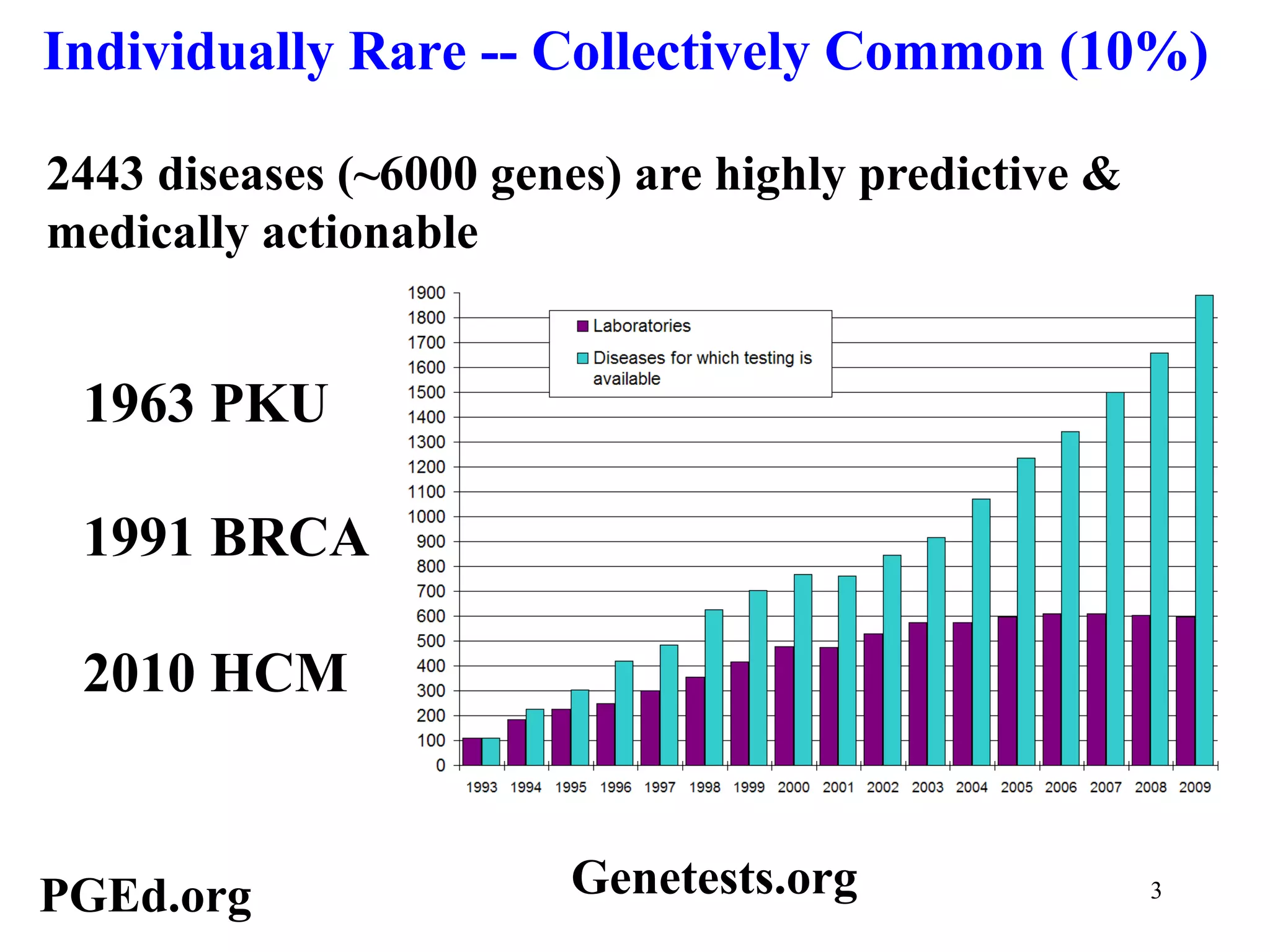
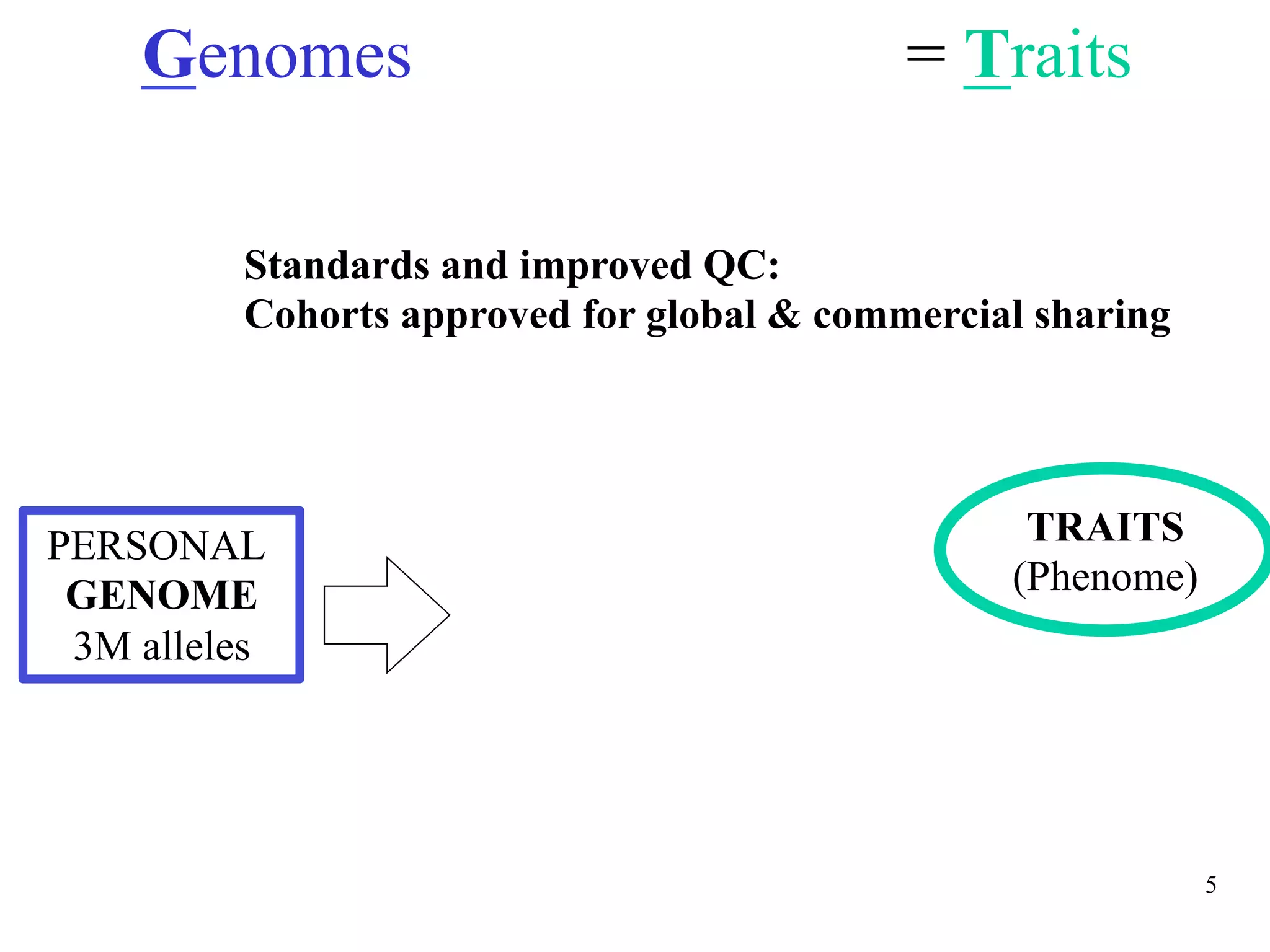
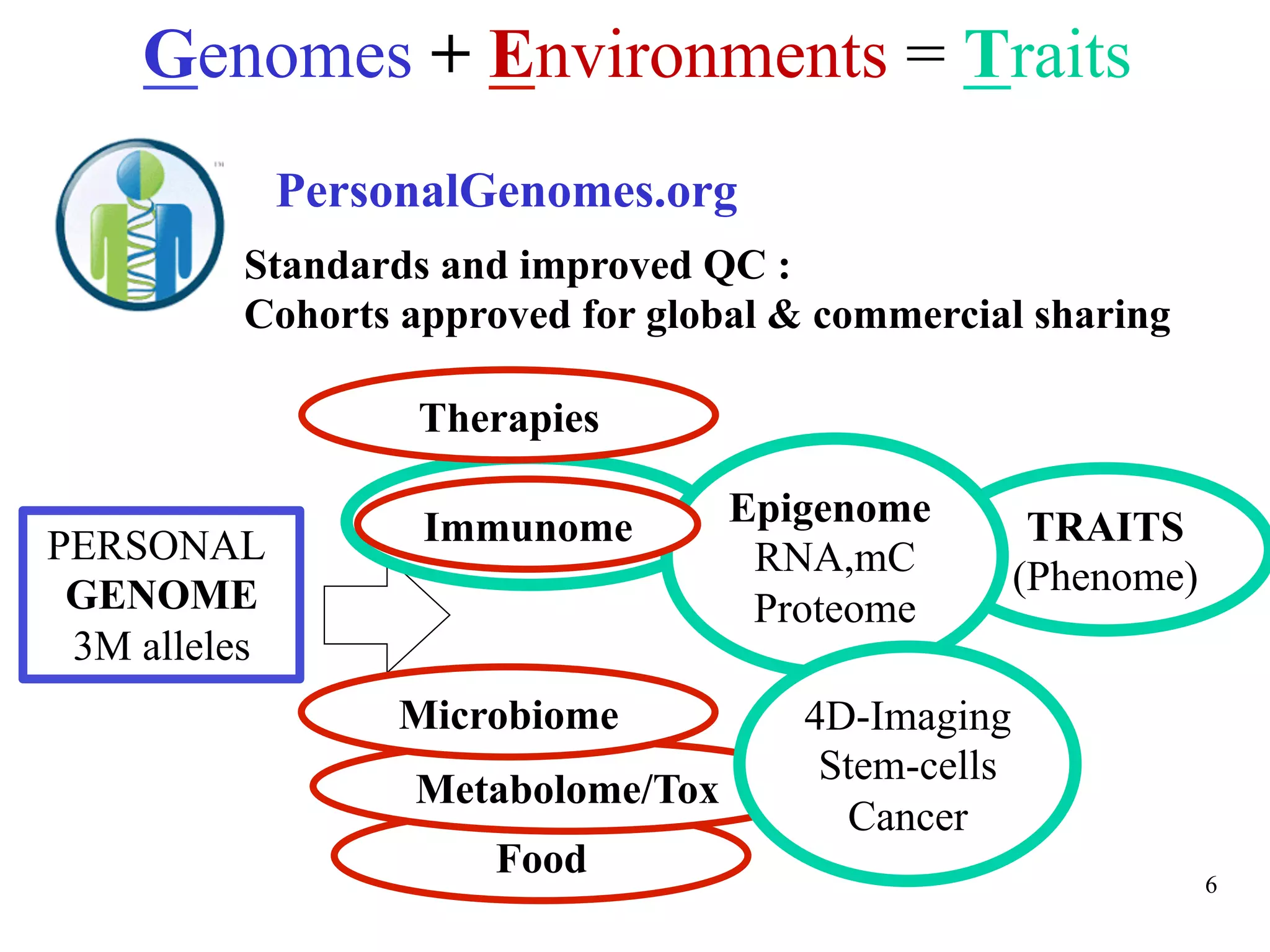
