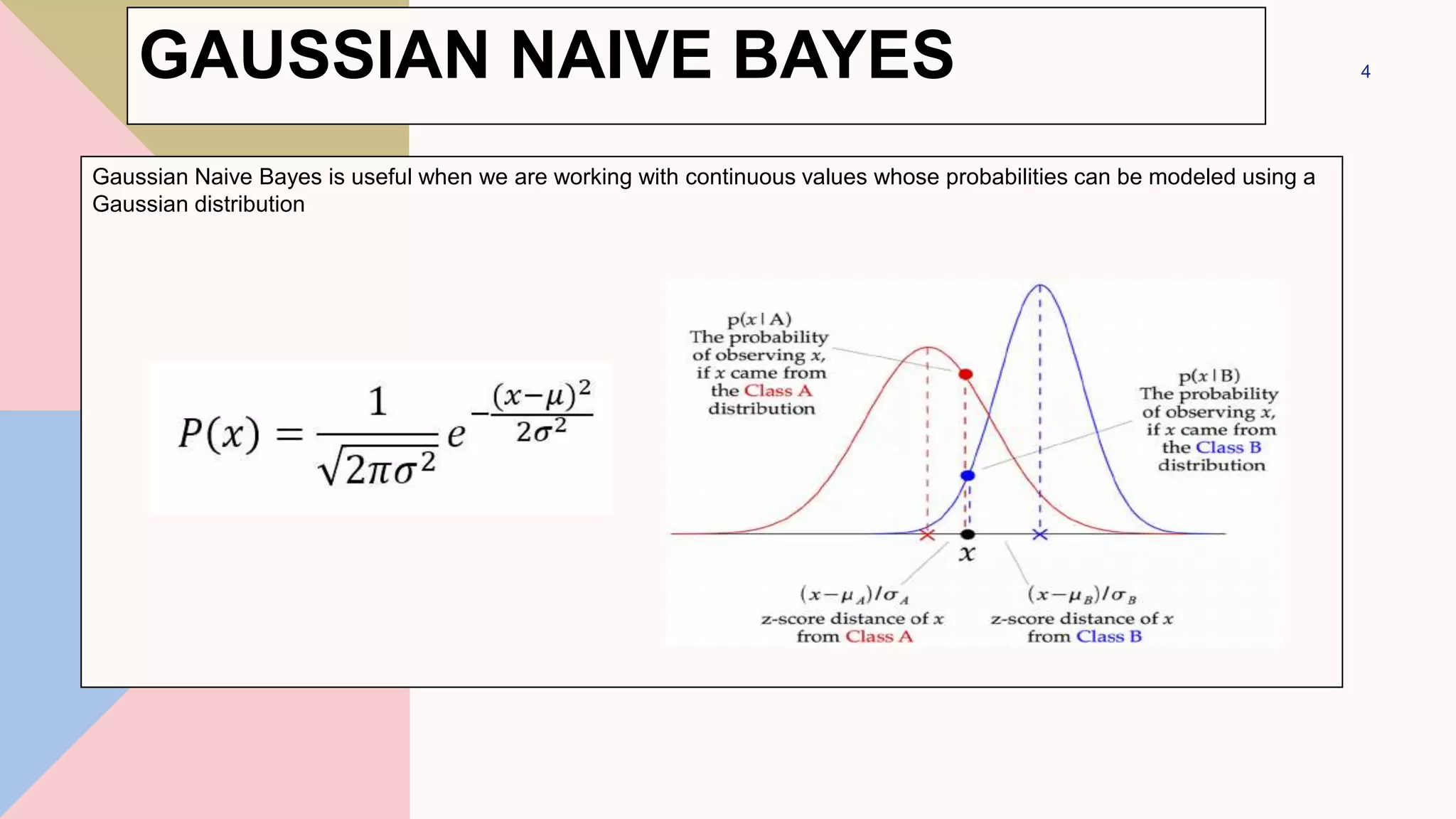
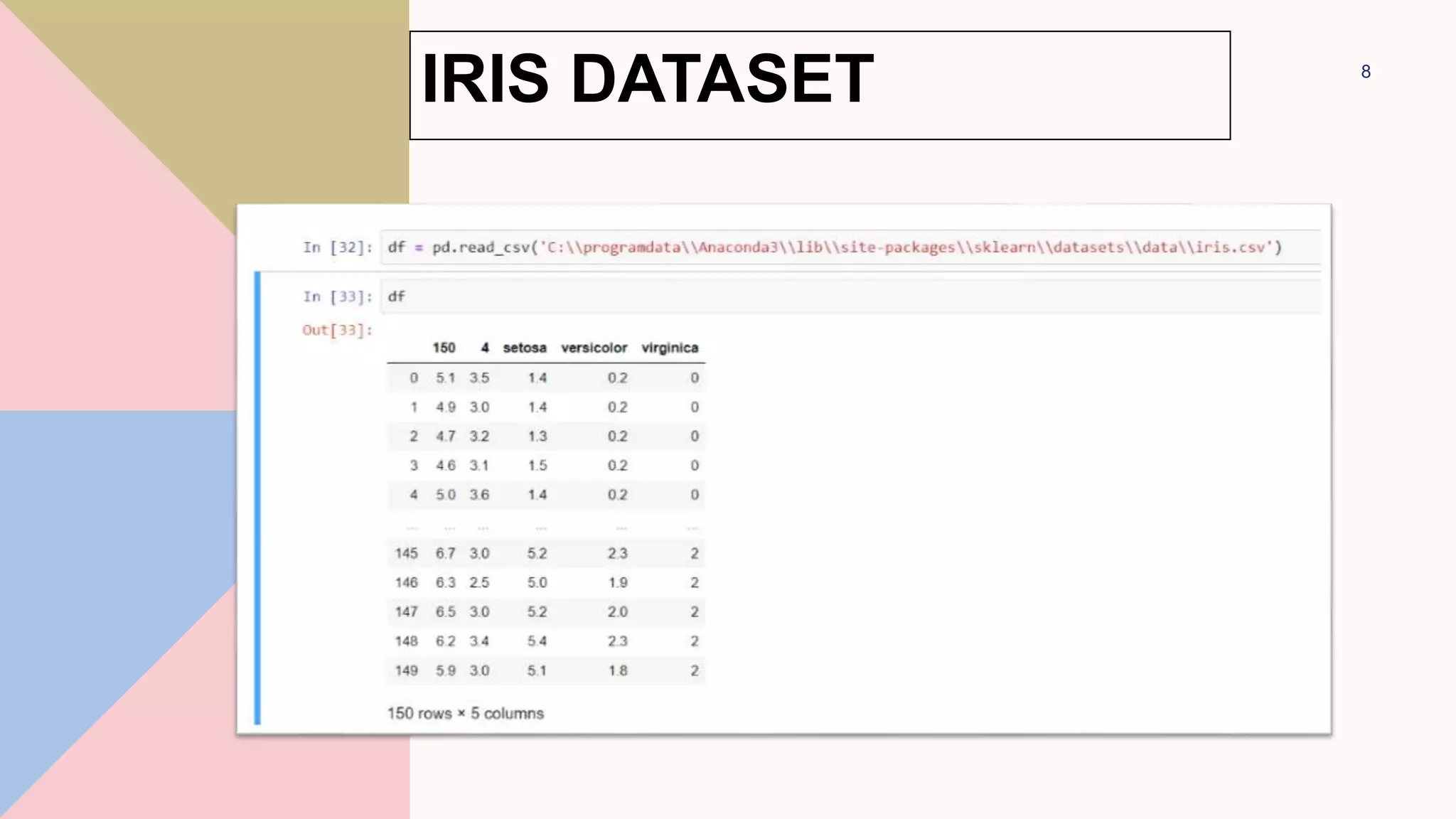
This document discusses different types of Naive Bayes classifiers and provides examples of using each type. The three types are Gaussian Naive Bayes, Multinomial Naive Bayes, and Bernoulli Naive Bayes. Gaussian Naive Bayes is useful for continuous data that can be modeled with a Gaussian distribution. Multinomial Naive Bayes models feature vectors with multiple possible values. Bernoulli Naive Bayes models features that take on only two values. Examples are provided using the Iris dataset and 20 Newsgroups dataset to classify data with Gaussian and Multinomial Naive Bayes classifiers respectively.







