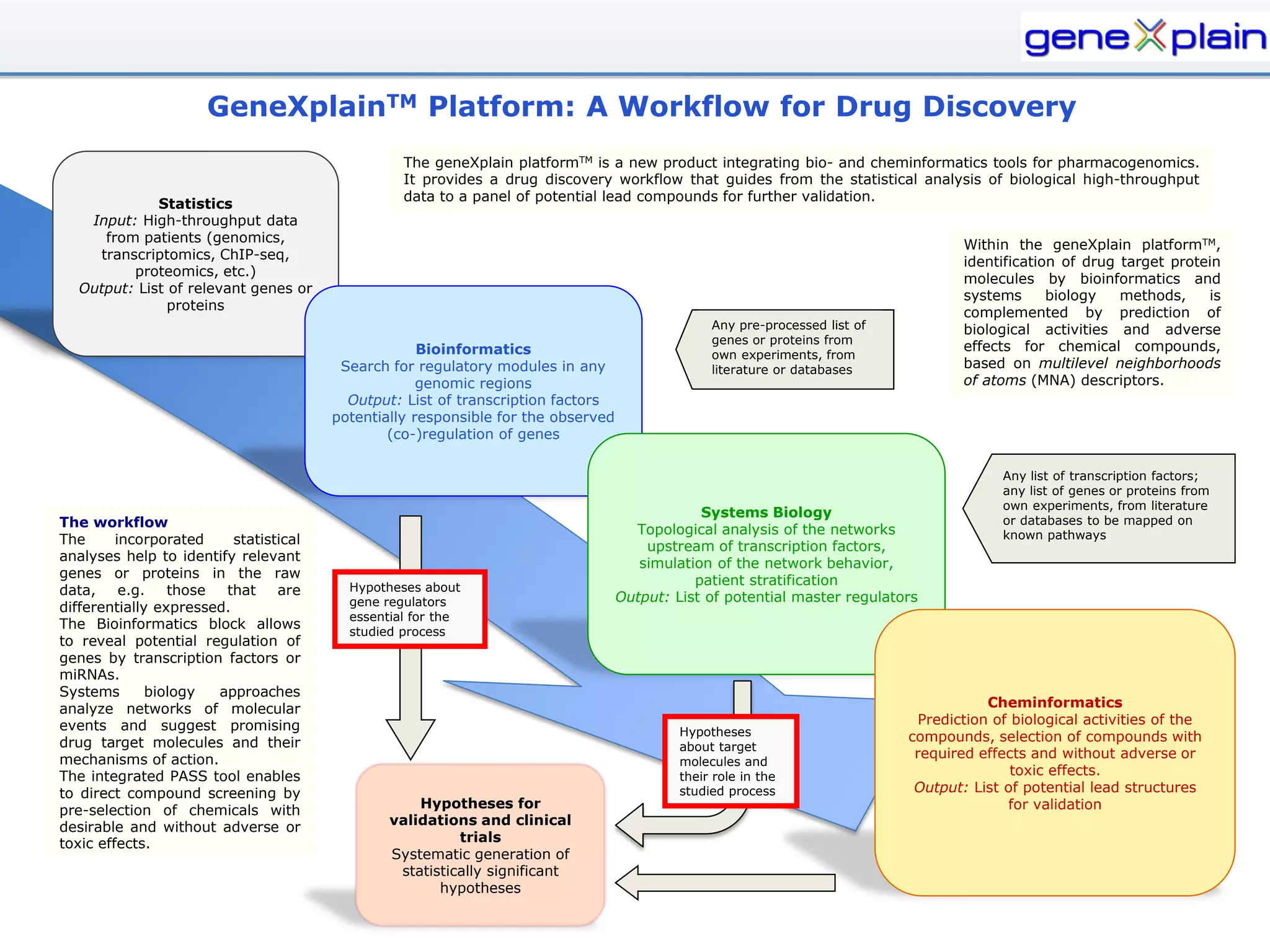
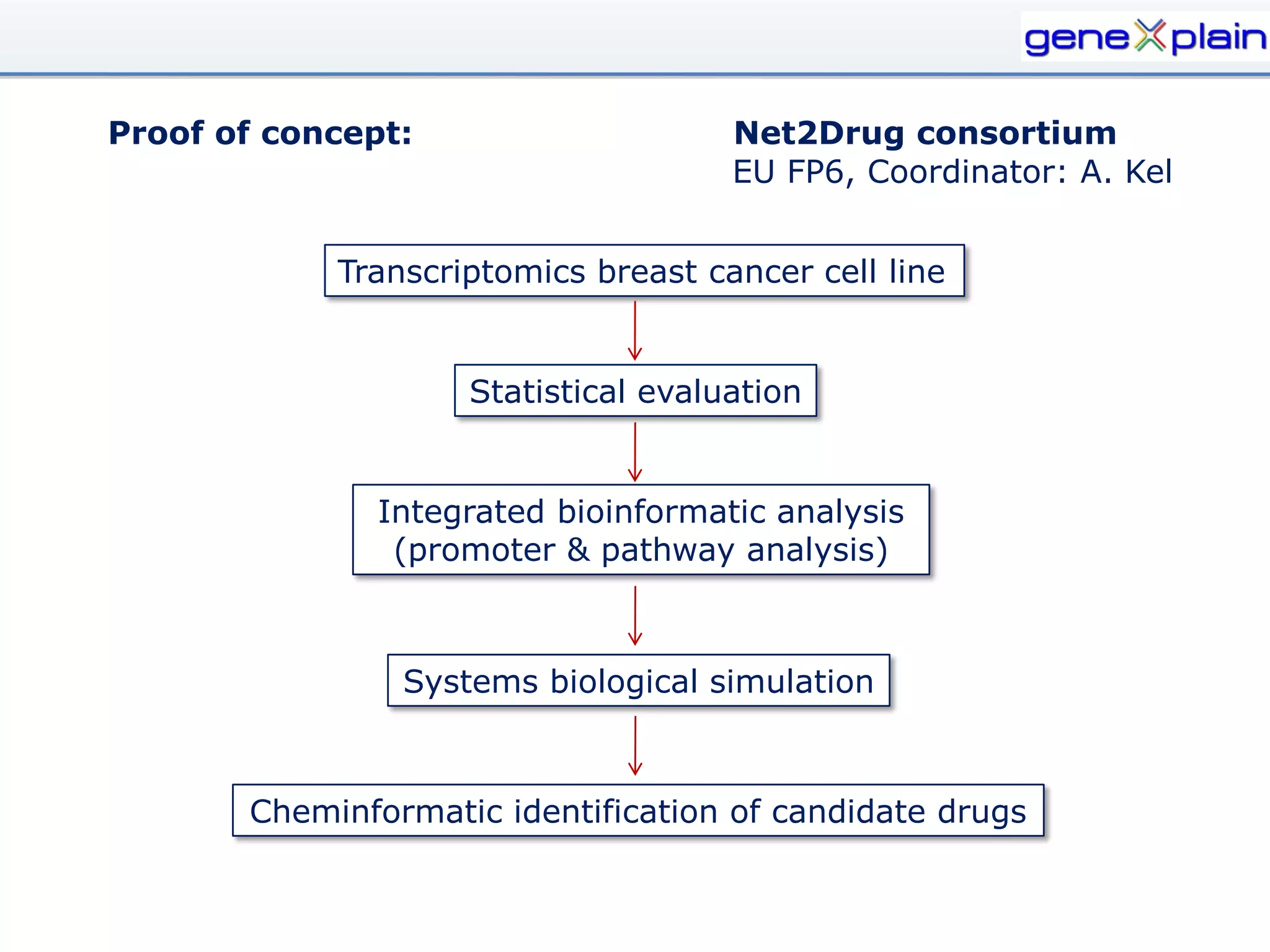
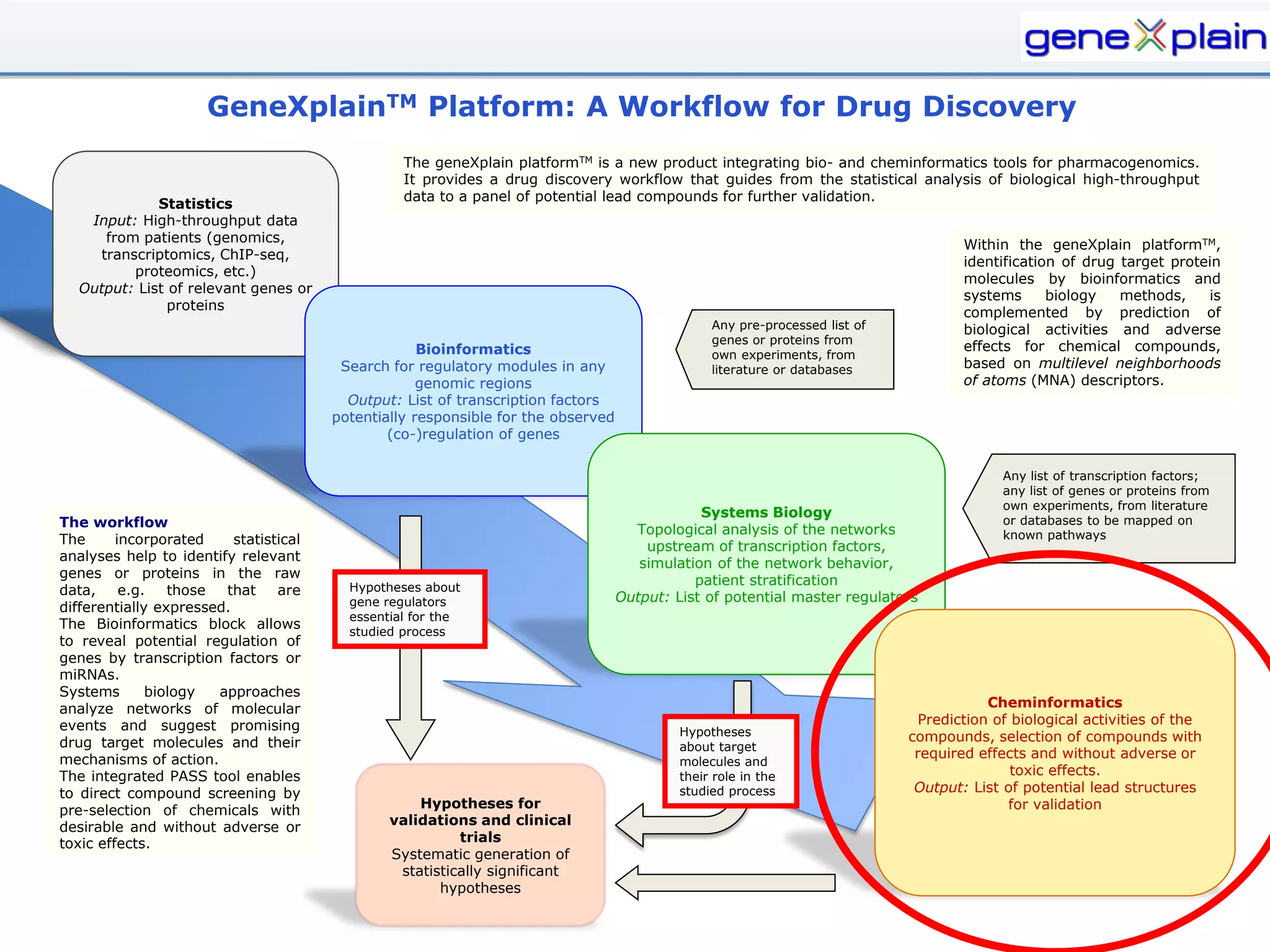
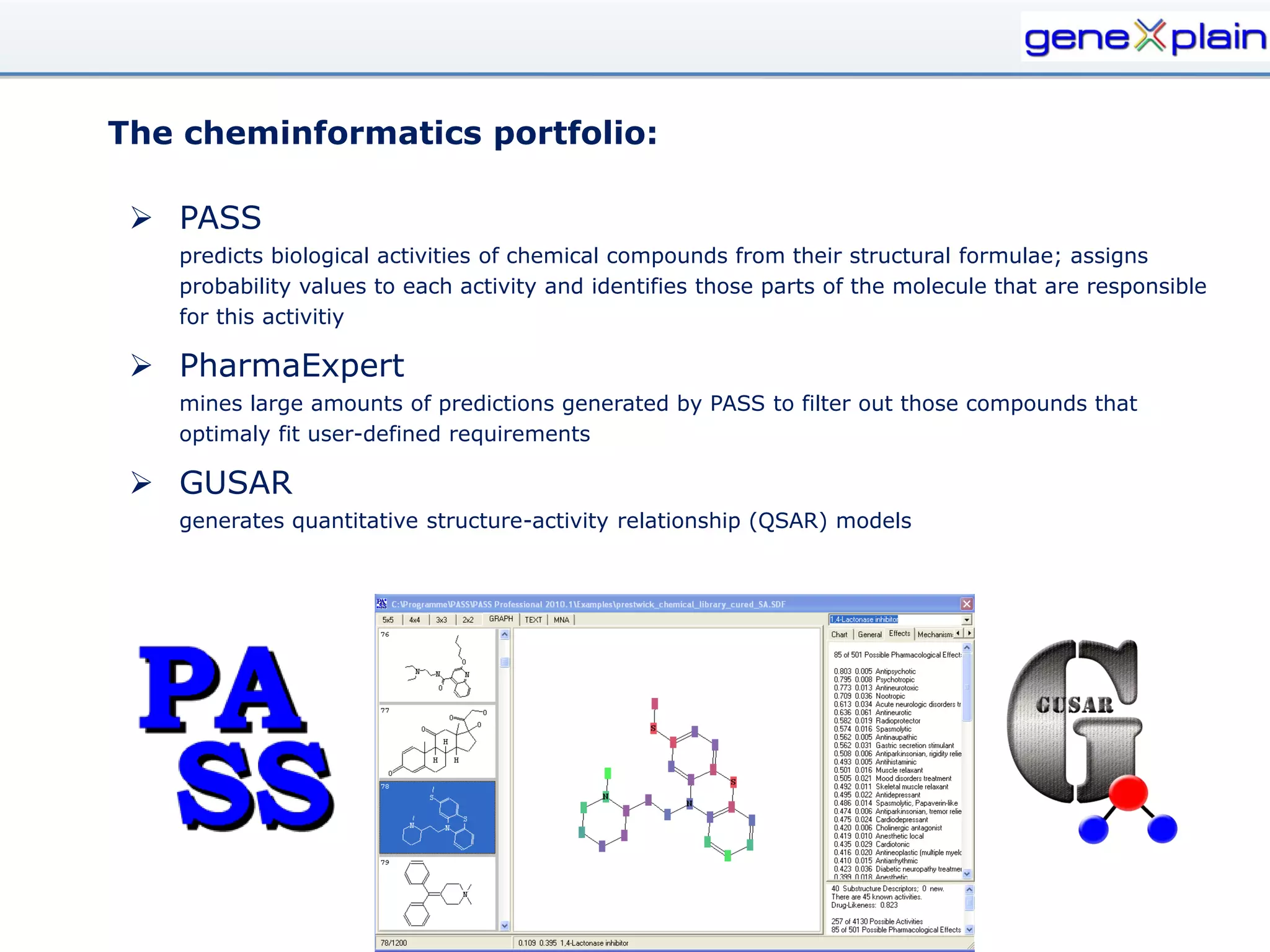