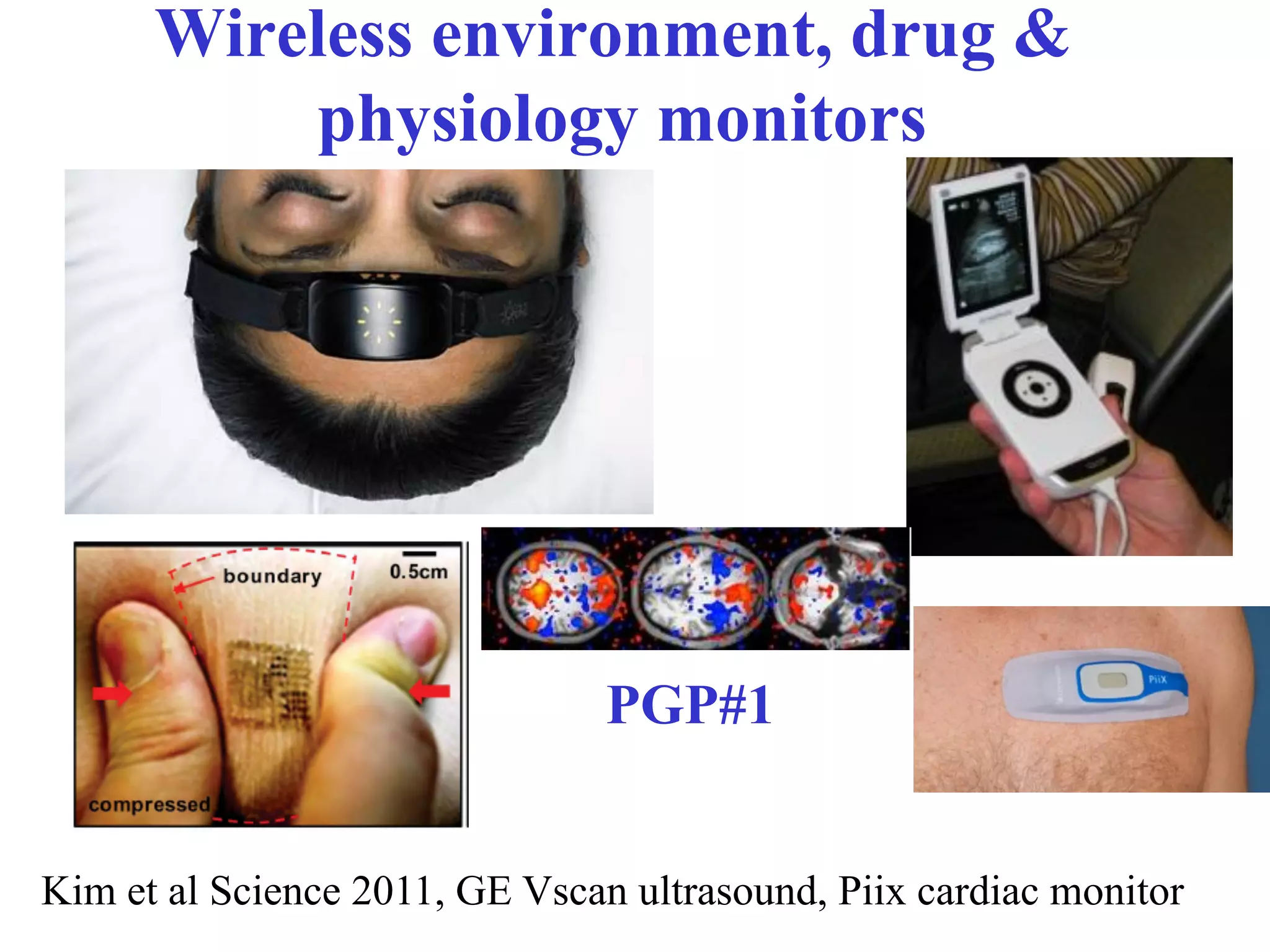
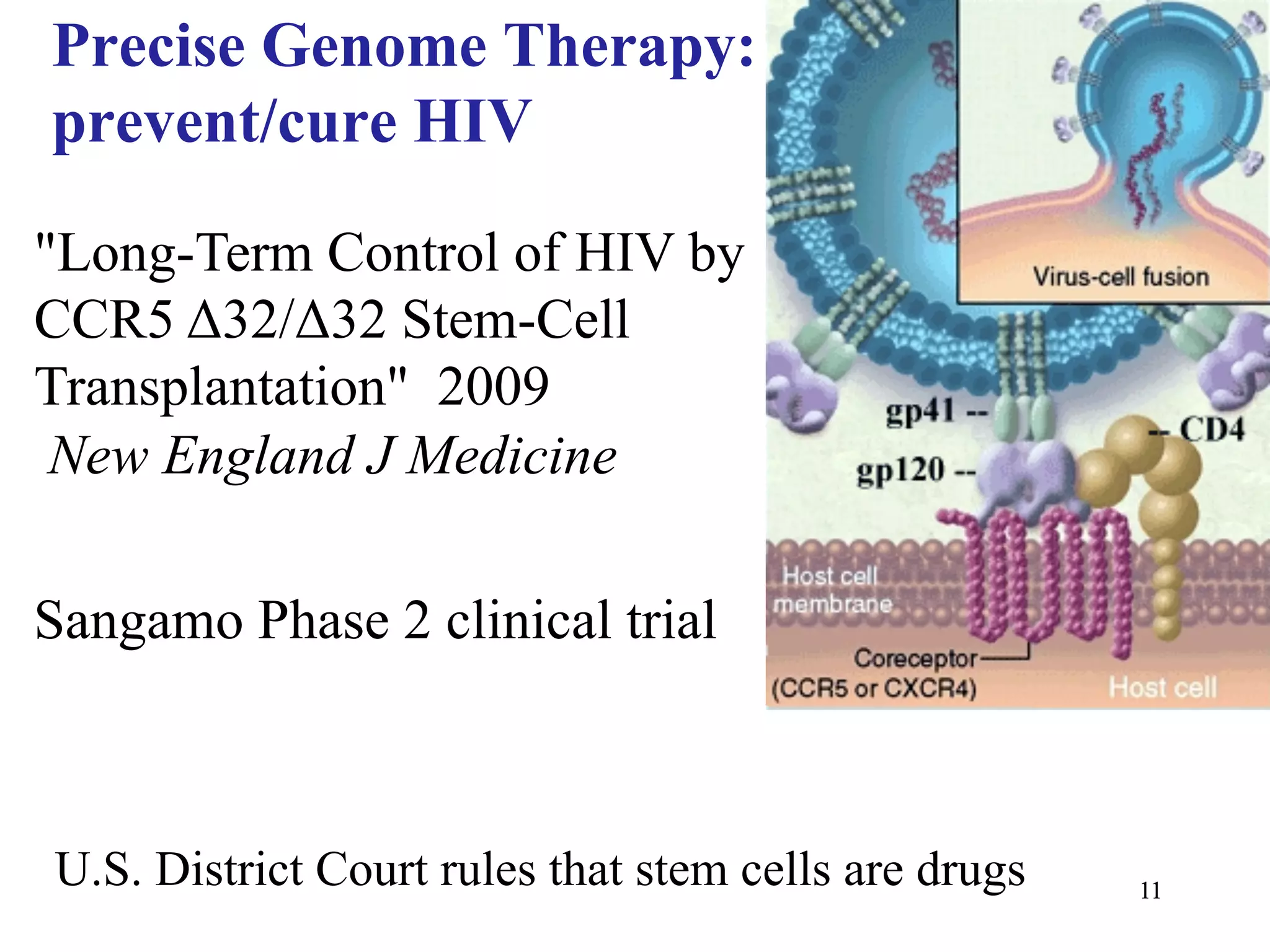
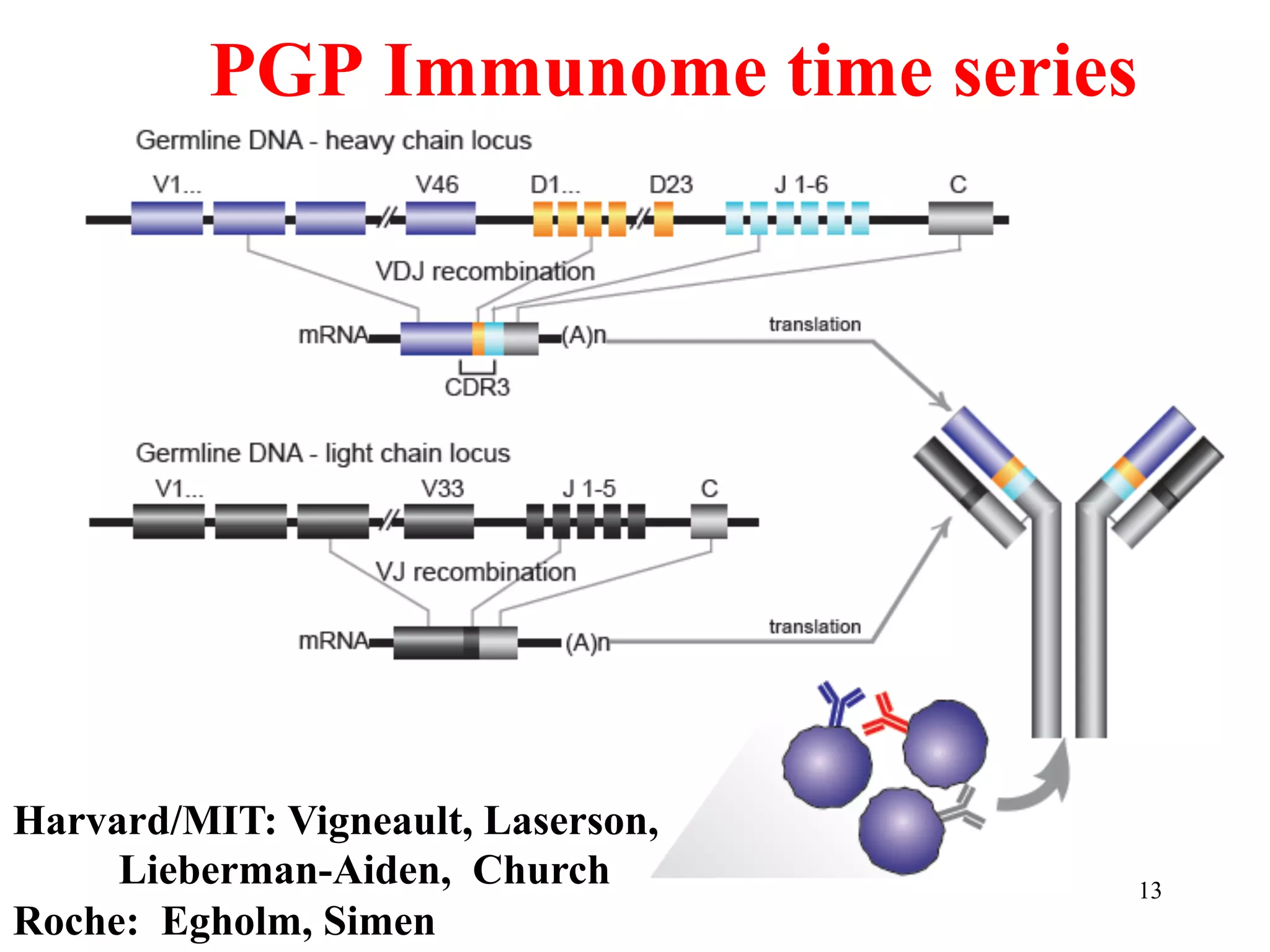
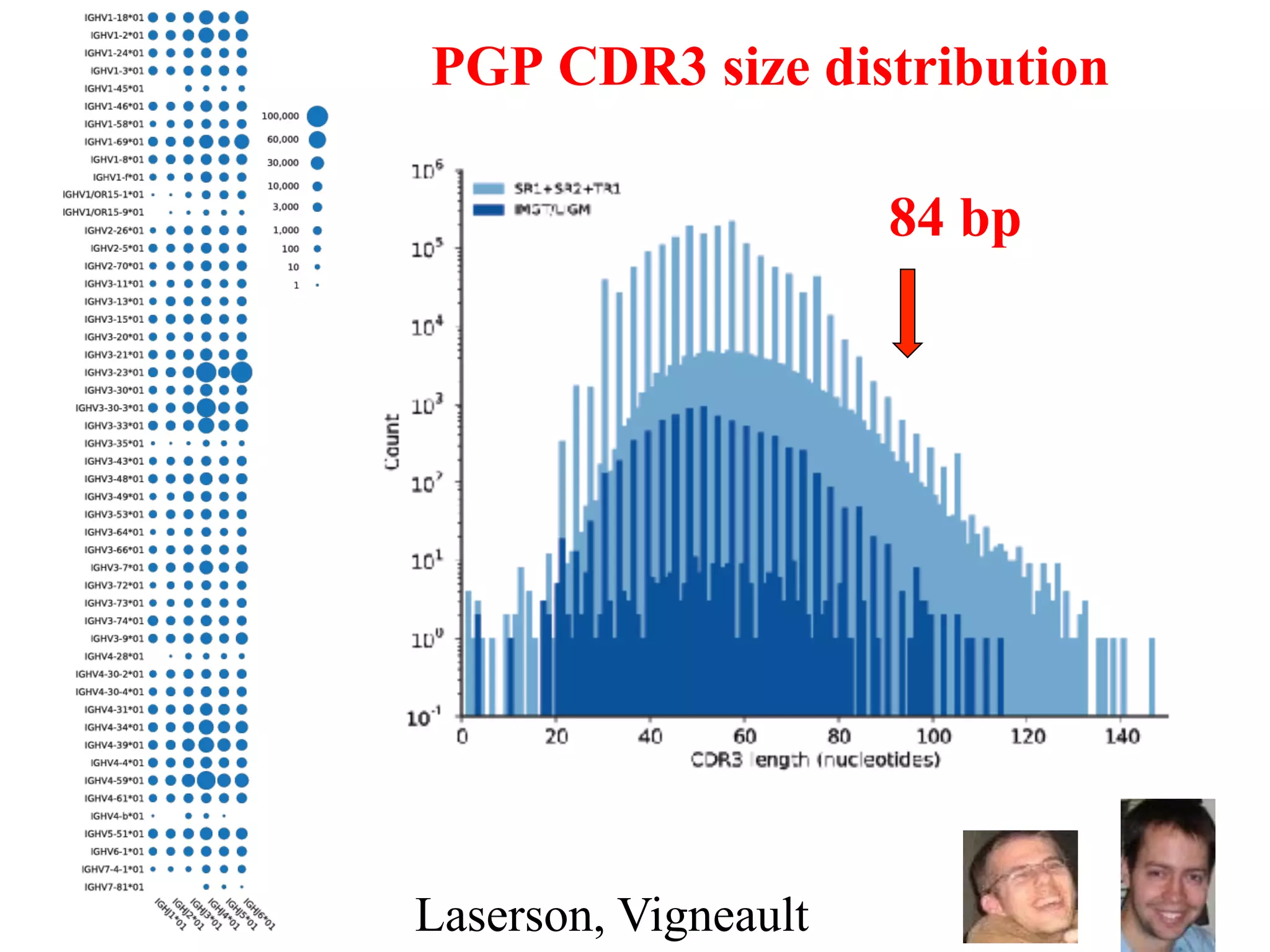