![# making predictions on the testing set
y_pred = model.predict(X_test)
# comparing actual response values (y_test) with predicted response
values (y_pred)
from sklearn.metrics import accuracy_score
print(f'Gaussian Naive Bayes model accuracy(in %):={accuracy_score(y_test,
y_pred)*100} %')
res = model.predict([[6.5,3.0,5.2,2.0]])
print(f'Result = {iris.target_names[res[0]]}')
10
GAUSSIAN NAIVE BAYES](https://image.slidesharecdn.com/navebayes-230201064135-58c1858c/75/Naive-Bayes-pptx-10-2048.jpg)
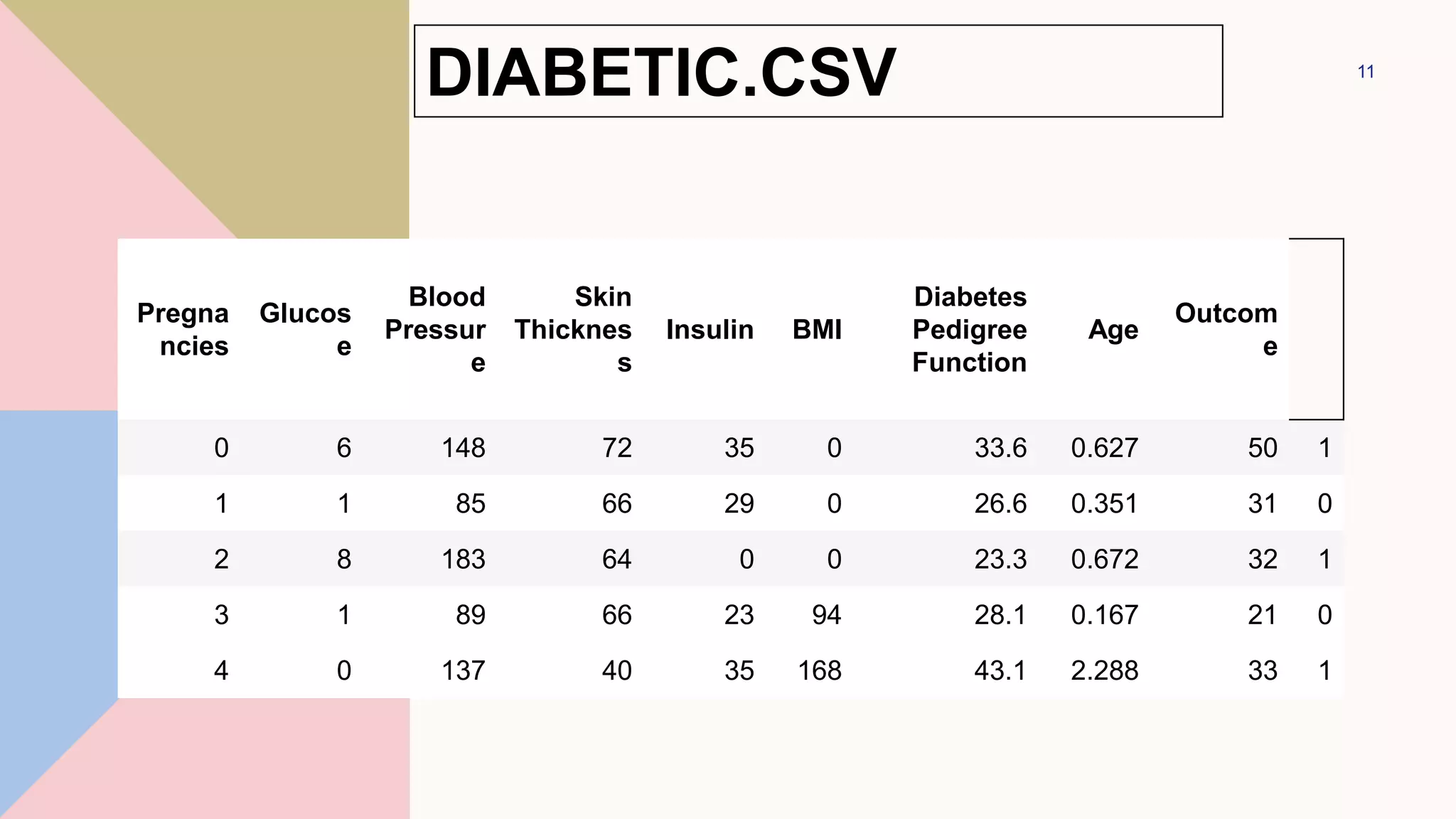
![diabetes = pd.read_csv("diabetes.csv")
diabetes.columns
Output-
12
Index(['Pregnancies', 'Glucose', 'BloodPressure',
'SkinThickness', 'Insulin', 'BMI',
'DiabetesPedigreeFunction', 'Age', 'Outcome'],
dtype='object')
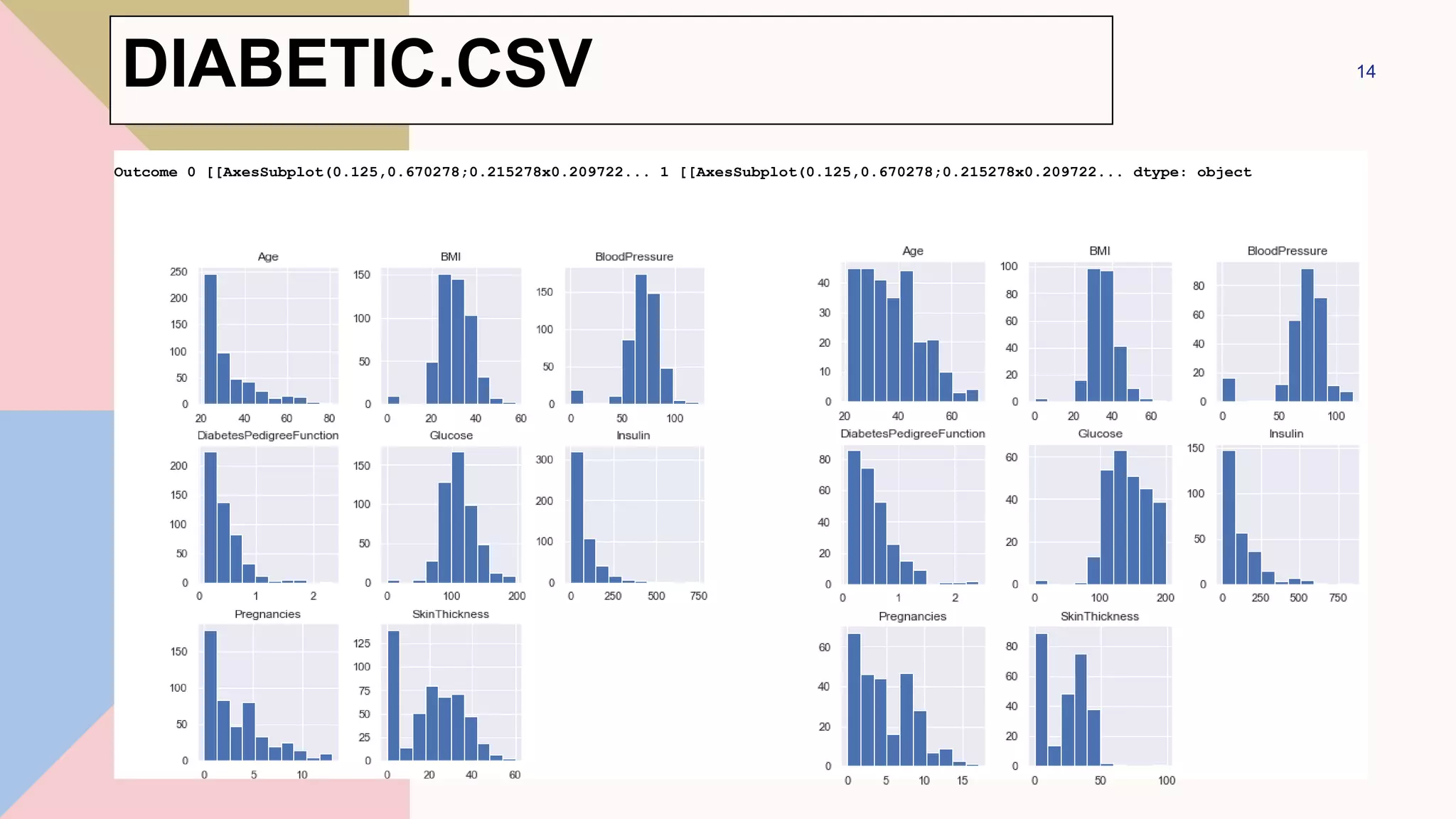
DIABETIC.CSV](https://image.slidesharecdn.com/navebayes-230201064135-58c1858c/75/Naive-Bayes-pptx-12-2048.jpg)


