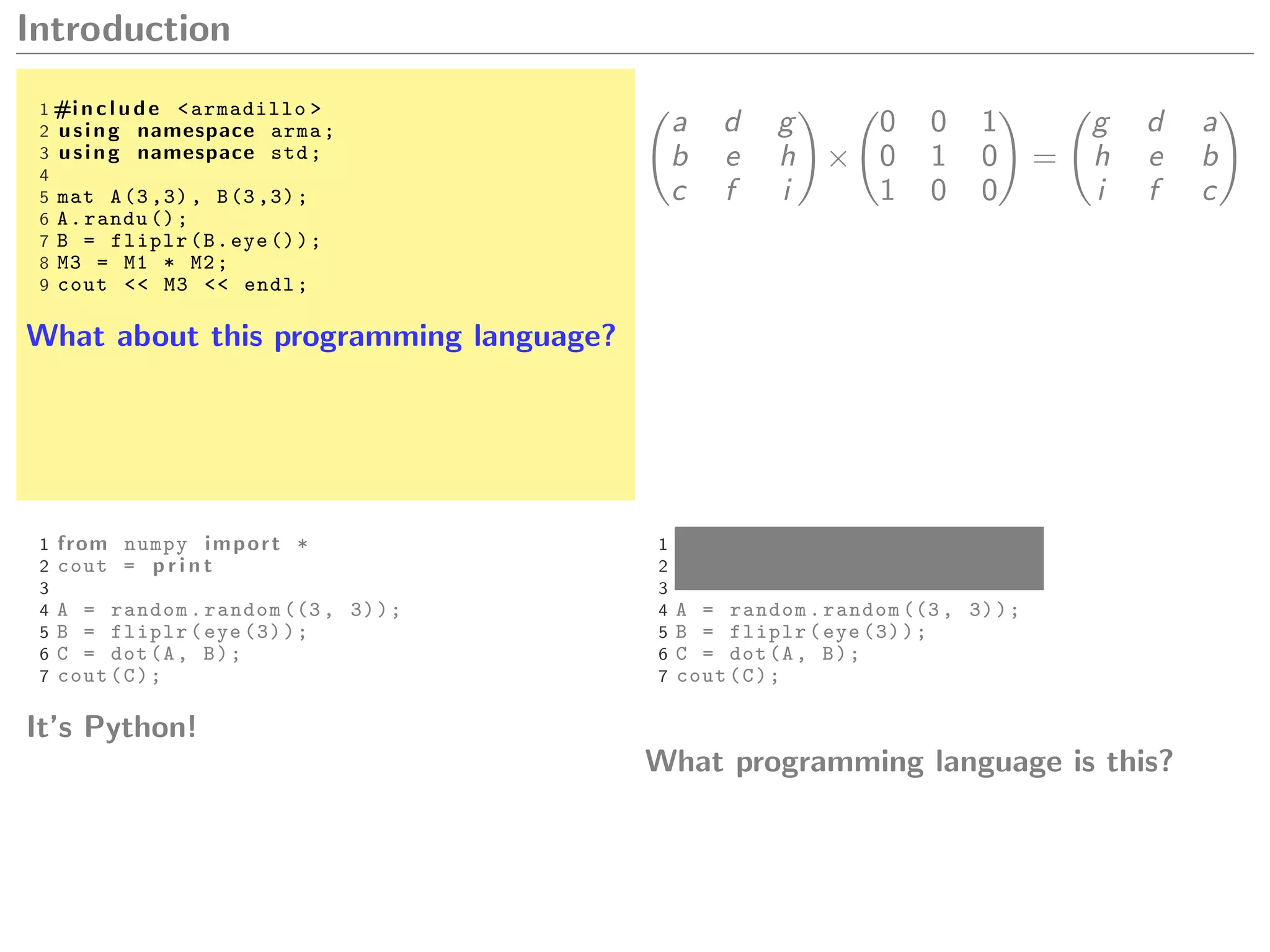
Let A and B be matrices,
Python/Numpy MATLAB R
A.dot(B) A * B A %*% B
A * B A .* B A * B
Operations are elementwise by default (like
R)
Python/Numpy MATLAB R
A.shape size(A) length, nrow, ncol
A[0:4,:] or
A[0:4] or A[:4] A(1:4,:) A[1:4,]
A[0:10:2] A[seq(0, 9, 2)]
A[-4:] A(end-4:end,:) A[nrow(A)-4:nrow(A),]
A.T A.’ t(A)
Numpy in general allows for more succinct
writing.
Furthermore:
Indexing starts at zero.
Intervals are of the form [i, j[](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-13-2048.jpg)
![Numpy Notes
Let A and B be matrices,
Python/Numpy MATLAB R
A.dot(B) A * B A %*% B
A * B A .* B A * B
Operations are elementwise by default (like
R)
Python/Numpy MATLAB R
A.shape size(A) length, nrow, ncol
A[0:4,:] or
A[0:4] or A[:4] A(1:4,:) A[1:4,]
A[0:10:2] A[seq(0, 9, 2)]
A[-4:] A(end-4:end,:) A[nrow(A)-4:nrow(A),]
A.T A.’ t(A)
Numpy in general allows for more succinct
writing.
Furthermore:
Indexing starts at zero.
Intervals are of the form [i, j[
This is further aided by the fact that Numpy
supports arithmetic broadcasting. (unlike
MATLAB or R.)
That is, you can do the following element-
wise multiplication: (6,3) * (6,1). It auto-
matically assumes you want to multiply by
column. In MATLAB, you would have to use
bsxfun(@times,r,A) or first use repmat().](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-14-2048.jpg)
![Numpy Notes
Let A and B be matrices,
Python/Numpy MATLAB R
A.dot(B) A * B A %*% B
A * B A .* B A * B
Operations are elementwise by default (like
R)
Python/Numpy MATLAB R
A.shape size(A) length, nrow, ncol
A[0:4,:] or
A[0:4] or A[:4] A(1:4,:) A[1:4,]
A[0:10:2] A[seq(0, 9, 2)]
A[-4:] A(end-4:end,:) A[nrow(A)-4:nrow(A),]
A.T A.’ t(A)
Numpy in general allows for more succinct
writing.
Furthermore:
Indexing starts at zero.
Intervals are of the form [i, j[
Something like the following is valid in
Numpy...
1 import skimage.data
2 img1 = skimage.data.astronaut ()
3 img2 = skimage.data.moon ()
4 p r i n t (img1.shape) # (512 , 512 , 3)
5 p r i n t (img2.shape) # (512 , 512)
6
7 import matplotlib.pyplot as plt
8 plt.subplot (1, 2, 1)
9 plt.imshow(img1)
10 plt.subplot (1, 2, 2)
11 plt.imshow(img2 , cmap=’gray ’)
12 plt.show ()
This is further aided by the fact that Numpy
supports arithmetic broadcasting. (unlike
MATLAB or R.)
That is, you can do the following element-
wise multiplication: (6,3) * (6,1). It auto-
matically assumes you want to multiply by
column. In MATLAB, you would have to use
bsxfun(@times,r,A) or first use repmat().](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-15-2048.jpg)
![Numpy Notes
Python/Numpy MATLAB R
A.shape size(A) length, nrow, ncol
A[0:4,:] or
A[0:4] or A[:4] A(1:4,:) A[1:4,]
A[0:10:2] A[seq(0, 9, 2)]
A[-4:] A(end-4:end,:) A[nrow(A)-4:nrow(A),]
A.T A.’ t(A)
Numpy in general allows for more succinct
writing.
Furthermore:
Indexing starts at zero.
Intervals are of the form [i, j[
Something like the following is valid in
Numpy...
1 import skimage.data
2 img1 = skimage.data.astronaut ()
3 img2 = skimage.data.moon ()
4 p r i n t (img1.shape) # (512 , 512 , 3)
5 p r i n t (img2.shape) # (512 , 512)
6
7 import matplotlib.pyplot as plt
8 plt.subplot (1, 2, 1)
9 plt.imshow(img1)
10 plt.subplot (1, 2, 2)
11 plt.imshow(img2 , cmap=’gray ’)
12 plt.show ()
This is further aided by the fact that Numpy
supports arithmetic broadcasting. (unlike
MATLAB or R.)