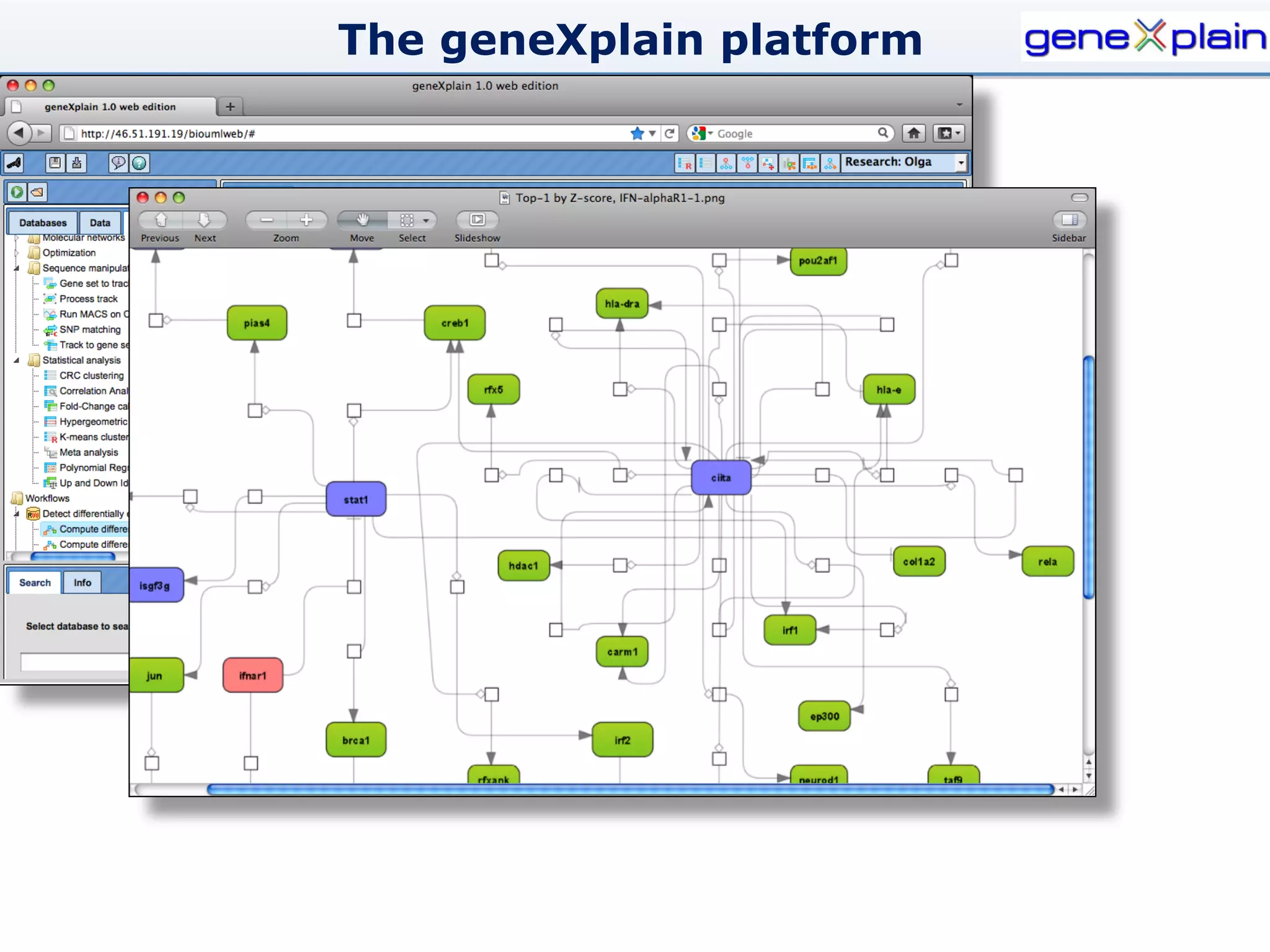
The document outlines the Genexplain platform, a comprehensive drug development and bioinformatics tool created through a Russian-German partnership, founded in 2010. It integrates systems biology and cheminformatics to facilitate personalized medicine by providing a workflow that guides drug discovery from high-throughput data analysis to potential lead compound identification. The platform also offers a mix of proprietary and free tools, enhancing accessibility and user flexibility while addressing common drawbacks of public domain services.