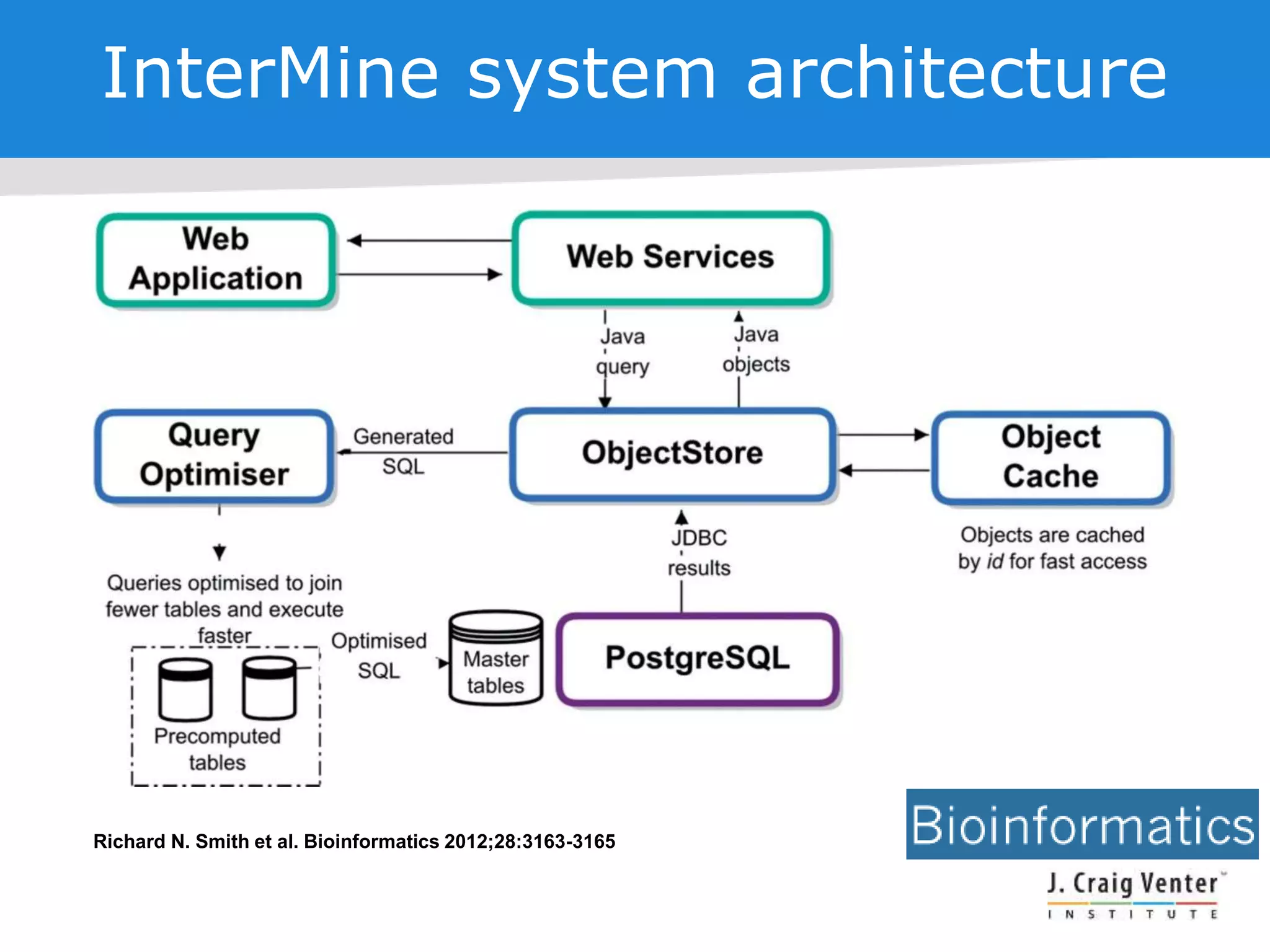
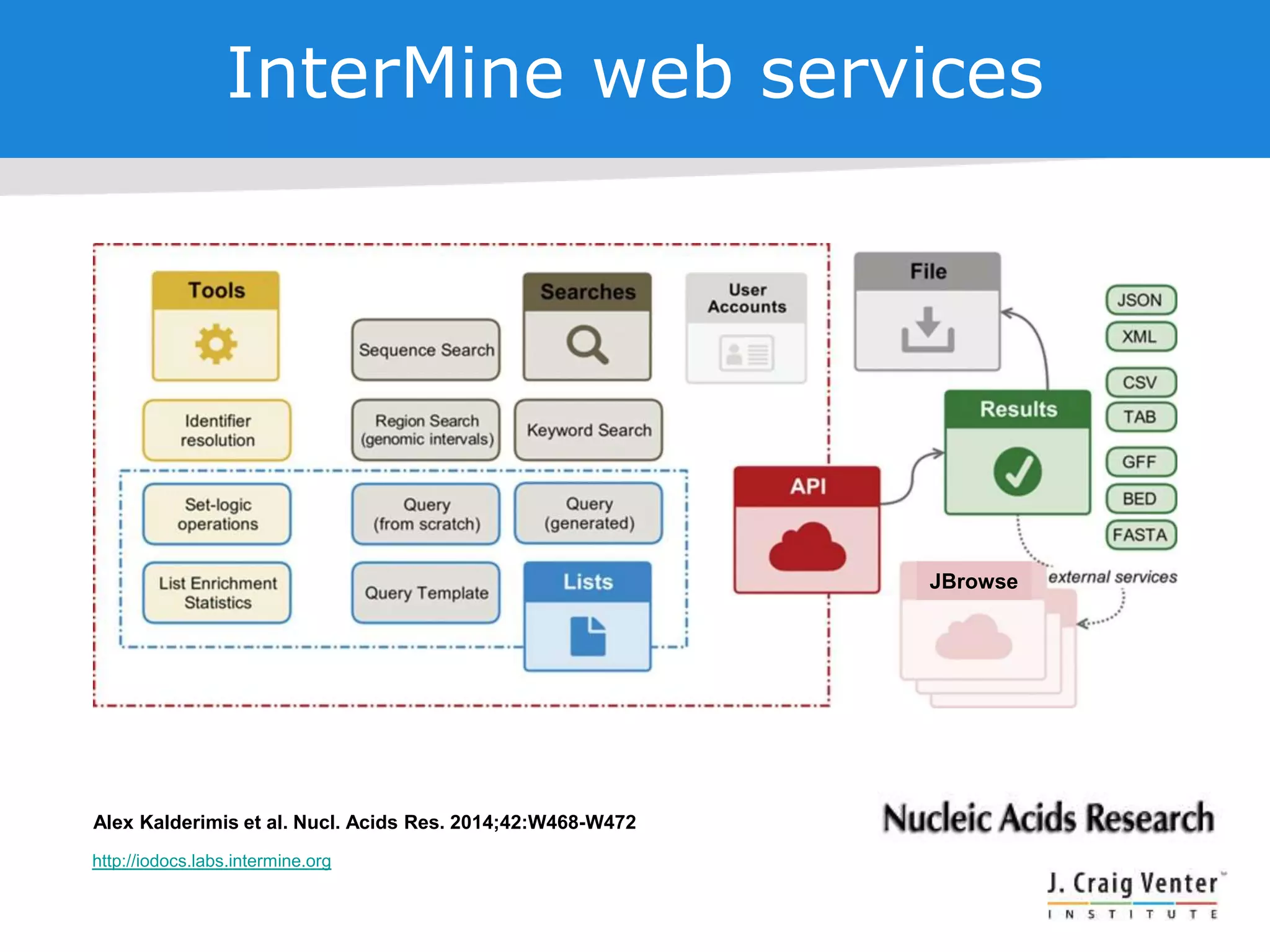
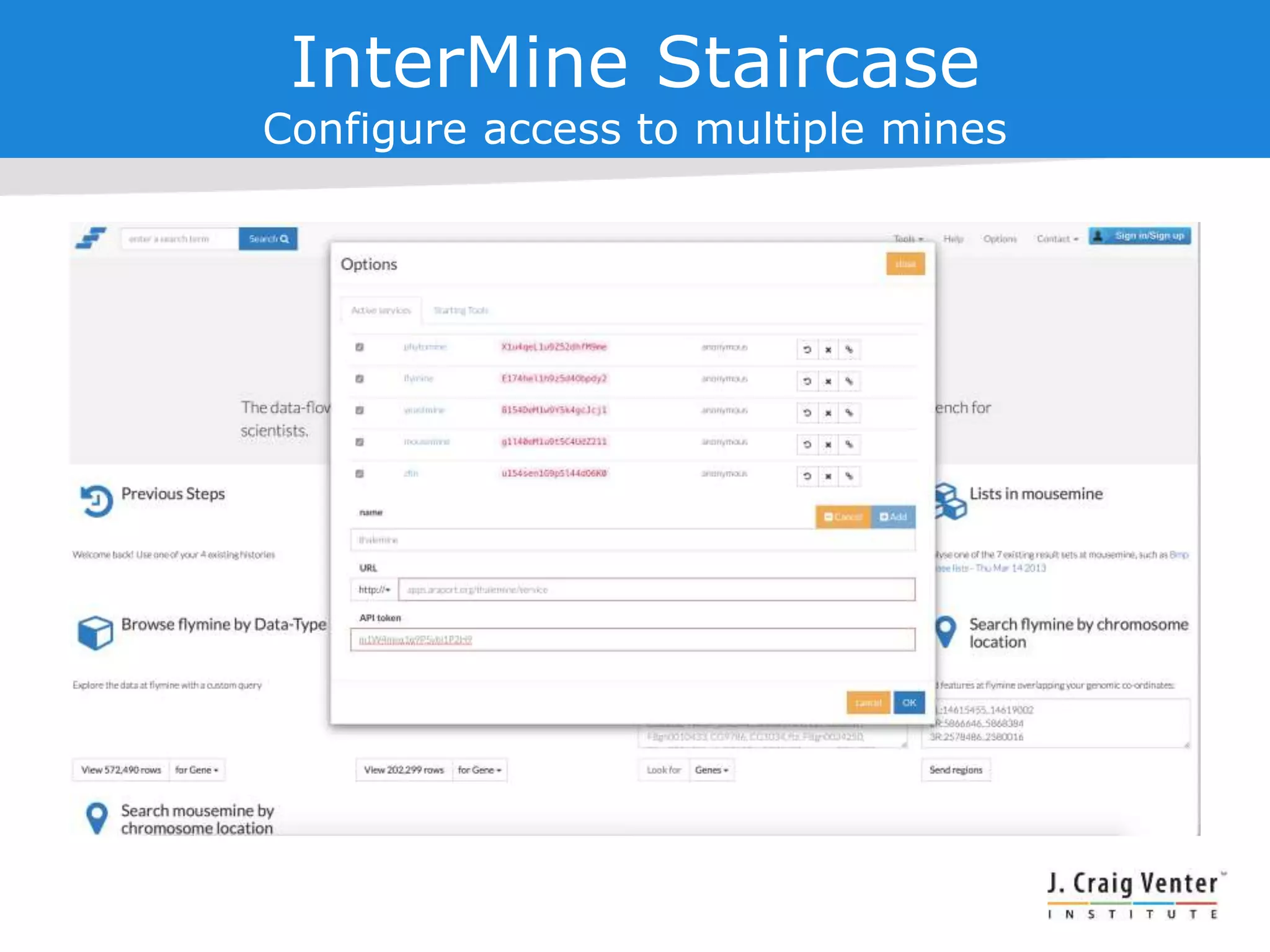