That is, you can do the following element-
wise multiplication: (6,3) * (6,1). It auto-
matically assumes you want to multiply by
column. In MATLAB, you would have to use
bsxfun(@times,r,A) or first use repmat().](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-16-2048.jpg)
![Numpy Notes
Arithmetic mean
1 img2 = img2[:, :, np.newaxis] #(512 ,512 ,1)
2 img1 = img1.astype(np.uint32)
3 img2 = img2.astype(np.uint32)
4 img3 = (img1 + img2)//2
5 img3 = img3.astype(np.uint8)
6 plt.imshow(img3)
7 plt.show ()
Something like the following is valid in
Numpy...
1 import skimage.data
2 img1 = skimage.data.astronaut ()
3 img2 = skimage.data.moon ()
4 p r i n t (img1.shape) # (512 , 512 , 3)
5 p r i n t (img2.shape) # (512 , 512)
6
7 import matplotlib.pyplot as plt
8 plt.subplot (1, 2, 1)
9 plt.imshow(img1)
10 plt.subplot (1, 2, 2)
11 plt.imshow(img2 , cmap=’gray ’)
12 plt.show ()
This is further aided by the fact that Numpy
supports arithmetic broadcasting. (unlike
MATLAB or R.)
That is, you can do the following element-
wise multiplication: (6,3) * (6,1). It auto-
matically assumes you want to multiply by
column. In MATLAB, you would have to use
bsxfun(@times,r,A) or first use repmat().](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-17-2048.jpg)