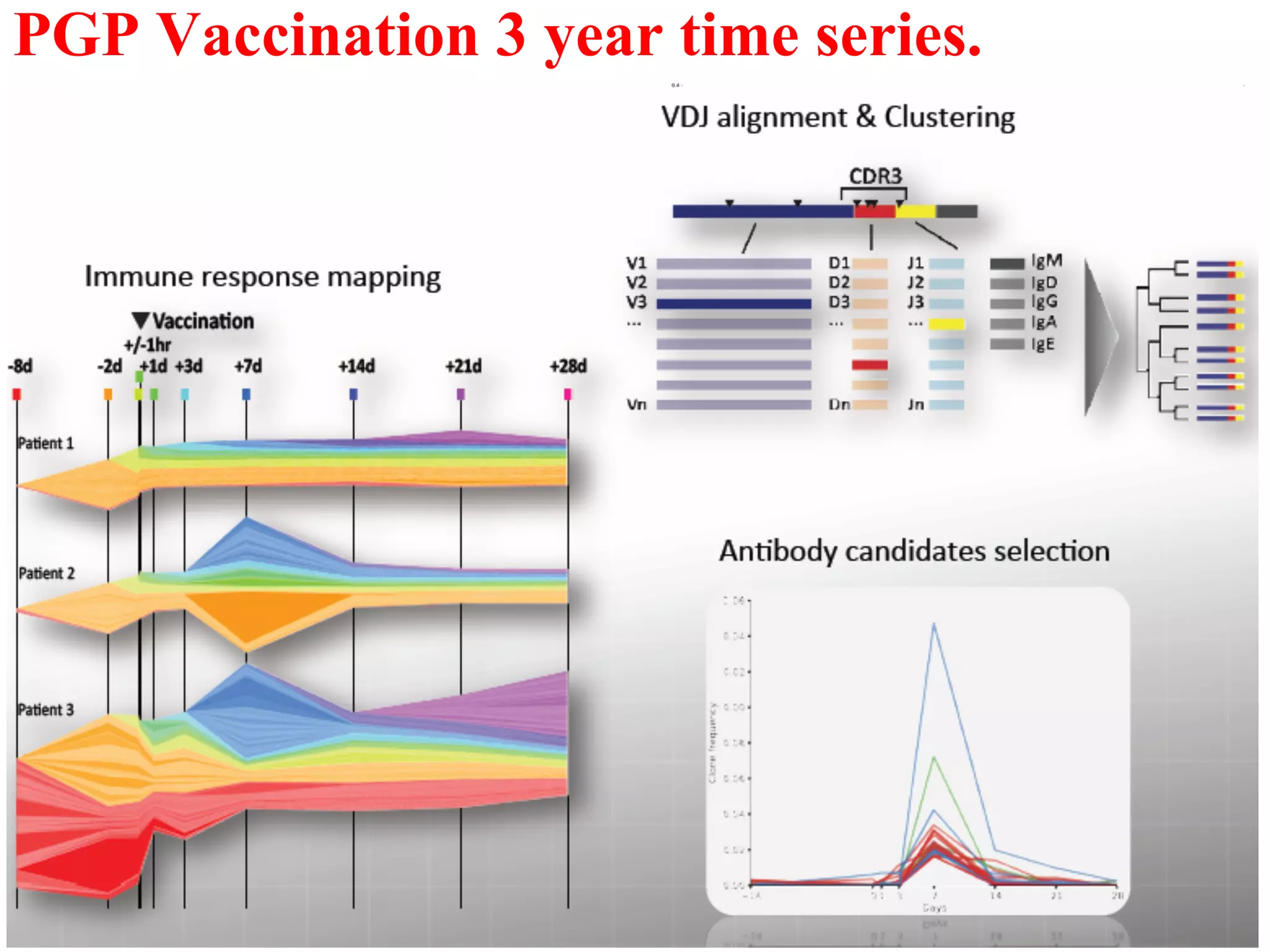
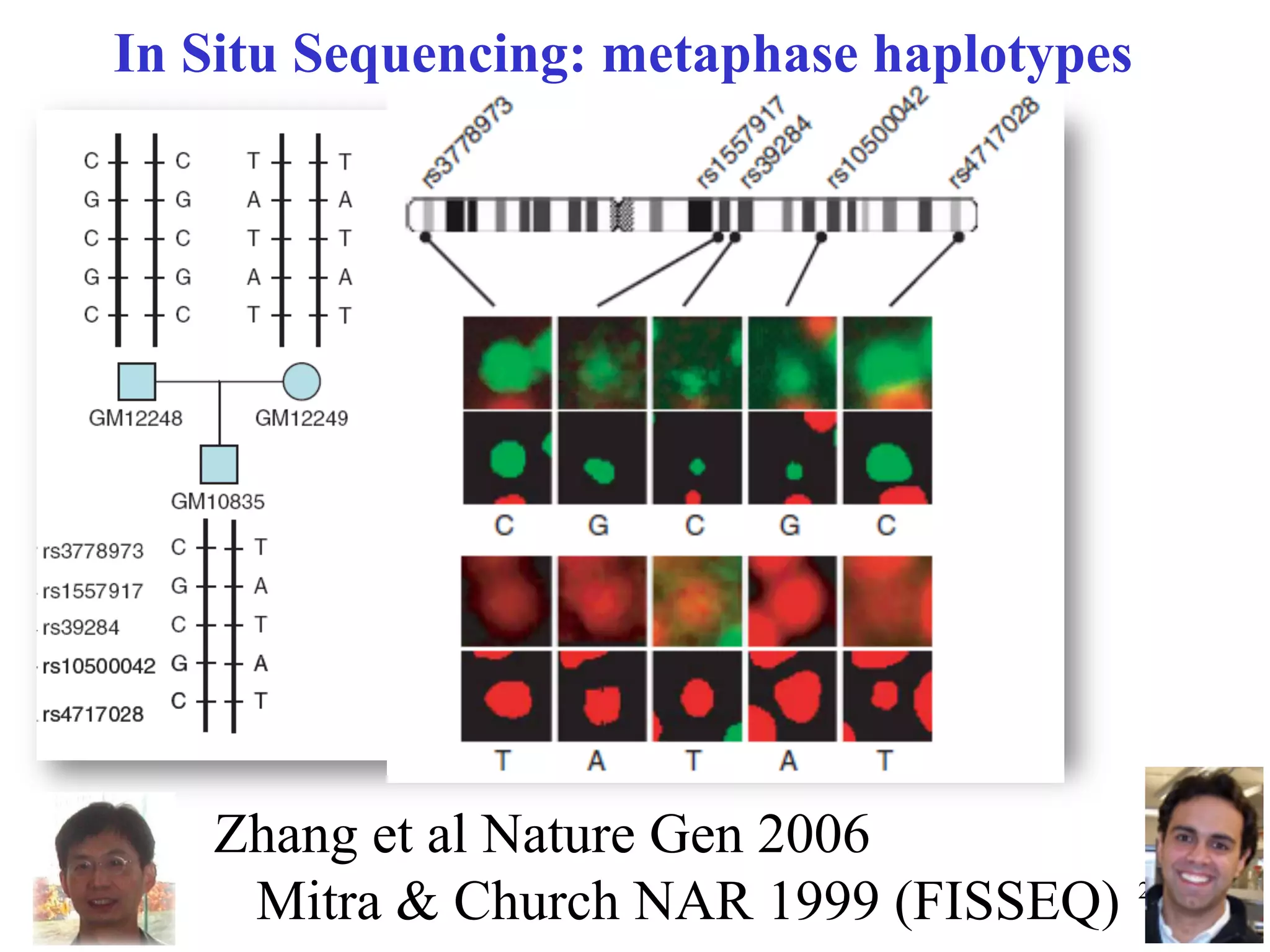
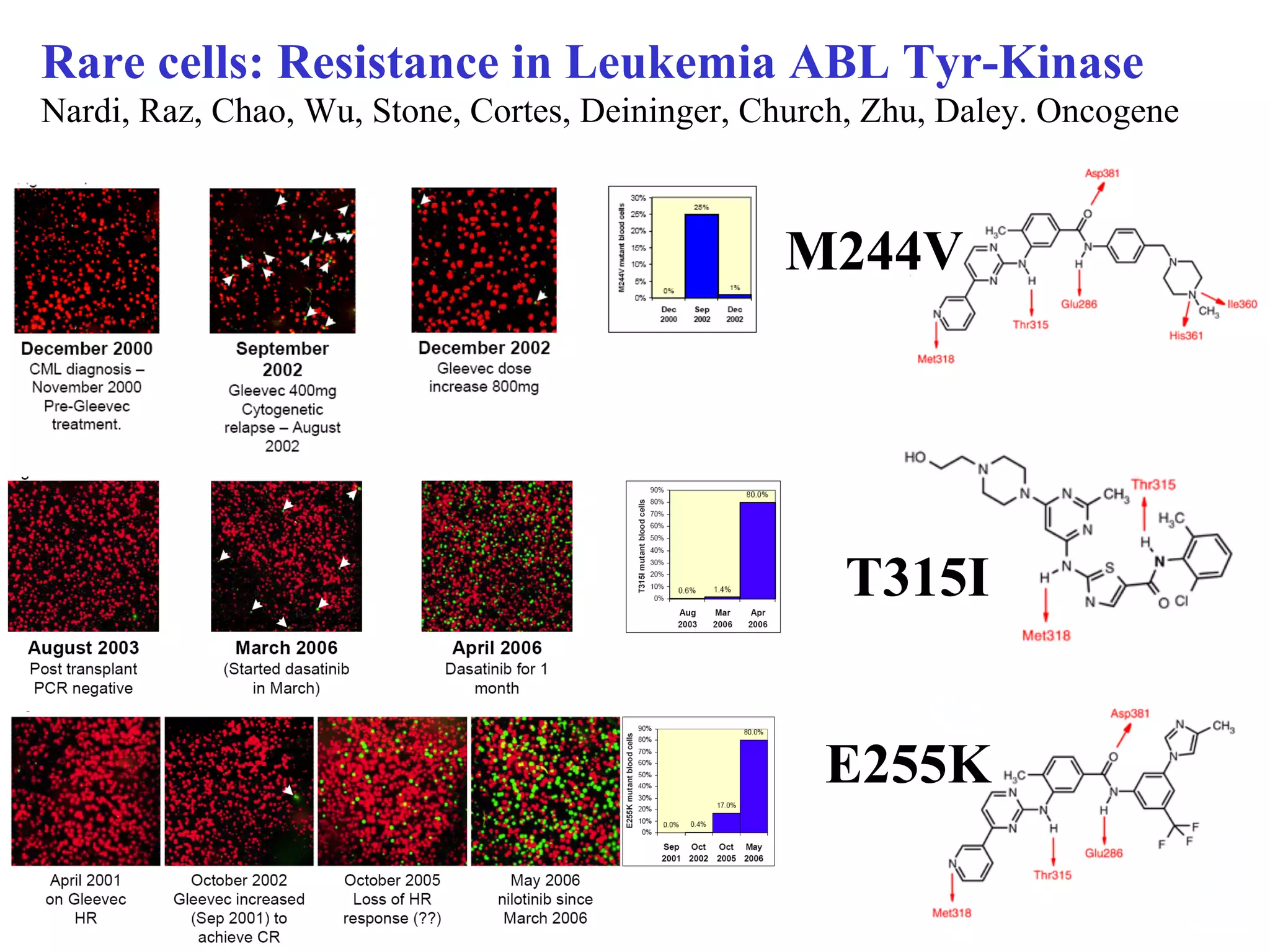