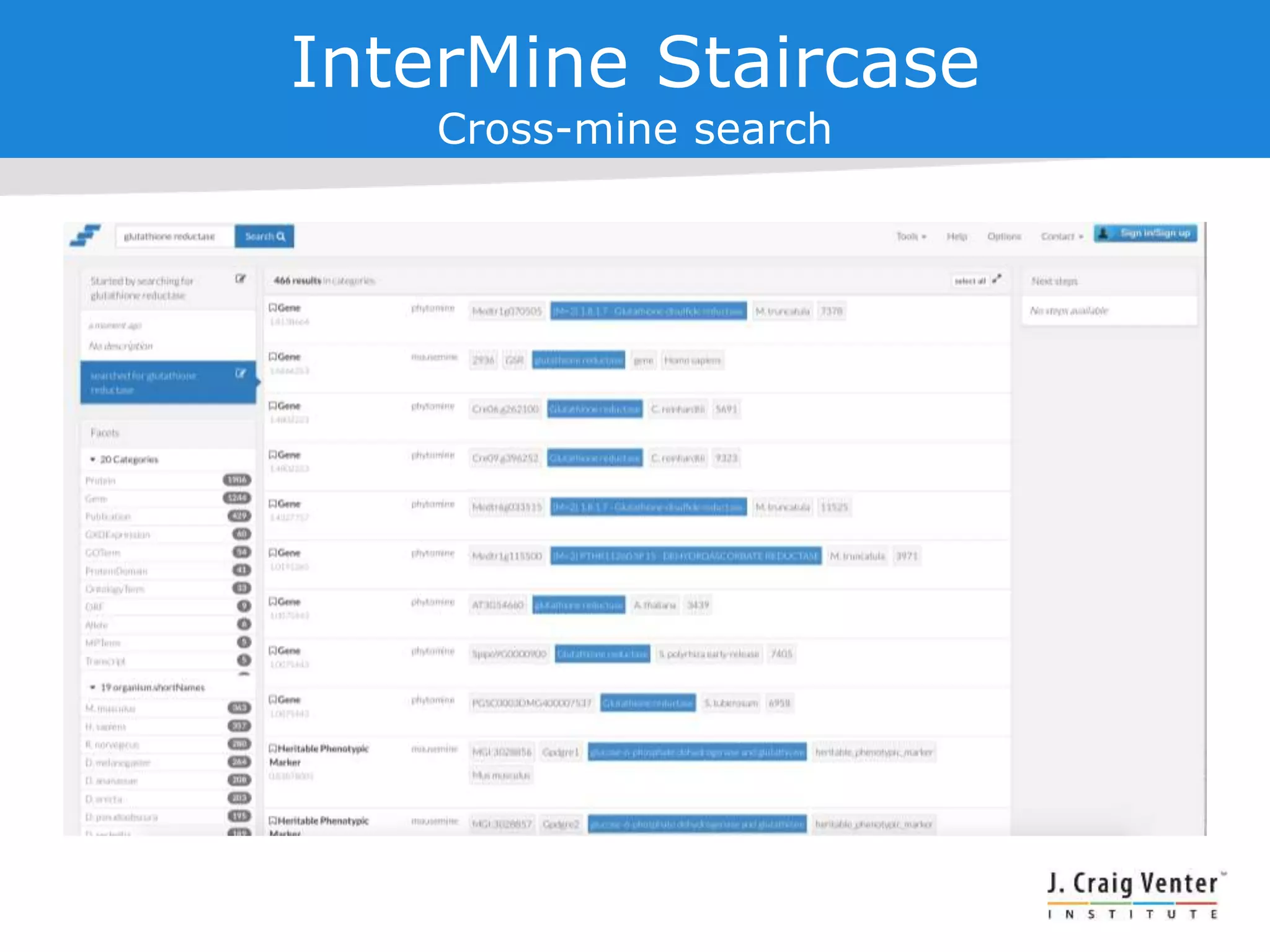
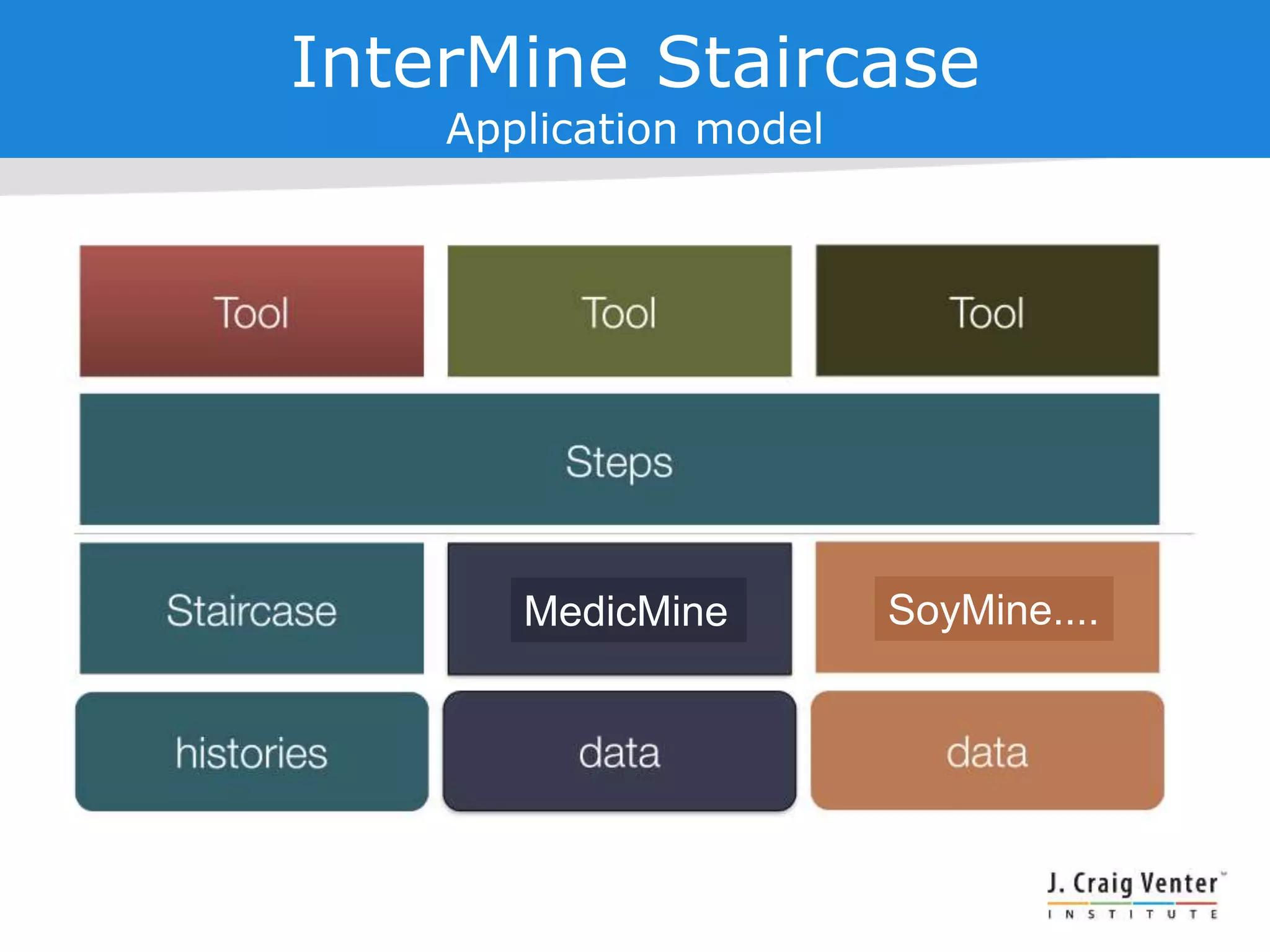
The document discusses InterMine, an open-source data warehouse software designed for the integration and management of diverse biological data. It outlines its architecture, features like web services, interoperability, and advantages of a common data model that fosters data mining and analysis across multiple biological datasets. Additionally, it highlights specific applications like ThaleMine and MedicMine, and provides recommendations and challenges for future development within the InterMine framework.