![Numpy Notes
Arithmetic mean
1 img2 = img2[:, :, np.newaxis] #(512 ,512 ,1)
2 img1 = img1.astype(np.uint32)
3 img2 = img2.astype(np.uint32)
4 img3 = (img1 + img2)//2
5 img3 = img3.astype(np.uint8)
6 plt.imshow(img3)
7 plt.show ()
Something like the following is valid in
Numpy...
1 import skimage.data
2 img1 = skimage.data.astronaut ()
3 img2 = skimage.data.moon ()
4 p r i n t (img1.shape) # (512 , 512 , 3)
5 p r i n t (img2.shape) # (512 , 512)
6
7 import matplotlib.pyplot as plt
8 plt.subplot (1, 2, 1)
9 plt.imshow(img1)
10 plt.subplot (1, 2, 2)
11 plt.imshow(img2 , cmap=’gray ’)
12 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-18-2048.jpg)
![Numpy Notes
Arithmetic mean
1 img2 = img2[:, :, np.newaxis] #(512 ,512 ,1)
2 img1 = img1.astype(np.uint32)
3 img2 = img2.astype(np.uint32)
4 img3 = (img1 + img2)//2
5 img3 = img3.astype(np.uint8)
6 plt.imshow(img3)
7 plt.show ()
Geometric mean
1 img2 = img2[:, :, np.newaxis]
2 img1 = img1.astype(np.uint32)
3 img2 = img2.astype(np.uint32)
4 img3 = np.sqrt(img1 * img2)
5 img3 = img3.astype(np.uint8)
6 plt.imshow(img3)
7 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-19-2048.jpg)
![Numpy Notes
Arithmetic mean