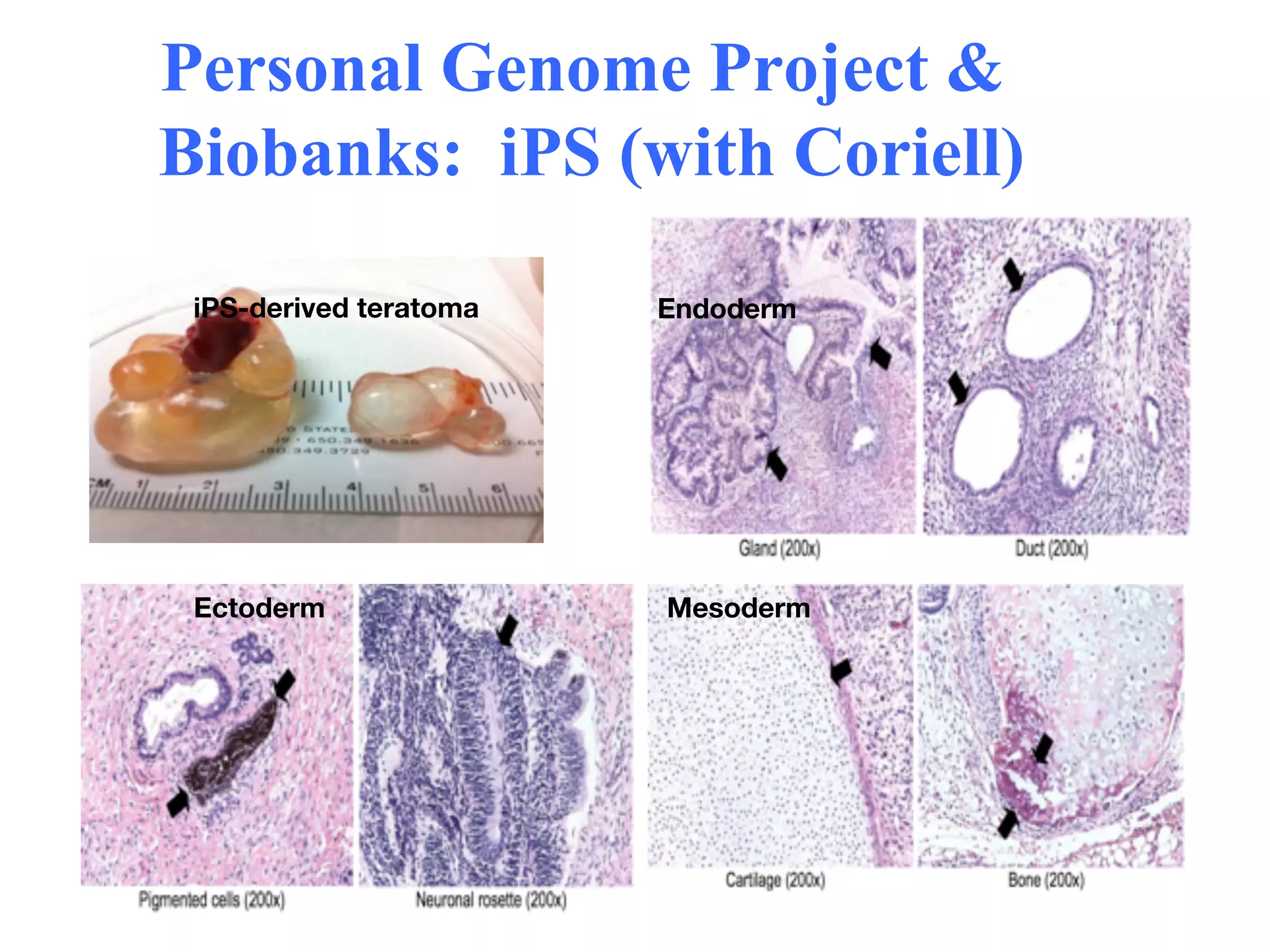
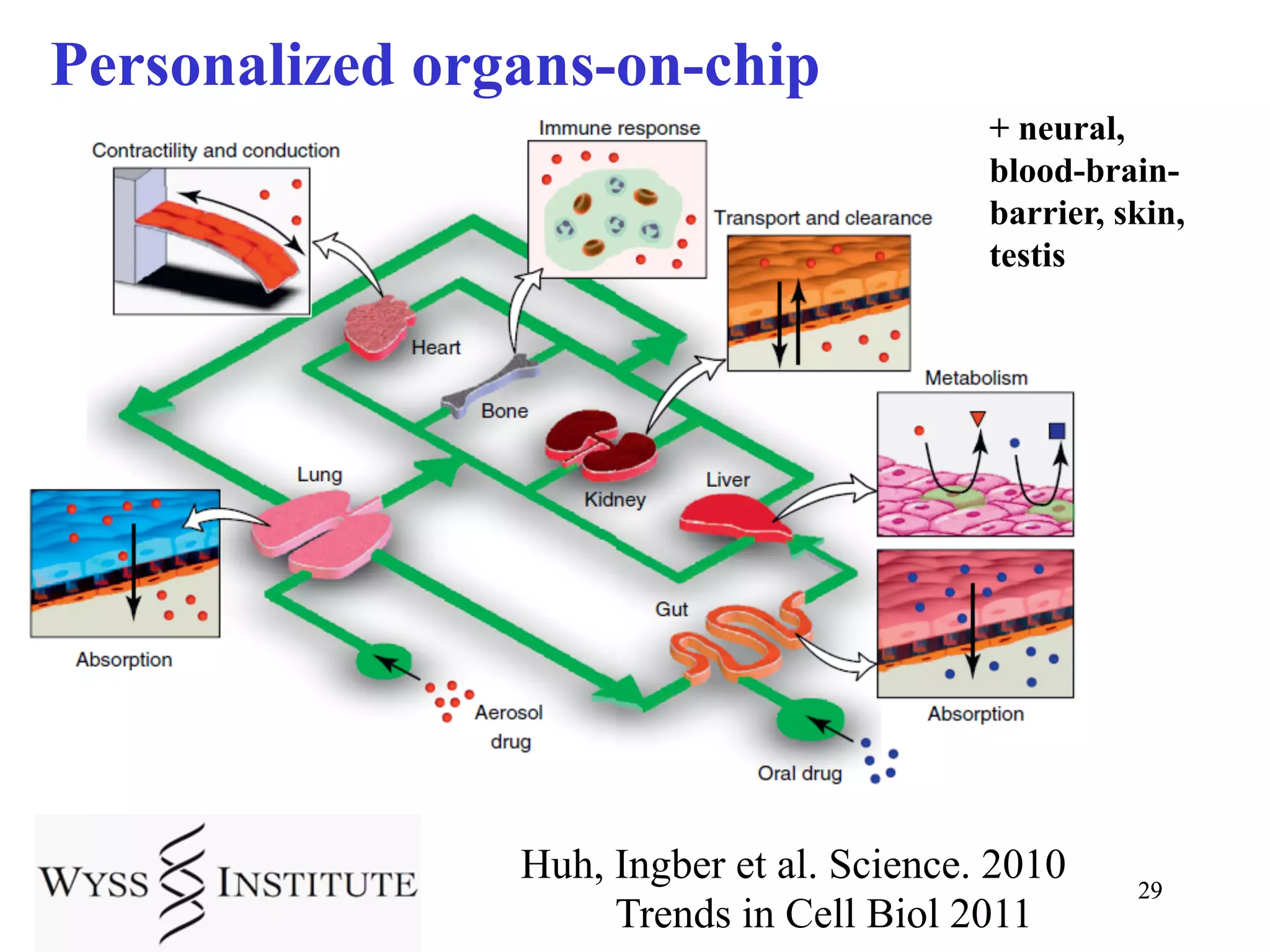
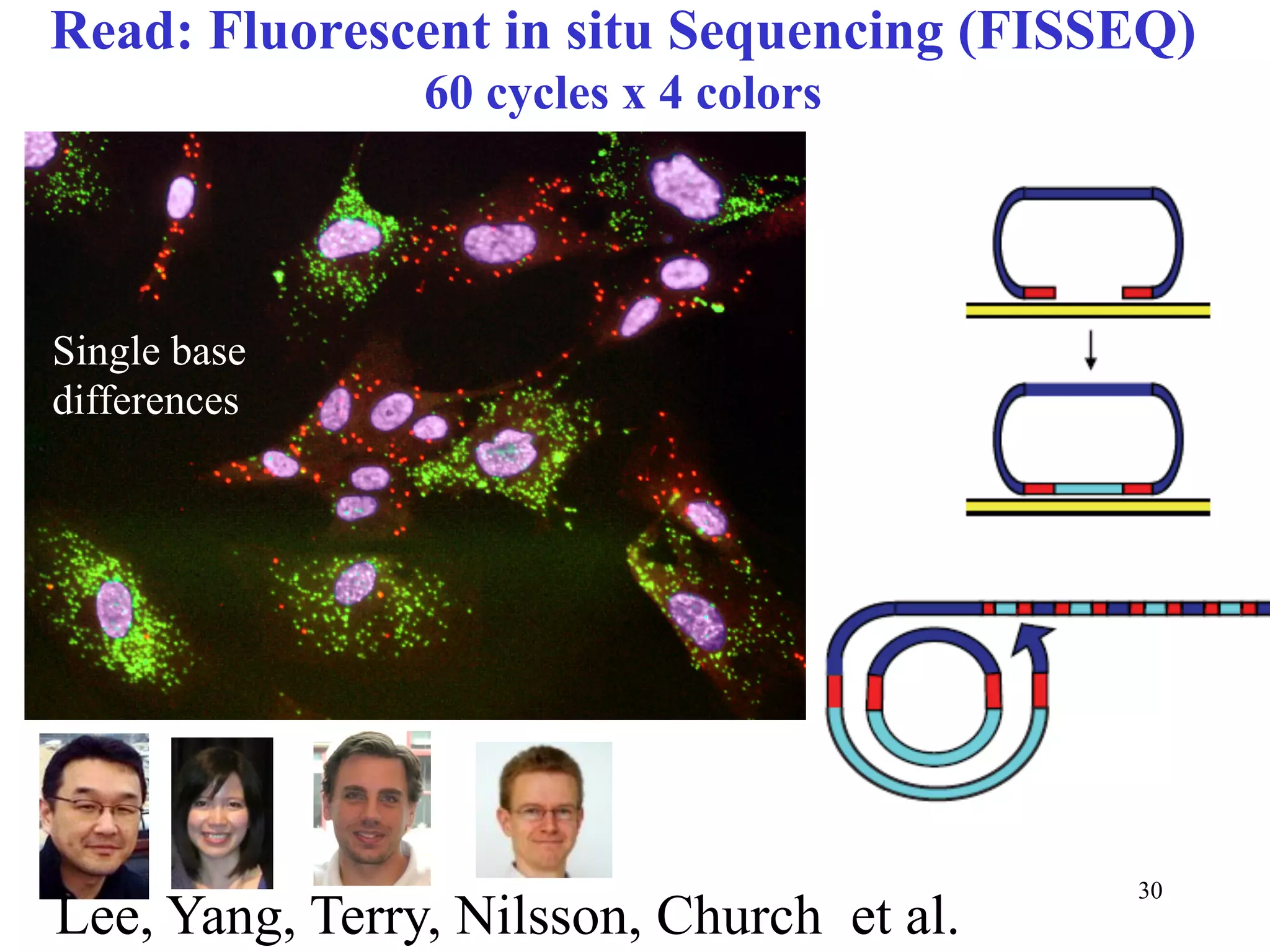
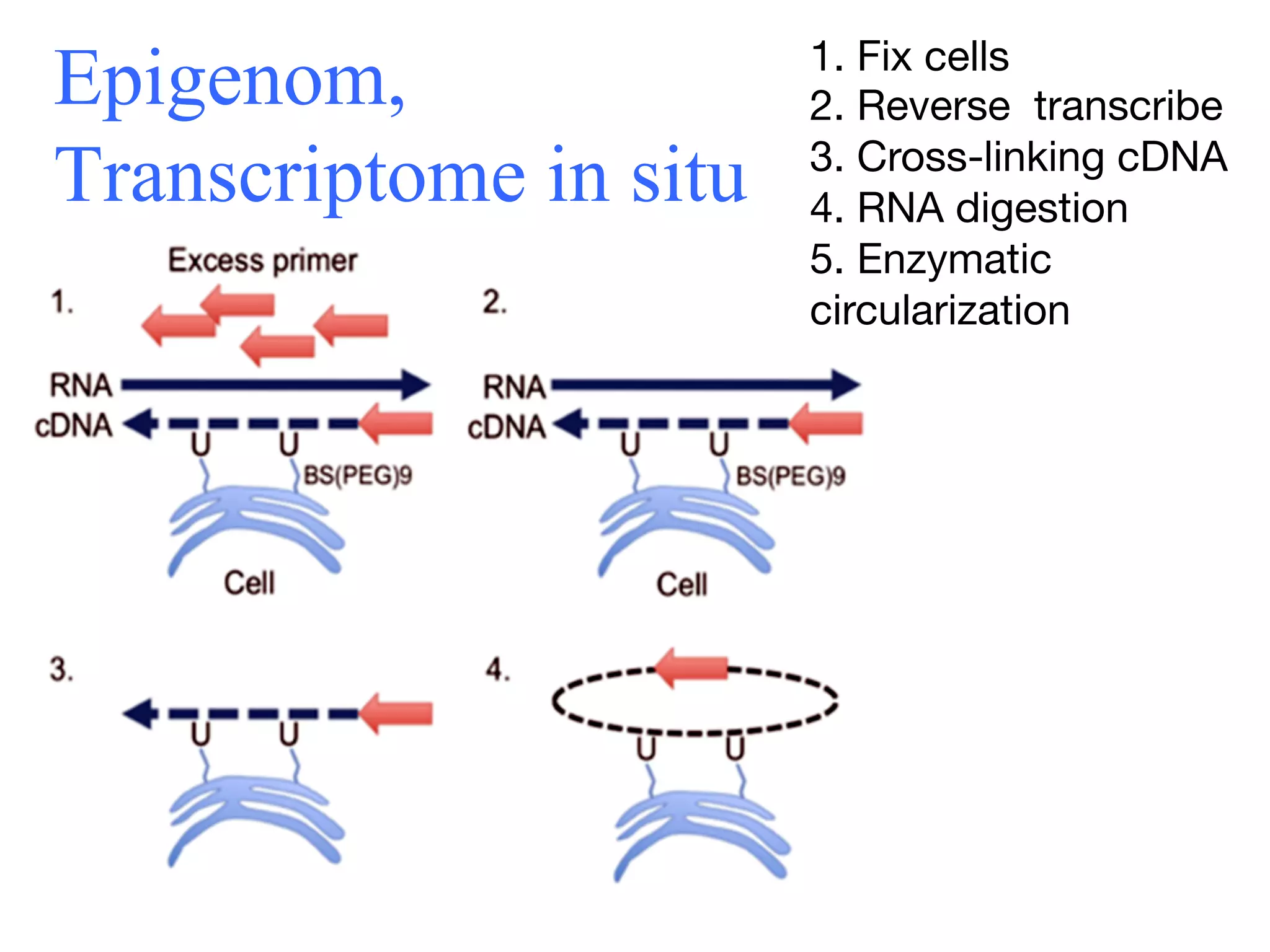