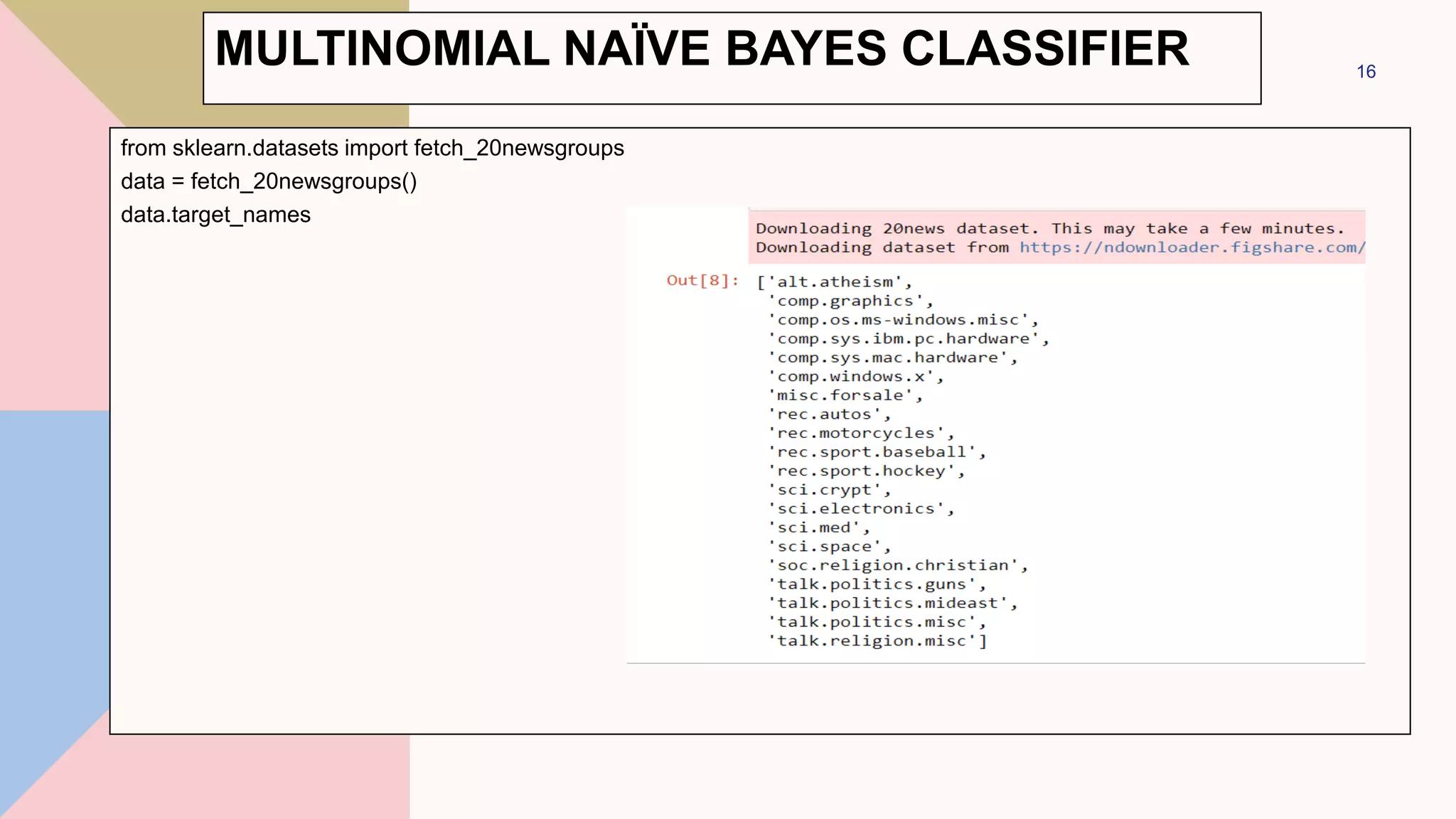
![categories = ['talk.religion.misc', 'soc.religion.christian’,
'sci.space', 'comp.graphics']
train = fetch_20newsgroups(subset='train', categories=categories)
test = fetch_20newsgroups(subset='test', categories=categories)
17
Multinomial Naïve Bayes Classifier](https://image.slidesharecdn.com/navebayes-230201064135-58c1858c/75/Naive-Bayes-pptx-17-2048.jpg)




![def predict_category(s, train=train, model=model):
pred = model.predict([s])
return train.target_names[pred[0]]
predict_category('sending a payload to the ISS’)
Output-
22
'sci.space'
Multinomial Naïve Bayes Classifier](https://image.slidesharecdn.com/navebayes-230201064135-58c1858c/75/Naive-Bayes-pptx-22-2048.jpg)

![import matplotlib.pyplot as plt
import numpy as np
fig, ax = plt.subplots()
xmin, xmax = -0.2, 1.4