1 img2 = img2[:, :, np.newaxis] #(512 ,512 ,1)
2 img1 = img1.astype(np.uint32)
3 img2 = img2.astype(np.uint32)
4 img3 = (img1 + img2)//2
5 img3 = img3.astype(np.uint8)
6 plt.imshow(img3)
7 plt.show ()
Geometric mean
1 img2 = img2[:, :, np.newaxis]
2 img1 = img1.astype(np.uint32)
3 img2 = img2.astype(np.uint32)
4 img3 = np.sqrt(img1 * img2)
5 img3 = img3.astype(np.uint8)
6 plt.imshow(img3)
7 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-20-2048.jpg)




![Models
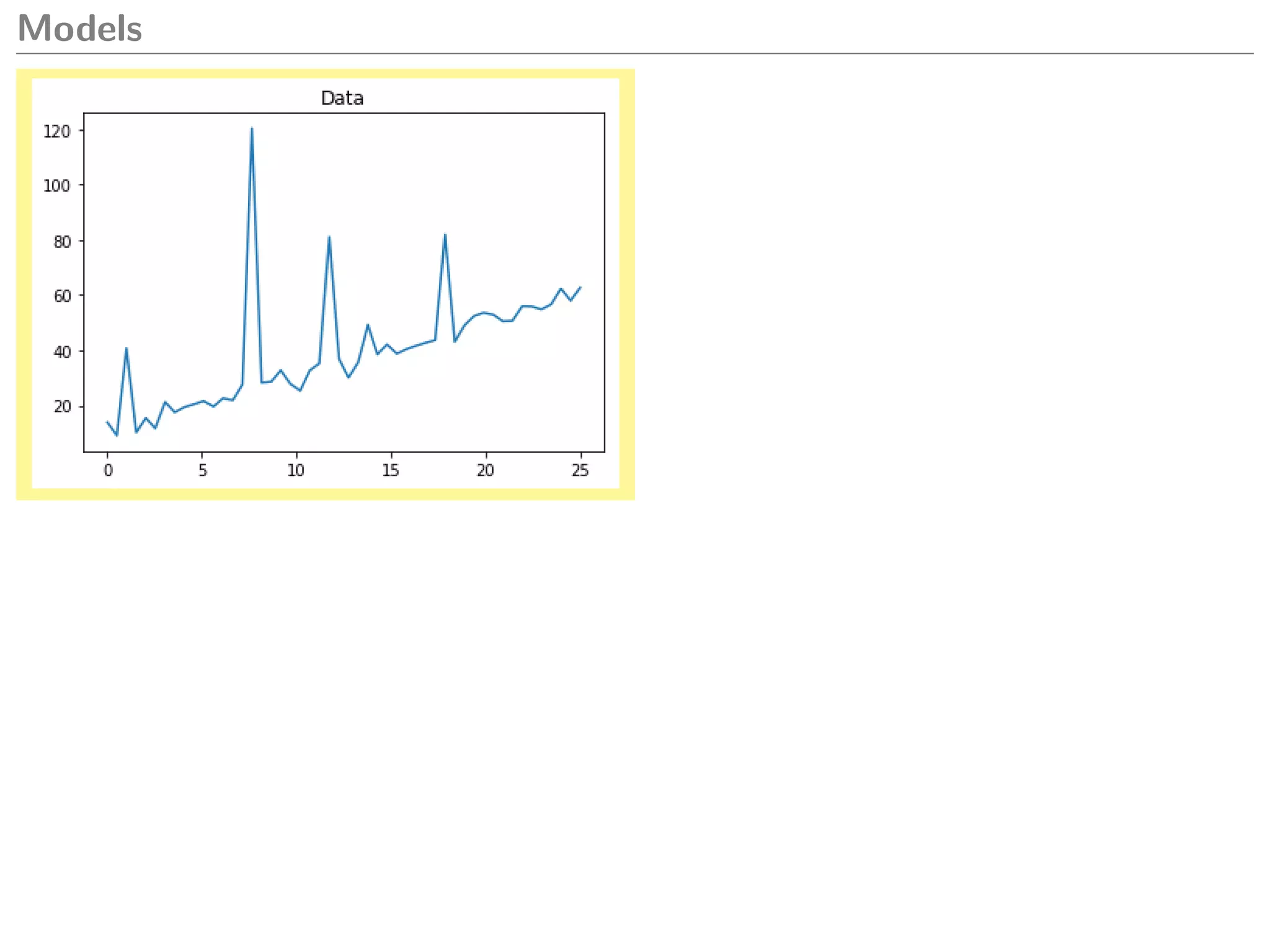
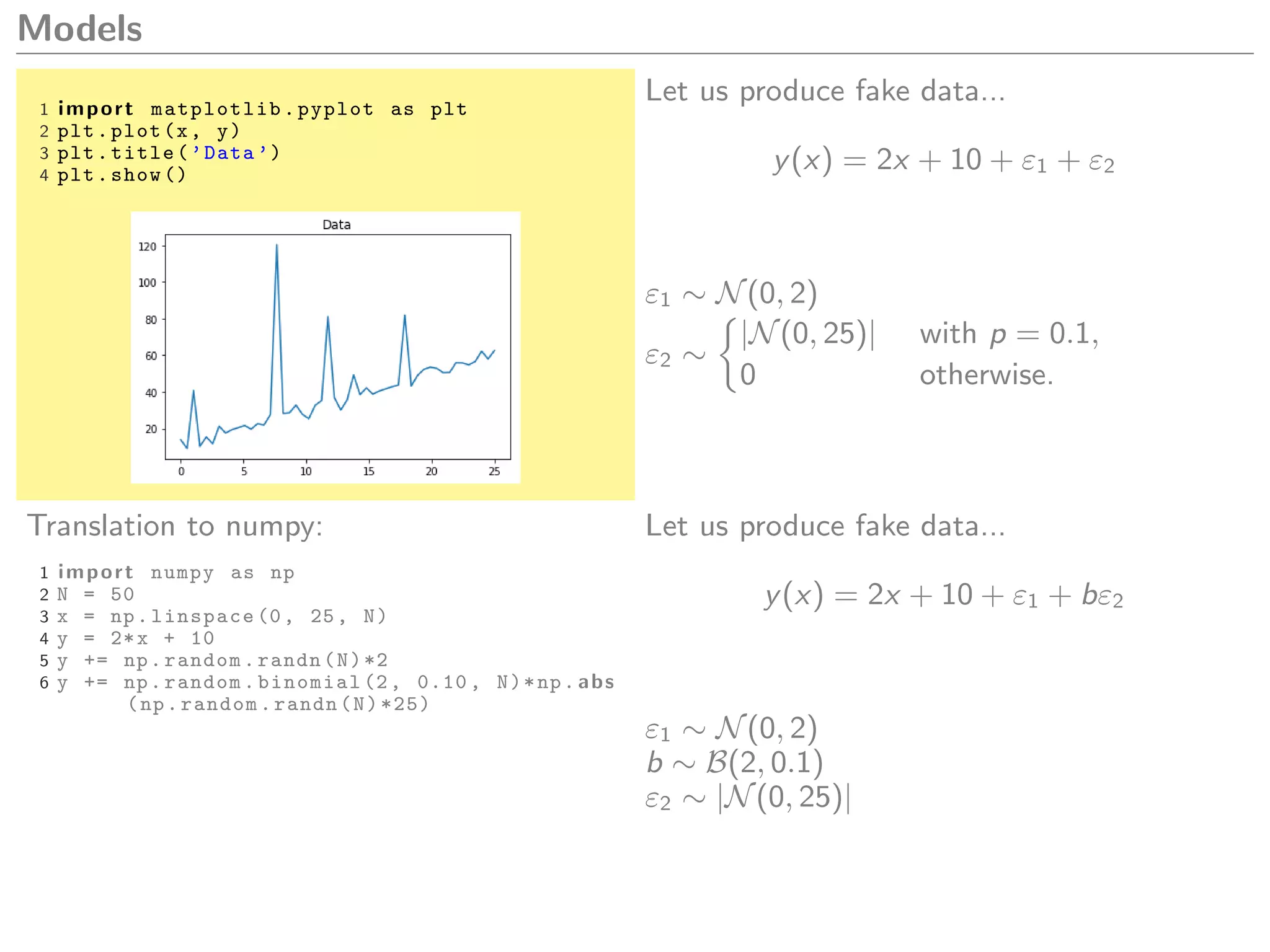
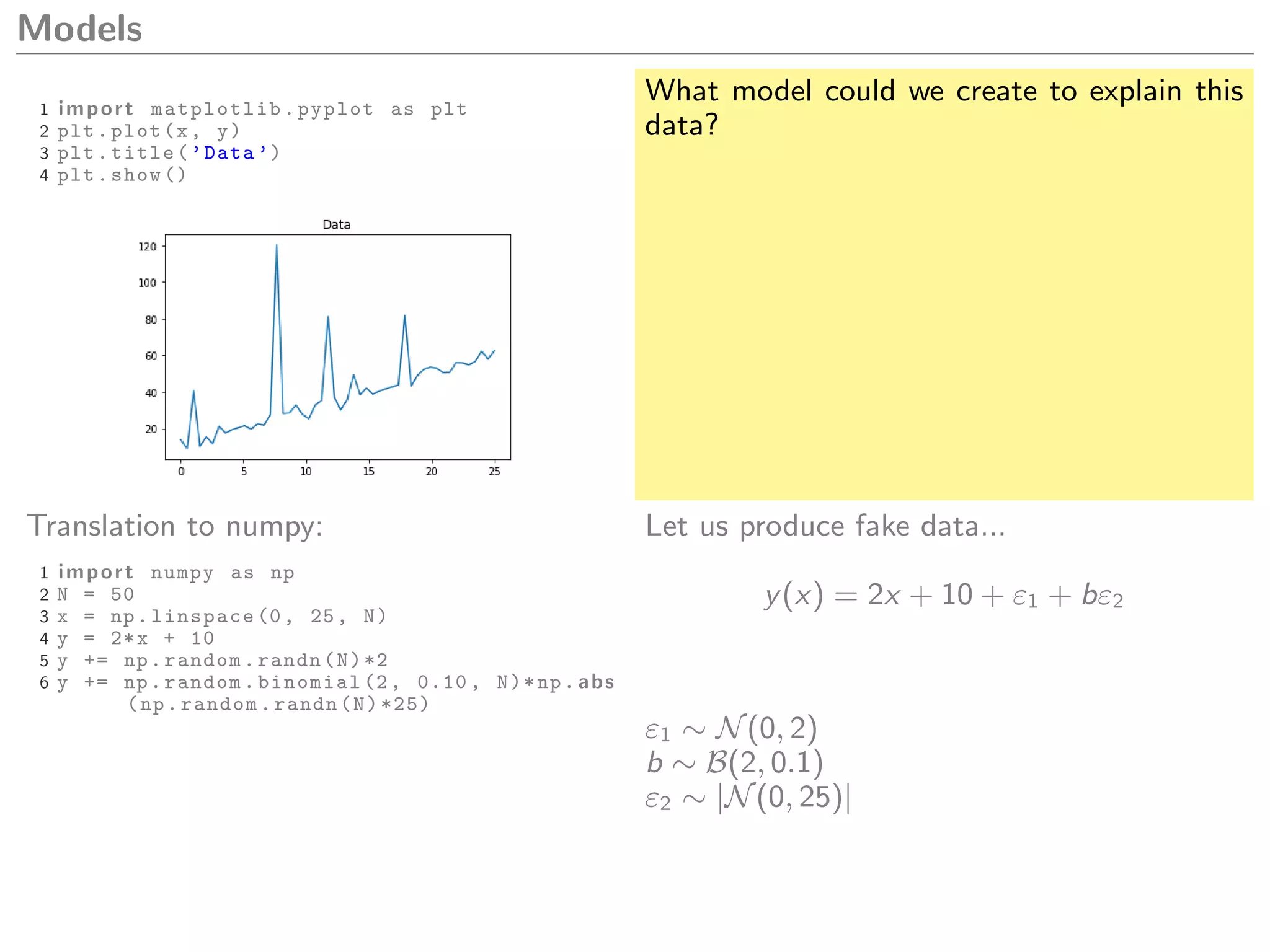
1 import matplotlib.pyplot as plt
2 plt.plot(x, y)
3 plt.title(’Data ’)
4 plt.show ()
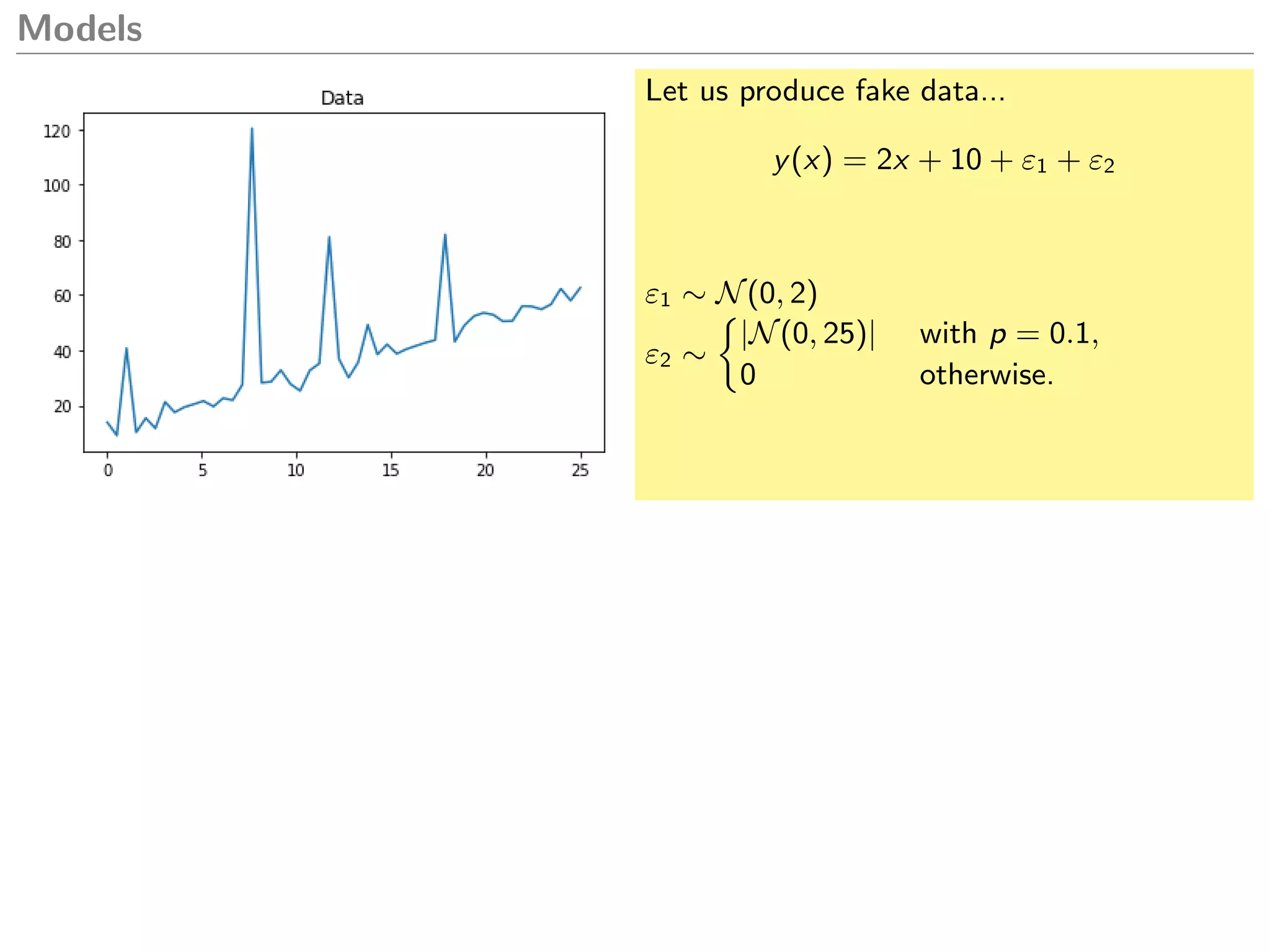
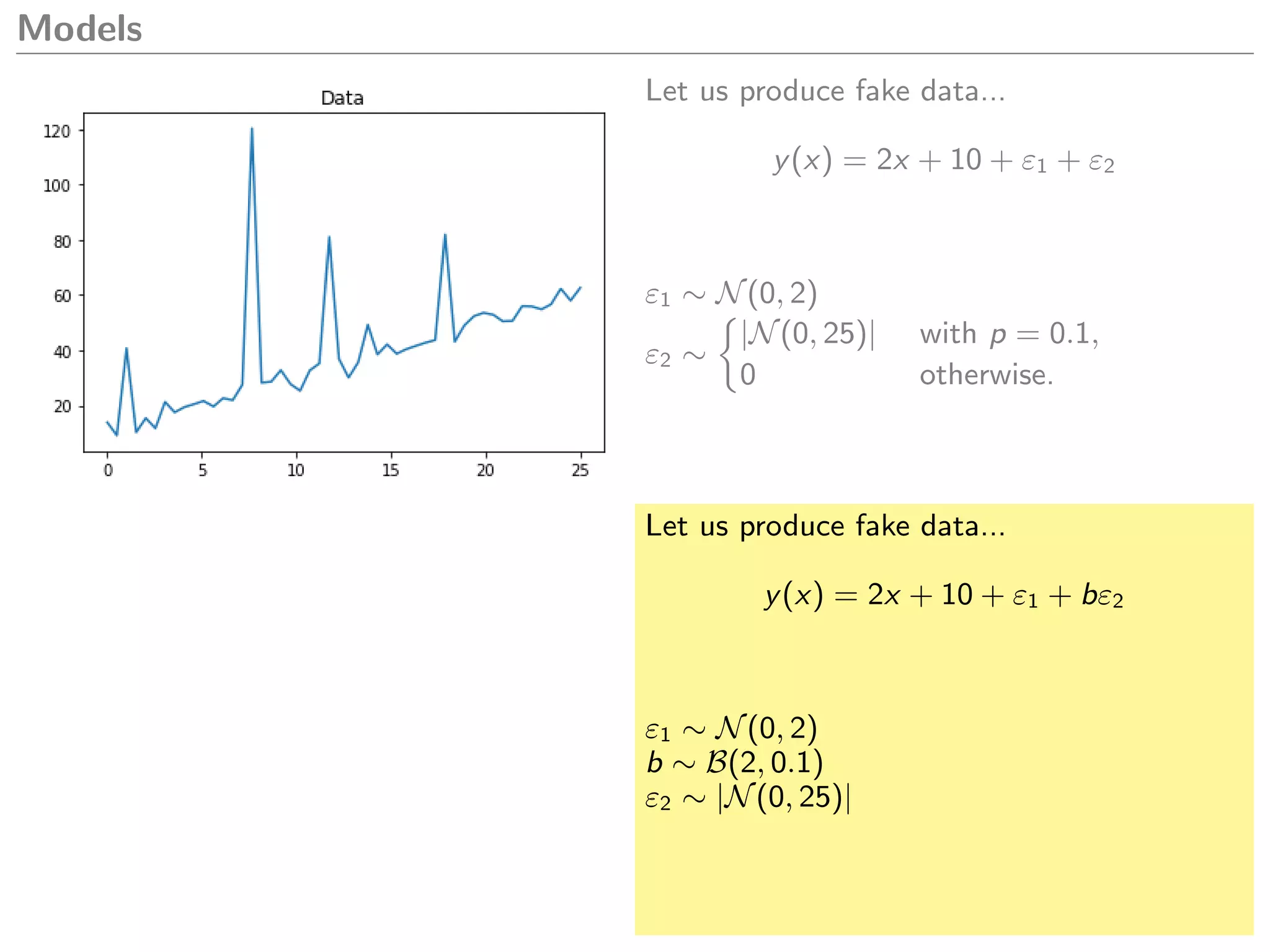
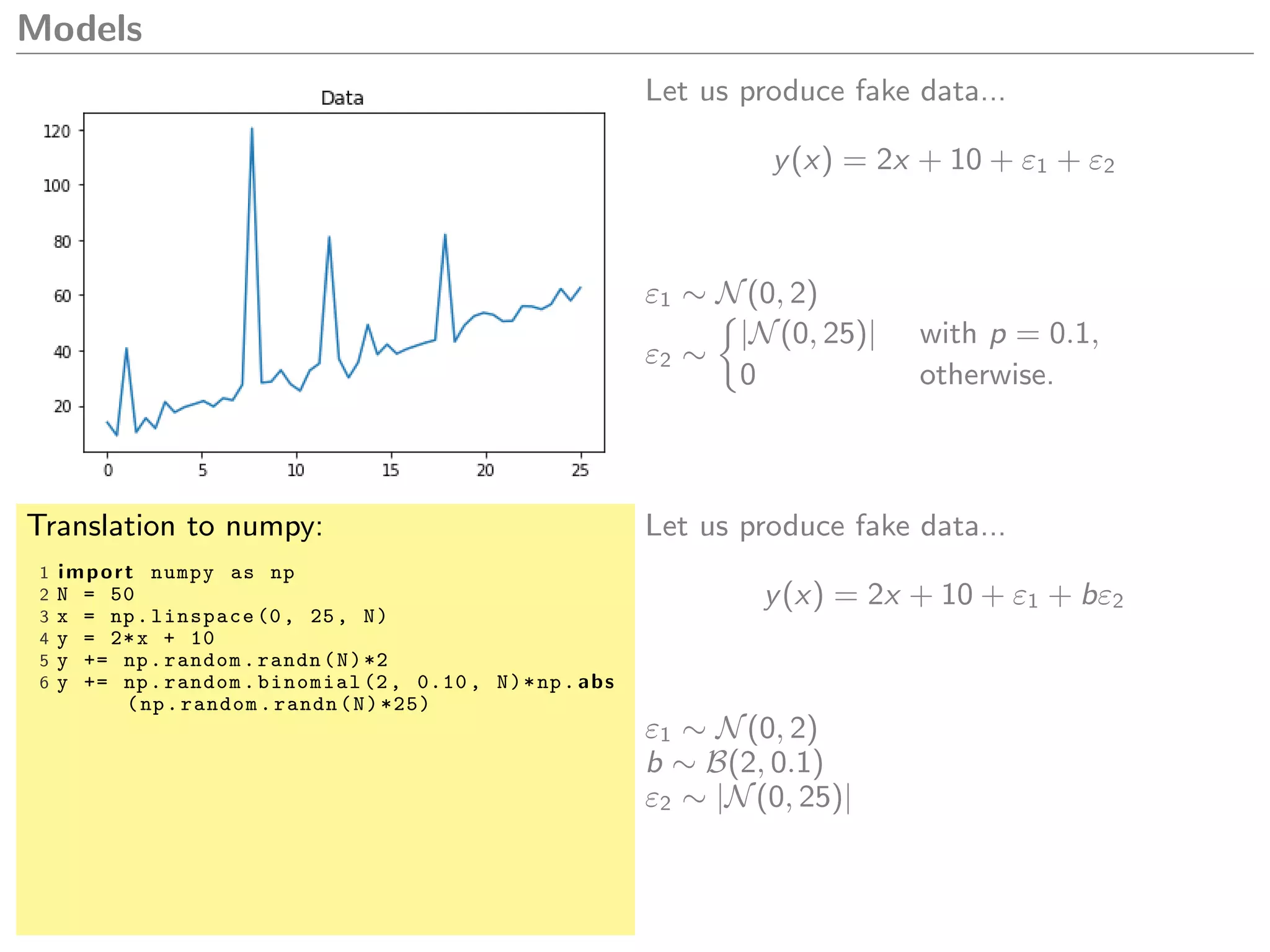
What model could we create to explain this
data?
1 from sklearn. linear_model import
LinearRegression
2 m = LinearRegression ()
3 m.fit(x[:, np.newaxis], y)
4 yp = m.predict(x[:, np.newaxis ])
5
6 plt.plot(x, y)
7 plt.plot(x, yp)
8 plt.title(’Linear regression ’)
9 plt.text(0, 70, ’m=%.1f b=%.1f’ % (m.coef_
[0], m.intercept_))
10 plt.show ()
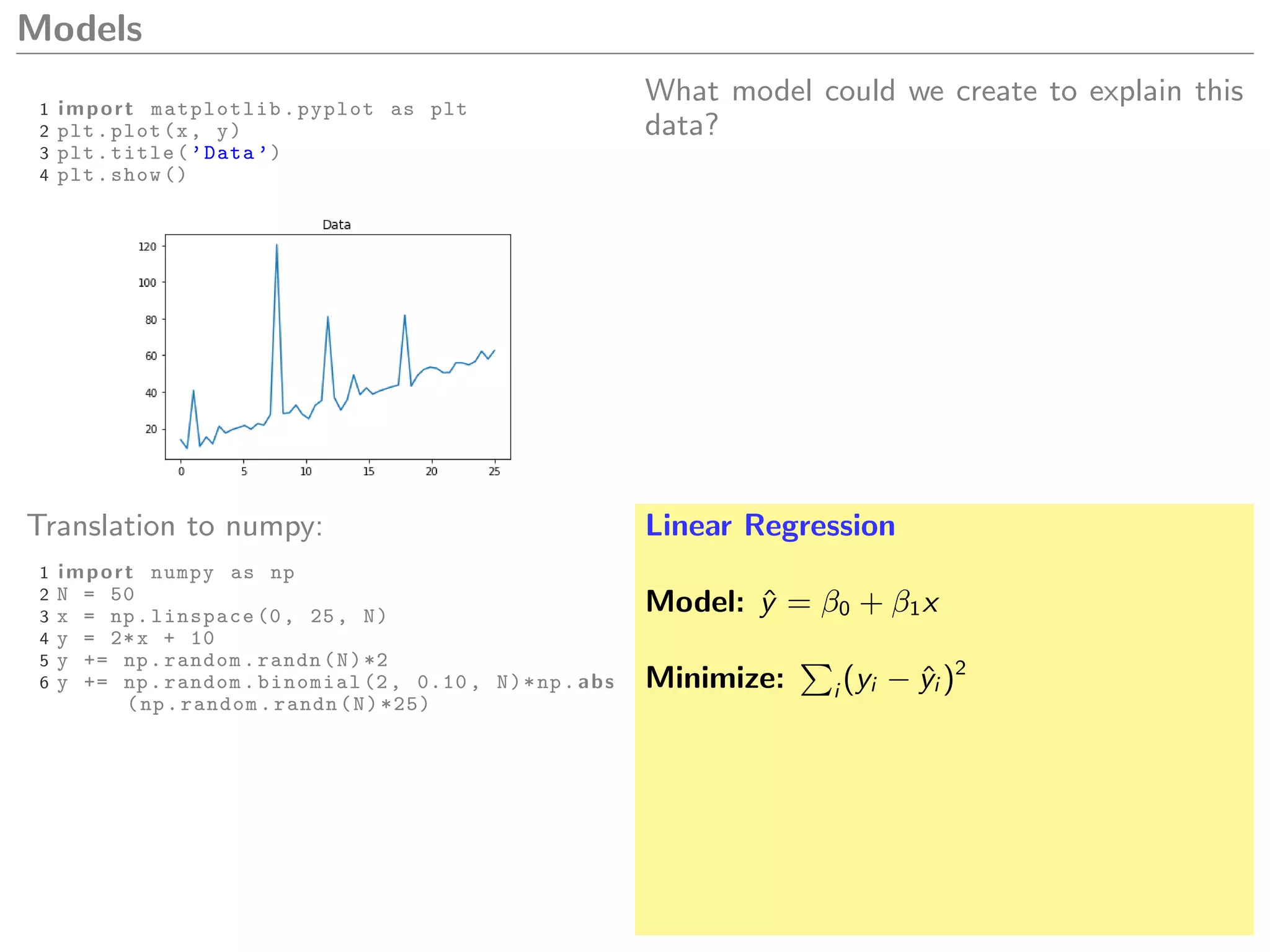
Linear Regression
Model: ˆy = β0 + β1x
