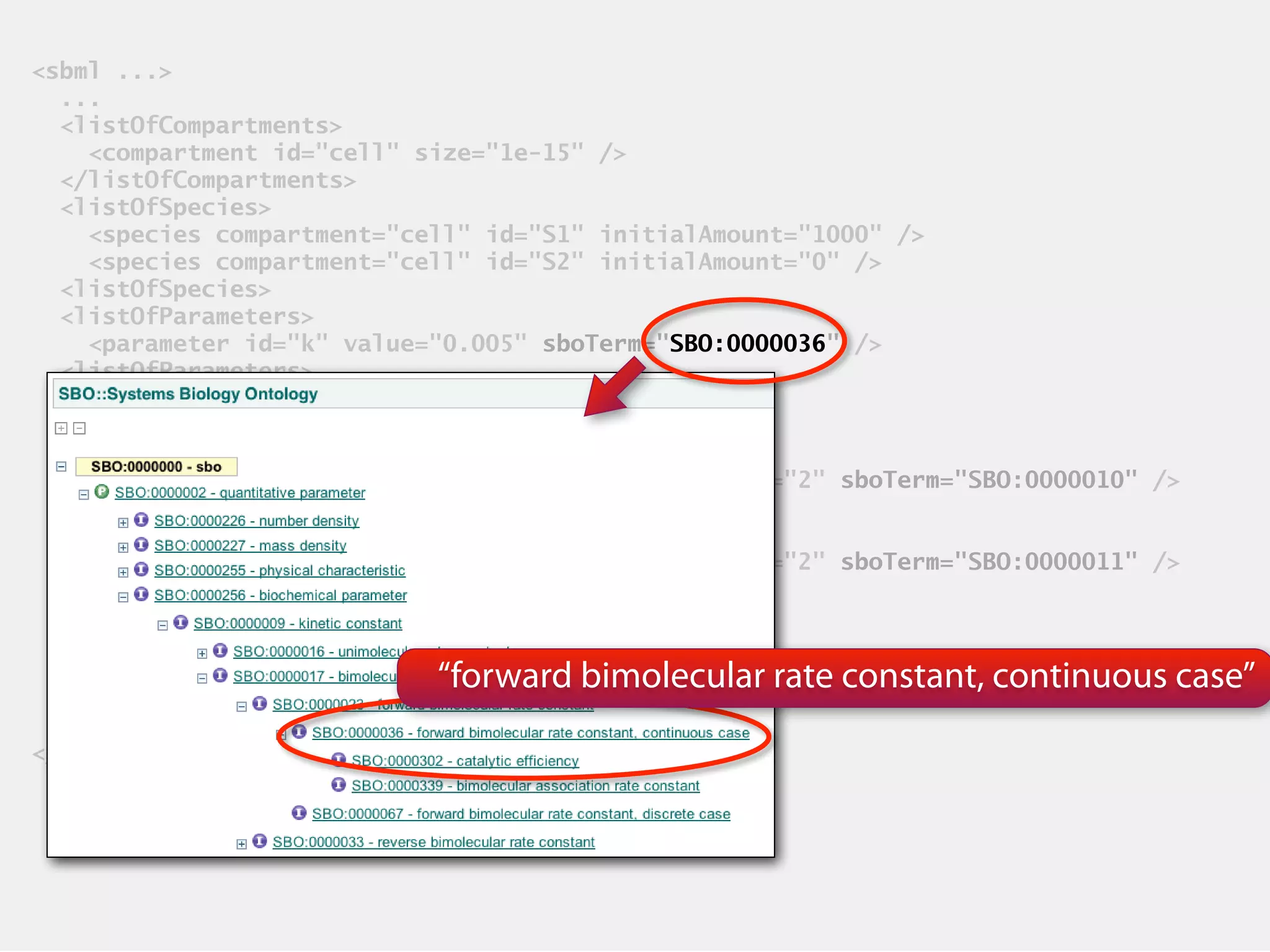
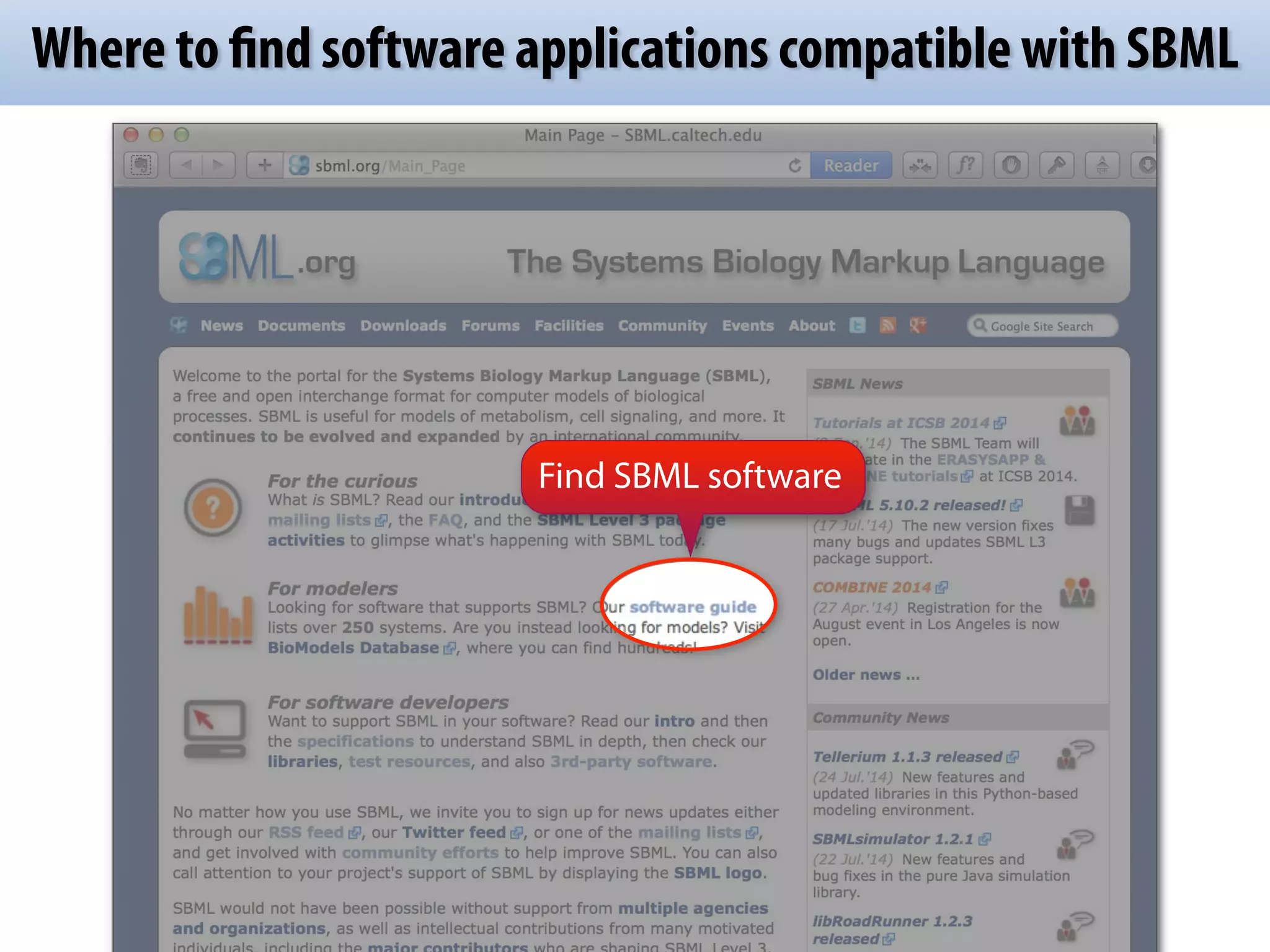
SBML (Systems Biology Markup Language) is an XML-based file format designed for representing models of biological processes, facilitating model exchange across various software tools. It provides a neutral framework for modeling that allows for increased reuse and communication of biological semantics, enhancing research collaboration. The document outlines SBML's core concepts, software compatibility, and usage practices, emphasizing its importance in computational biology and systems biology research.





























![Interpreting reactions in SBML
Why are SBML rate expression not identical to rate laws?
• Consider reaction S1 S2
between 2 compartments, #1 and #2
• Assume told . What does this mean?
rartaetel alaww == k · [S1]
!
• But what if volume ?
- Look at # of molecules in each compartment:
! - How molecules leave enter each compartment?
V1
S1
S2
molecules leave the first compartment
molecules enter the second compartment
V2
d[S2]
dt
=
d[S1]
dt
= k · [S1]
V2 = 3V1
nS1 = [S1] · V1 nS2 = [S2] · V2
k · [S1] · V1
3 · k · [S1] · V1](https://image.slidesharecdn.com/mhucka-sbml-tutorial-part1-icsb-2014-140919174259-phpapp01/75/SBML-the-Systems-Biology-Markup-Language-33-2048.jpg)