X = np.arange(xmin, xmax, 0.1)
ax.scatter(0, 0, color="r")
ax.scatter(0, 1, color="r")
ax.scatter(1, 0, color="r")
ax.scatter(1, 1, color="g")
ax.set_xlim([xmin, xmax])
ax.set_ylim([-0.1, 1.1])
m = -1
#ax.plot(X, m * X + 1.2, label="decision boundary")
plt.plot()
24
PERCEPTRON FOR THE AND FUNCTION](https://image.slidesharecdn.com/navebayes-230201064135-58c1858c/75/Naive-Bayes-pptx-24-2048.jpg)
![fig, ax = plt.subplots()
xmin, xmax = -0.2, 1.4
X = np.arange(xmin, xmax, 0.1)
ax.set_xlim([xmin, xmax])
ax.set_ylim([-0.1, 1.1])
m = -1
for m in np.arange(0, 6, 0.1):
ax.plot(X, m * X )
ax.scatter(0, 0, color="r")
ax.scatter(0, 1, color="r")
ax.scatter(1, 0, color="r")
ax.scatter(1, 1, color="g")
plt.plot()
25
PERCEPTRON FOR THE AND FUNCTION](https://image.slidesharecdn.com/navebayes-230201064135-58c1858c/75/Naive-Bayes-pptx-25-2048.jpg)
![fig, ax = plt.subplots()
xmin, xmax = -0.2, 1.4
X = np.arange(xmin, xmax, 0.1)
ax.scatter(0, 0, color="r")
ax.scatter(0, 1, color="r")
ax.scatter(1, 0, color="r")
ax.scatter(1, 1, color="g")
ax.set_xlim([xmin, xmax])
ax.set_ylim([-0.1, 1.1])
m, c = -1, 1.2
ax.plot(X, m * X + c )
plt.plot()
26
PERCEPTRON FOR THE AND FUNCTION](https://image.slidesharecdn.com/navebayes-230201064135-58c1858c/75/Naive-Bayes-pptx-26-2048.jpg)




![CREATE A NEURAL NETWORK CLASS IN PYTHON TO
TRAIN THE NEURON TO GIVE AN ACCURATE
PREDICTION
training_inputs = np.array([[0,0,1],
[1,1,1],
[1,0,1],
[0,1,1]])
training_outputs = np.array([[0,1,1,0]]).T
#training taking place
neural_network.train(training_inputs, training_outputs, 15000)
print("Ending Weights After Training: ")
print(neural_network.synaptic_weights)
32](https://image.slidesharecdn.com/navebayes-230201064135-58c1858c/75/Naive-Bayes-pptx-32-2048.jpg)