Minimize: i (yi − ˆyi )2](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-32-2048.jpg)
![Models
What model could we create to explain this
data?
1 from sklearn. linear_model import
LinearRegression
2 m = LinearRegression ()
3 m.fit(x[:, np.newaxis], y)
4 yp = m.predict(x[:, np.newaxis ])
5
6 plt.plot(x, y)
7 plt.plot(x, yp)
8 plt.title(’Linear regression ’)
9 plt.text(0, 70, ’m=%.1f b=%.1f’ % (m.coef_
[0], m.intercept_))
10 plt.show ()
Linear Regression
Model: ˆy = β0 + β1x
Minimize: i (yi − ˆyi )2](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-33-2048.jpg)
![Models
y(x) = 2x + 10 + ε1 + bε2
ˆy(x) = 2x + 18
What if I want to explain only the trend?
How can I avoid the impact of these spikes?
1 from sklearn. linear_model import
LinearRegression
2 m = LinearRegression ()
3 m.fit(x[:, np.newaxis], y)
4 yp = m.predict(x[:, np.newaxis ])
5
6 plt.plot(x, y)
7 plt.plot(x, yp)
8 plt.title(’Linear regression ’)
9 plt.text(0, 70, ’m=%.1f b=%.1f’ % (m.coef_
[0], m.intercept_))
10 plt.show ()
Linear Regression
Model: ˆy = β0 + β1x
Minimize: i (yi − ˆyi )2](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-34-2048.jpg)
![Models
y(x) = 2x + 10 + ε1 + bε2
ˆy(x) = 2x + 18
What if I want to explain only the trend?