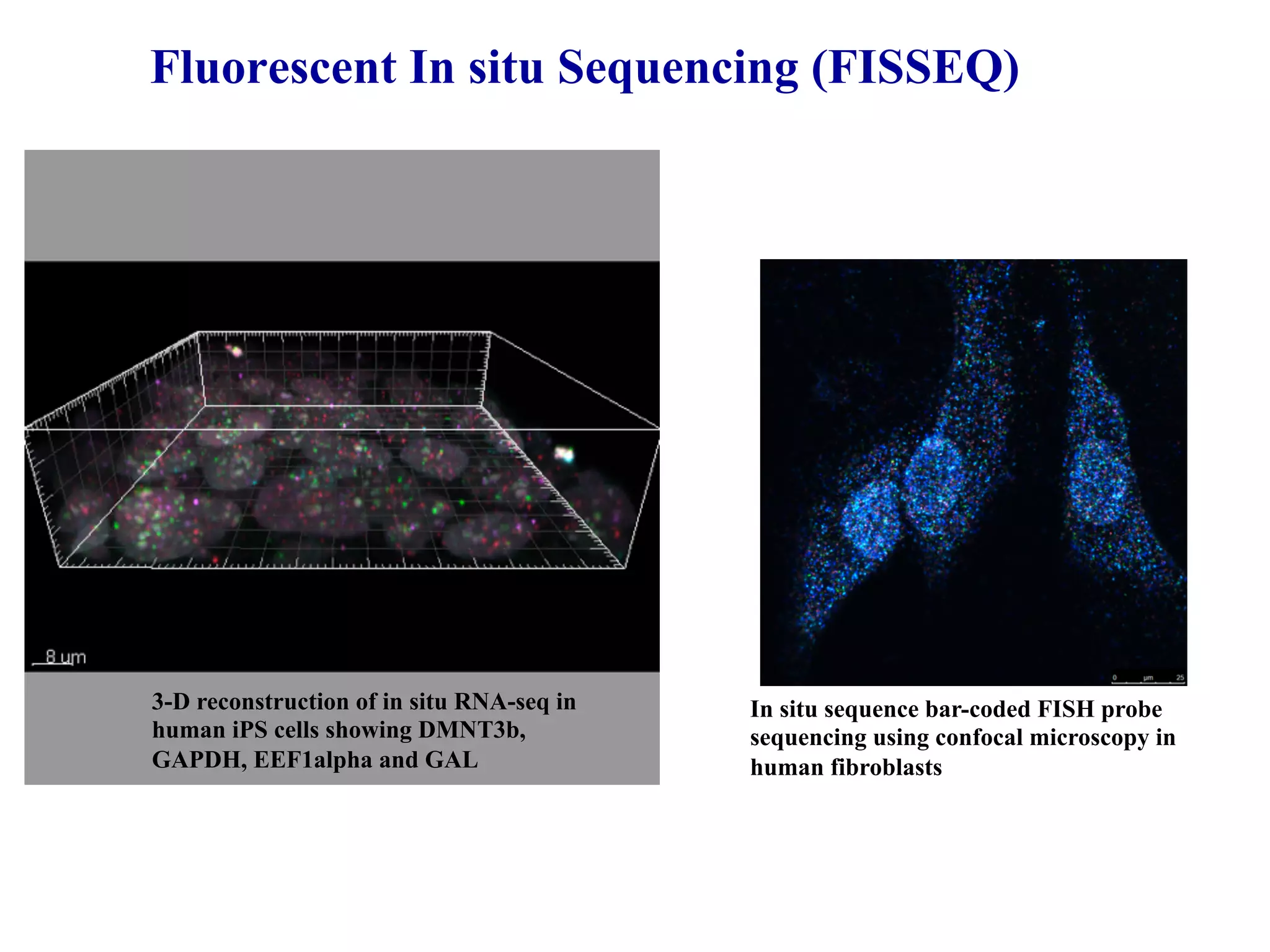
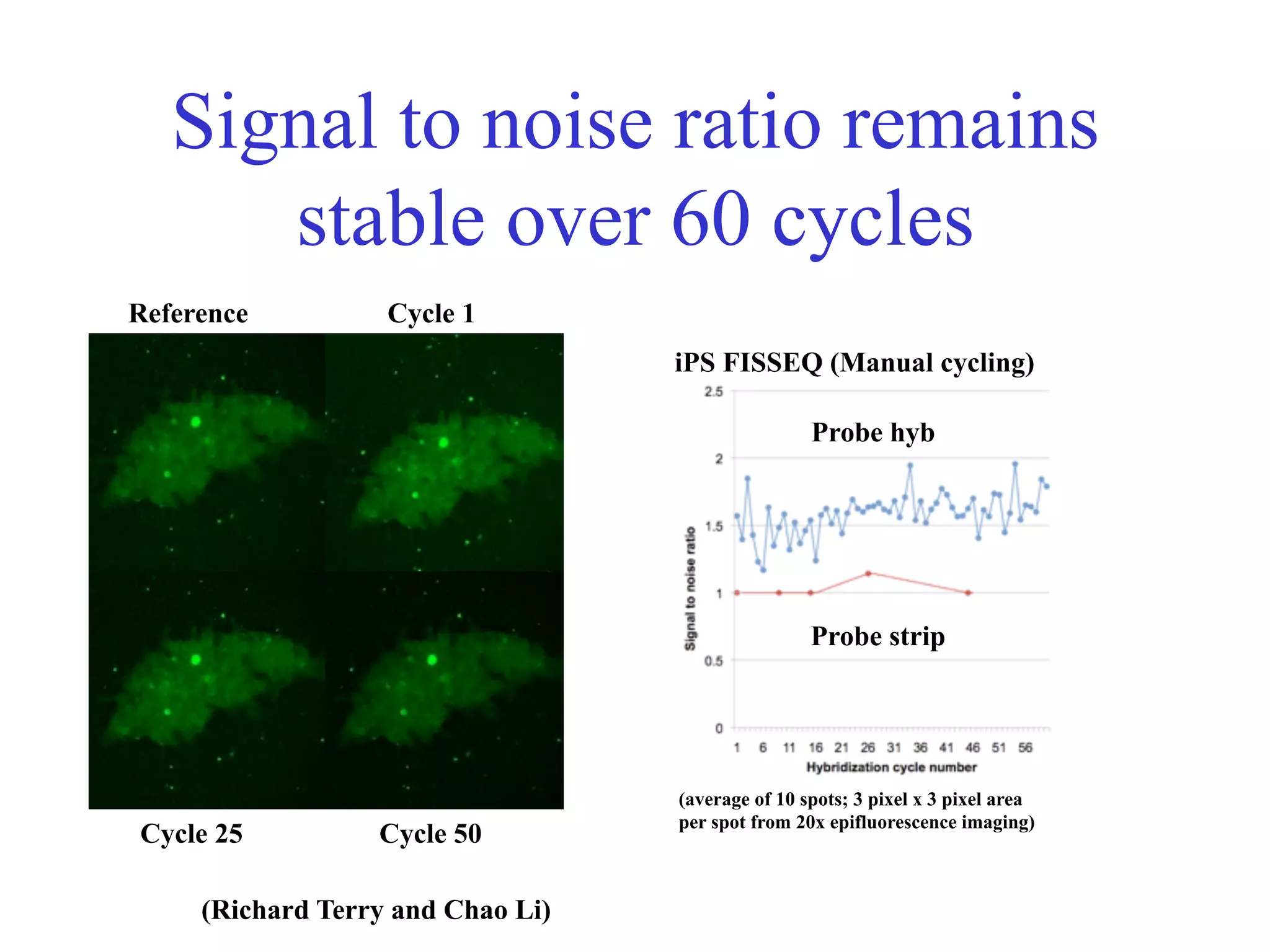
This document discusses standards and open access for genome, environment, and trait data. It covers reference materials, sequencing technologies, bioinformatics, and performance metrics. It provides examples of rare protective alleles and case studies showing how genome data can help diagnose and treat diseases. It also discusses technologies like nanopore sequencing, fluorescent in situ sequencing, and organ chips that can advance personalized and precision medicine.