![Interpreting reactions in SBML
Why are SBML rate expression not identical to rate laws?
• Consider reaction S1 S2
between 2 compartments, #1 and #2
• Assume told . What does this mean?
rartaetel alaww == k · [S1]
!
• But what if volume ?
- Look at # of molecules in each compartment:
! - How molecules leave enter each compartment?
V1
S1
S2
molecules leave the first compartment
molecules enter the second compartment
V2
d[S2]
dt
=
d[S1]
dt
= k · [S1]
V2 = 3V1
nS1 = [S1] · V1 nS2 = [S2] · V2
k · [S1] · V1
3 · k · [S1] · V1
Oops! Something’s
wrong …](https://image.slidesharecdn.com/mhucka-sbml-tutorial-part1-icsb-2014-140919174259-phpapp01/75/SBML-the-Systems-Biology-Markup-Language-34-2048.jpg)
![“Kinetic law” in SBML
Rate expressions are (usually) substance/time, not substance/size/time
Conversion for basic cases is simple:
• Multiply by volume of compartment where reactants are located:
!
rate = k · [S1] · V1
! • Express rates of changes of reactants products in terms of substances:
!
!
!
dnS1
dt
= k1 · [S1] · V1
!
• Can easily recover concentrations:
[S1] = nS1
V1
[S2] = nS2
V2
moles, or
items, or
mass, or
“dimensionless”
dnS2
dt
= k1 · [S1] · V1](https://image.slidesharecdn.com/mhucka-sbml-tutorial-part1-icsb-2014-140919174259-phpapp01/75/SBML-the-Systems-Biology-Markup-Language-35-2048.jpg)