How can I avoid the impact of these spikes?
1 from sklearn. linear_model import
LinearRegression
2 m = LinearRegression ()
3 m.fit(x[:, np.newaxis], y)
4 yp = m.predict(x[:, np.newaxis ])
5
6 plt.plot(x, y)
7 plt.plot(x, yp)
8 plt.title(’Linear regression ’)
9 plt.text(0, 70, ’m=%.1f b=%.1f’ % (m.coef_
[0], m.intercept_))
10 plt.show ()
What would a statistician do?
1 res = yp -y
2 plt.boxplot(res)
3 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-35-2048.jpg)
![Models
y(x) = 2x + 10 + ε1 + bε2
ˆy(x) = 2x + 18
What if I want to explain only the trend?
How can I avoid the impact of these spikes?
1 q1 = np.percentile(res , 25)
2 q3 = np.percentile(res , 75)
3 t = np.logical_and(res > q1 , res < q3)
4 x2 = x[t]
5 y2 = y[t]
6
7 m = LinearRegression ()
8 m.fit(x2[:, np.newaxis], y2)
9 yp = m.predict(x[:, np.newaxis ])
What would a statistician do?
1 res = yp -y
2 plt.boxplot(res)
3 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-36-2048.jpg)
![Models
y(x) = 2x + 10 + ε1 + bε2
ˆy(x) = 2x + 18
What if I want to explain only the trend?
How can I avoid the impact of these spikes?
1 q1 = np.percentile(res , 25)
2 q3 = np.percentile(res , 75)
3 t = np.logical_and(res > q1 , res < q3)
4 x2 = x[t]
5 y2 = y[t]
6
7 m = LinearRegression ()
8 m.fit(x2[:, np.newaxis], y2)
9 yp = m.predict(x[:, np.newaxis ])
What would a statistician do?
1 res = yp -y
2 plt.boxplot(res)
3 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-37-2048.jpg)
![Models
Approach #2: What would a statistician
with some computer science knowledge do?
1 q1 = np.percentile(res , 25)
2 q3 = np.percentile(res , 75)
3 t = np.logical_and(res > q1 , res < q3)
4 x2 = x[t]
5 y2 = y[t]
6
7 m = LinearRegression ()
8 m.fit(x2[:, np.newaxis], y2)
9 yp = m.predict(x[:, np.newaxis ])
What would a statistician do?
1 res = yp -y
2 plt.boxplot(res)
3 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-38-2048.jpg)
![Models
Approach #2: What would a statistician
with some computer science knowledge do?
1 q1 = np.percentile(res , 25)
2 q3 = np.percentile(res , 75)
3 t = np.logical_and(res > q1 , res < q3)
4 x2 = x[t]
5 y2 = y[t]
6
7 m = LinearRegression ()
8 m.fit(x2[:, np.newaxis], y2)
9 yp = m.predict(x[:, np.newaxis ])
Model: ˆy = β0 + β1x
Minimize: i |yi − ˆyi |](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-39-2048.jpg)
![Models
Approach #2: What would a statistician
with some computer science knowledge do?
1 from statsmodels.regression.
quantile_regression import QuantReg
2
3 m = QuantReg(y, np.c_[np.ones(N), x])
4 m = m.fit (0.5)
5 yp = m.predict ()
Model: ˆy = β0 + β1x
Minimize: i |yi − ˆyi |](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-40-2048.jpg)
![Models
Approach #2: What would a statistician
with some computer science knowledge do?
1 from statsmodels.regression.
quantile_regression import QuantReg
2
3 m = QuantReg(y, np.c_[np.ones(N), x])
4 m = m.fit (0.5)
5 yp = m.predict ()
Model: ˆy = β0 + β1x
Minimize: i |yi − ˆyi |](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-41-2048.jpg)
![Models
Approach #3: What would a crazy com-
puter scientist do?
1 from statsmodels.regression.
quantile_regression import QuantReg
2
3 m = QuantReg(y, np.c_[np.ones(N), x])
4 m = m.fit (0.5)
5 yp = m.predict ()
Model: ˆy = β0 + β1x
Minimize: i |yi − ˆyi |](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-42-2048.jpg)
![Models
Approach #3: What would a crazy com-
puter scientist do?
1 from statsmodels.regression.
quantile_regression import QuantReg
2
3 m = QuantReg(y, np.c_[np.ones(N), x])
4 m = m.fit (0.5)
5 yp = m.predict ()
1 plt.plot(x, y)
2 f o r it i n range (10):
3 t = np.random.choice(N, N//10 , replace
=False)
4 x2 = x[t]
5 y2 = y[t]
6 m.fit(x2[:, np.newaxis], y2)
7 yp = m.predict(x[:, np.newaxis ])
8 plt.plot(x, yp , color=’black ’, alpha
=0.4)
9 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-43-2048.jpg)
![Models
Approach #3: What would a crazy com-
puter scientist do?
1 plt.plot(x, y)
2 f o r it i n range (10):
3 t = np.random.choice(N, N//10 , replace
=False)
4 x2 = x[t]
5 y2 = y[t]
6 m.fit(x2[:, np.newaxis], y2)
7 yp = m.predict(x[:, np.newaxis ])
8 plt.plot(x, yp , color=’black ’, alpha
=0.4)
9 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-44-2048.jpg)
![Models
Sklearn already comes with this crazy model
too:
1 from sklearn. linear_model import
RANSACRegressor
2 m = RANSACRegressor ()
3 m.fit(x[:, np.newaxis], y)
4
5 plt.plot(x, y)
6 plt.plot(x, m.predict(x[:, np.newaxis ]))
7 plt.title(’RANSAC ’)
8 plt.show ()
Approach #3: What would a crazy com-
puter scientist do?
1 plt.plot(x, y)
2 f o r it i n range (10):
3 t = np.random.choice(N, N//10 , replace
=False)
4 x2 = x[t]
5 y2 = y[t]
6 m.fit(x2[:, np.newaxis], y2)
7 yp = m.predict(x[:, np.newaxis ])
8 plt.plot(x, yp , color=’black ’, alpha
=0.4)
9 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-45-2048.jpg)
![Models
Sklearn already comes with this crazy model
too:
1 from sklearn. linear_model import
RANSACRegressor
2 m = RANSACRegressor ()
3 m.fit(x[:, np.newaxis], y)
4
5 plt.plot(x, y)
6 plt.plot(x, m.predict(x[:, np.newaxis ]))
7 plt.title(’RANSAC ’)
8 plt.show ()
1 plt.plot(x, y)
2 f o r it i n range (10):
3 t = np.random.choice(N, N//10 , replace
=False)
4 x2 = x[t]
5 y2 = y[t]
6 m.fit(x2[:, np.newaxis], y2)
7 yp = m.predict(x[:, np.newaxis ])
8 plt.plot(x, yp , color=’black ’, alpha
=0.4)
9 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-46-2048.jpg)








![Text Mining
1 import tweepy
2 auth = tweepy. OAuthHandler (api_key ,
api_secret)
3 auth. set_access_token (access_token ,
access_secret )
4 api = tweepy.API(auth)
5
6 timeline = api. user_timeline (’
realDonaldTrump ’, count =100)
7 texts = [tweet.text f o r tweet i n timeline]](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-55-2048.jpg)
![Text Mining
1 import tweepy
2 auth = tweepy. OAuthHandler (api_key ,
api_secret)
3 auth. set_access_token (access_token ,
access_secret )
4 api = tweepy.API(auth)
5
6 timeline = api. user_timeline (’
realDonaldTrump ’, count =100)
7 texts = [tweet.text f o r tweet i n timeline]
1 from sklearn. feature_extraction .text
import CountVectorizer
2 m = CountVectorizer (stop_words=’english ’,
min_df =5, max_df =16)
3 X = m. fit_transform (texts)
4 words = sorted (m.vocabulary_ , key=m.
vocabulary_.get)
5
6 import pandas as pd
7 p r i n t (pd.DataFrame(X.todense (), columns=
words).ix[:5, :5]. to_latex ())
america big comey day dems
0 0 0 0 0 0
1 0 1 0 0 0
2 1 0 0 0 0](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-56-2048.jpg)
![Text Mining
1 import tweepy
2 auth = tweepy. OAuthHandler (api_key ,
api_secret)
3 auth. set_access_token (access_token ,
access_secret )
4 api = tweepy.API(auth)
5
6 timeline = api. user_timeline (’
realDonaldTrump ’, count =100)
7 texts = [tweet.text f o r tweet i n timeline]
1 from sklearn. feature_extraction .text
import CountVectorizer
2 m = CountVectorizer (stop_words=’english ’,
min_df =5, max_df =16)
3 X = m. fit_transform (texts)
4 words = sorted (m.vocabulary_ , key=m.
vocabulary_.get)
5
6 import pandas as pd
7 p r i n t (pd.DataFrame(X.todense (), columns=
words).ix[:5, :5]. to_latex ())
america big comey day dems
0 0 0 0 0 0
1 0 1 0 0 0
2 1 0 0 0 0
1 import matplotlib.pyplot as plt
2 counts = np.asarray(X.sum(0))[0]
3 plt.barh( range ( len (counts)), counts)
4 plt.xticks( range (0, 14, 2))
5 plt.yticks( range ( len (counts)), words)
6 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-57-2048.jpg)
![Text Mining
1 import tweepy
2 auth = tweepy. OAuthHandler (api_key ,
api_secret)
3 auth. set_access_token (access_token ,
access_secret )
4 api = tweepy.API(auth)
5
6 timeline = api. user_timeline (’
realDonaldTrump ’, count =100)
7 texts = [tweet.text f o r tweet i n timeline]
1 from sklearn. feature_extraction .text
import CountVectorizer
2 m = CountVectorizer (stop_words=’english ’,
min_df =5, max_df =16)
3 X = m. fit_transform (texts)
4 words = sorted (m.vocabulary_ , key=m.
vocabulary_.get)
5
6 import pandas as pd
7 p r i n t (pd.DataFrame(X.todense (), columns=
words).ix[:5, :5]. to_latex ())
america big comey day dems
0 0 0 0 0 0
1 0 1 0 0 0
2 1 0 0 0 0
1 import matplotlib.pyplot as plt
2 counts = np.asarray(X.sum(0))[0]
3 plt.barh( range ( len (counts)), counts)
4 plt.xticks( range (0, 14, 2))
5 plt.yticks( range ( len (counts)), words)
6 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-58-2048.jpg)
![Text Mining
1 from sklearn. decomposition import
LatentDirichletAllocation
2 lda = LatentDirichletAllocation (2,
learning_method =’online ’)
3 lda.fit(X)
4 topics = lda. components_
newword1 = β11word1 + β12word2 + . . .
newword2 = β21word1 + β22word2 + . . .
1 from sklearn. feature_extraction .text
import CountVectorizer
2 m = CountVectorizer (stop_words=’english ’,
min_df =5, max_df =16)
3 X = m. fit_transform (texts)
4 words = sorted (m.vocabulary_ , key=m.
vocabulary_.get)
5
6 import pandas as pd
7 p r i n t (pd.DataFrame(X.todense (), columns=
words).ix[:5, :5]. to_latex ())
america big comey day dems
0 0 0 0 0 0
1 0 1 0 0 0
2 1 0 0 0 0
1 import matplotlib.pyplot as plt
2 counts = np.asarray(X.sum(0))[0]
3 plt.barh( range ( len (counts)), counts)
4 plt.xticks( range (0, 14, 2))
5 plt.yticks( range ( len (counts)), words)
6 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-59-2048.jpg)
![Text Mining
1 from sklearn. decomposition import
LatentDirichletAllocation
2 lda = LatentDirichletAllocation (2,
learning_method =’online ’)
3 lda.fit(X)
4 topics = lda. components_
newword1 = β11word1 + β12word2 + . . .
newword2 = β21word1 + β22word2 + . . .
1 topics = topics / topics.max(1)[:, np.
newaxis]
2 topics += np.random.randn (* topics.shape)
*0.02
3 f o r i, word i n enumerate(words):
4 plt.text(topics [0, i], topics [1, i],
word , ha=’center ’)
5 plt.show ()
1 import matplotlib.pyplot as plt
2 counts = np.asarray(X.sum(0))[0]
3 plt.barh( range ( len (counts)), counts)
4 plt.xticks( range (0, 14, 2))
5 plt.yticks( range ( len (counts)), words)
6 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-60-2048.jpg)
![Text Mining
1 from sklearn. decomposition import
LatentDirichletAllocation
2 lda = LatentDirichletAllocation (2,
learning_method =’online ’)
3 lda.fit(X)
4 topics = lda. components_
newword1 = β11word1 + β12word2 + . . .
newword2 = β21word1 + β22word2 + . . .
1 topics = topics / topics.max(1)[:, np.
newaxis]
2 topics += np.random.randn (* topics.shape)
*0.02
3 f o r i, word i n enumerate(words):
4 plt.text(topics [0, i], topics [1, i],
word , ha=’center ’)
5 plt.show ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-61-2048.jpg)
![Text Mining
1 from sklearn. decomposition import
LatentDirichletAllocation
2 lda = LatentDirichletAllocation (2,
learning_method =’online ’)
3 lda.fit(X)
4 topics = lda. components_
newword1 = β11word1 + β12word2 + . . .
newword2 = β21word1 + β22word2 + . . .
1 topics = topics / topics.max(1)[:, np.
newaxis]
2 topics += np.random.randn (* topics.shape)
*0.02
3 f o r i, word i n enumerate(words):
4 plt.text(topics [0, i], topics [1, i],
word , ha=’center ’)
5 plt.show ()
1 timeline = api. user_timeline (’
marcelorebelo_ ’, count =100)](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-62-2048.jpg)

![Traditional Learning
Cats vs Dogs – Kaggle Competition – https:
//www.kaggle.com/c/dogs-vs-cats
25,000 images of cats and dogs
Feature #1: Extract histogram of colors
1 from skimage.io import imread
2 from skimage.transform import rgb2gray
3
4 f o r filename i n os.listdir(’train ’):
5 im = imread(os.path.join(’train ’,
filename))
6 im = rgb2gray(im)
7 f1 = np.histogram(im.flatten (), 10) [0]
8 f1 = (f1/f1.sum()).cumsum ()](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-65-2048.jpg)
![Traditional Learning
Cats vs Dogs – Kaggle Competition – https:
//www.kaggle.com/c/dogs-vs-cats
25,000 images of cats and dogs
Feature #1: Extract histogram of colors
1 from skimage.io import imread
2 from skimage.transform import rgb2gray
3
4 f o r filename i n os.listdir(’train ’):
5 im = imread(os.path.join(’train ’,
filename))
6 im = rgb2gray(im)
7 f1 = np.histogram(im.flatten (), 10) [0]
8 f1 = (f1/f1.sum()).cumsum ()
Feature #2: Histogram of Oriented Gradi-
ents
1 im2 = resize(im , (32, 32) , mode=’reflect
’)
2 im2 = np.sqrt(im2)
3 f2 = hog(im2 , block_norm=’L2 -Hys ’)](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-66-2048.jpg)
![Traditional Learning
Cats vs Dogs – Kaggle Competition – https:
//www.kaggle.com/c/dogs-vs-cats
25,000 images of cats and dogs
Feature #1: Extract histogram of colors
1 from skimage.io import imread
2 from skimage.transform import rgb2gray
3
4 f o r filename i n os.listdir(’train ’):
5 im = imread(os.path.join(’train ’,
filename))
6 im = rgb2gray(im)
7 f1 = np.histogram(im.flatten (), 10) [0]
8 f1 = (f1/f1.sum()).cumsum ()
1 from sklearn.tree import
DecisionTreeClassifier ,
export_graphviz
2 m = DecisionTreeClassifier (max_depth =3)
3 m.fit(X, y)
Feature #2: Histogram of Oriented Gradi-
ents
1 im2 = resize(im , (32, 32) , mode=’reflect
’)
2 im2 = np.sqrt(im2)
3 f2 = hog(im2 , block_norm=’L2 -Hys ’)](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-67-2048.jpg)
![Traditional Learning
1 from sklearn. model_selection import
cross_val_score
2 from sklearn.ensemble import
RandomForestClassifier
3 p r i n t ( cross_val_score (
RandomForestClassifier (100) , X, y))
1 [ 0.69642429 0.70086393 0.69851176]
Feature #1: Extract histogram of colors
1 from skimage.io import imread
2 from skimage.transform import rgb2gray
3
4 f o r filename i n os.listdir(’train ’):
5 im = imread(os.path.join(’train ’,
filename))
6 im = rgb2gray(im)
7 f1 = np.histogram(im.flatten (), 10) [0]
8 f1 = (f1/f1.sum()).cumsum ()
1 from sklearn.tree import
DecisionTreeClassifier ,
export_graphviz
2 m = DecisionTreeClassifier (max_depth =3)
3 m.fit(X, y)
Feature #2: Histogram of Oriented Gradi-
ents
1 im2 = resize(im , (32, 32) , mode=’reflect
’)
2 im2 = np.sqrt(im2)
3 f2 = hog(im2 , block_norm=’L2 -Hys ’)](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-68-2048.jpg)
![Deep Learning
1 from sklearn. model_selection import
cross_val_score
2 from sklearn.ensemble import
RandomForestClassifier
3 p r i n t ( cross_val_score (
RandomForestClassifier (100) , X, y))
1 [ 0.69642429 0.70086393 0.69851176]
Linear Regression
ˆy = β0 + β1x1 + β2x2 + . . .
Multilayer perceptron / neural network
ˆy = β00σ(β10 + β11x1 + β12x2 + . . . )
+ β01σ(β20 + β21x1 + β22x2 + . . . ) + . . .
1 from sklearn.tree import
DecisionTreeClassifier ,
export_graphviz
2 m = DecisionTreeClassifier (max_depth =3)
3 m.fit(X, y)
Feature #2: Histogram of Oriented Gradi-
ents
1 im2 = resize(im , (32, 32) , mode=’reflect
’)
2 im2 = np.sqrt(im2)
3 f2 = hog(im2 , block_norm=’L2 -Hys ’)](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-69-2048.jpg)
![Deep Learning
1 from sklearn. model_selection import
cross_val_score
2 from sklearn.ensemble import
RandomForestClassifier
3 p r i n t ( cross_val_score (
RandomForestClassifier (100) , X, y))
1 [ 0.69642429 0.70086393 0.69851176]
Linear Regression
ˆy = β0 + β1x1 + β2x2 + . . .
Multilayer perceptron / neural network
ˆy = β00σ(β10 + β11x1 + β12x2 + . . . )
+ β01σ(β20 + β21x1 + β22x2 + . . . ) + . . .
1 from sklearn.tree import
DecisionTreeClassifier ,
export_graphviz
2 m = DecisionTreeClassifier (max_depth =3)
3 m.fit(X, y)](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-70-2048.jpg)
![Deep Learning
1 from sklearn. model_selection import
cross_val_score
2 from sklearn.ensemble import
RandomForestClassifier
3 p r i n t ( cross_val_score (
RandomForestClassifier (100) , X, y))
1 [ 0.69642429 0.70086393 0.69851176]
Linear Regression
ˆy = β0 + β1x1 + β2x2 + . . .
Multilayer perceptron / neural network
ˆy = β00σ(β10 + β11x1 + β12x2 + . . . )
+ β01σ(β20 + β21x1 + β22x2 + . . . ) + . . .
1 model = Sequential ()
2 model.add(Conv2D (8, 3, 1, activation=’relu
’, input_shape =(32 , 32, 1)))
3 model.add( MaxPooling2D ())
4 model.add(Conv2D (16, 3, 1, activation=’
relu ’))
5 model.add( MaxPooling2D ())
6 model.add(Flatten ())
7 model.add(Dense (16, activation=’relu ’))
8 model.add(Dense (8, activation=’relu ’))
9 model.add(Dense (1, activation=’sigmoid ’))
10
11 sgd = SGD ()
12 model. compile (sgd , ’binary_crossentropy ’)
13
14 model.fit(X[tr], y[tr], validation_data =(X
[ts], y[ts]),
15 epochs =10, batch_size =100)](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-71-2048.jpg)
![Deep Learning
1 f o r tr , ts i n StratifiedKFold ().split(X, y
):
2 model = ...
3 ...
4 yp = (model.predict(X[ts])[:, -1] > 0.5)
.astype( i n t )
5 p r i n t ( accuracy_score (y[ts], yp))
1 [0.57 , 0.57 , 0.63]
Linear Regression
ˆy = β0 + β1x1 + β2x2 + . . .
Multilayer perceptron / neural network
ˆy = β00σ(β10 + β11x1 + β12x2 + . . . )
+ β01σ(β20 + β21x1 + β22x2 + . . . ) + . . .
1 model = Sequential ()
2 model.add(Conv2D (8, 3, 1, activation=’relu
’, input_shape =(32 , 32, 1)))
3 model.add( MaxPooling2D ())
4 model.add(Conv2D (16, 3, 1, activation=’
relu ’))
5 model.add( MaxPooling2D ())
6 model.add(Flatten ())
7 model.add(Dense (16, activation=’relu ’))
8 model.add(Dense (8, activation=’relu ’))
9 model.add(Dense (1, activation=’sigmoid ’))
10
11 sgd = SGD ()
12 model. compile (sgd , ’binary_crossentropy ’)
13
14 model.fit(X[tr], y[tr], validation_data =(X
[ts], y[ts]),
15 epochs =10, batch_size =100)](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-72-2048.jpg)
![Deep Learning
1 f o r tr , ts i n StratifiedKFold ().split(X, y
):
2 model = ...
3 ...
4 yp = (model.predict(X[ts])[:, -1] > 0.5)
.astype( i n t )
5 p r i n t ( accuracy_score (y[ts], yp))
1 [0.57 , 0.57 , 0.63]
Overview of Python deep learning landscape:
Theano TensorFlow PyTorch
KerasLasagne
1 model = Sequential ()
2 model.add(Conv2D (8, 3, 1, activation=’relu
’, input_shape =(32 , 32, 1)))
3 model.add( MaxPooling2D ())
4 model.add(Conv2D (16, 3, 1, activation=’
relu ’))
5 model.add( MaxPooling2D ())
6 model.add(Flatten ())
7 model.add(Dense (16, activation=’relu ’))
8 model.add(Dense (8, activation=’relu ’))
9 model.add(Dense (1, activation=’sigmoid ’))
10
11 sgd = SGD ()
12 model. compile (sgd , ’binary_crossentropy ’)
13
14 model.fit(X[tr], y[tr], validation_data =(X
[ts], y[ts]),
15 epochs =10, batch_size =100)](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-73-2048.jpg)
![Deep Learning
1 f o r tr , ts i n StratifiedKFold ().split(X, y
):
2 model = ...
3 ...
4 yp = (model.predict(X[ts])[:, -1] > 0.5)
.astype( i n t )
5 p r i n t ( accuracy_score (y[ts], yp))
1 [0.57 , 0.57 , 0.63]
Overview of Python deep learning landscape:
Theano TensorFlow PyTorch
KerasLasagne
1 model = Sequential ()
2 model.add(Conv2D (8, 3, 1, activation=’relu
’, input_shape =(32 , 32, 1)))
3 model.add( MaxPooling2D ())
4 model.add(Conv2D (16, 3, 1, activation=’
relu ’))
5 model.add( MaxPooling2D ())
6 model.add(Flatten ())
7 model.add(Dense (16, activation=’relu ’))
8 model.add(Dense (8, activation=’relu ’))
9 model.add(Dense (1, activation=’sigmoid ’))
10
11 sgd = SGD ()
12 model. compile (sgd , ’binary_crossentropy ’)
13
14 model.fit(X[tr], y[tr], validation_data =(X
[ts], y[ts]),
15 epochs =10, batch_size =100)
Deep learning architectures:
Fully connected perceptrons
Convolutional neural networks
Recurrent neural networks
Neural Turing Machines
Autoencoders](https://image.slidesharecdn.com/dm-170713174507/75/Python-for-Scientific-Computing-Ricardo-Cruz-74-2048.jpg)




